Abstract
In the United States, ~1.4 million sporadic human Salmonella enterica infections occur annually, with an estimated 6% attributable to reptile exposure. Detection of Salmonella in reptiles can be challenging given the limitations among detection methods. We evaluated sampling and detection methods for S. enterica in a cross-sectional study of reptilian patients (n = 45) over the course of 13 mo. Two sampling methods (cloacal swabs, electrostatic cloth body-feet samples) and 3 detection methods (enriched culture, lateral flow immunoassay [LFI], real-time PCR) were compared using McNemar and Fisher exact tests. Results varied by species, sample type, and detection method. In total, 14 of 45 (33%) patients were positive by culture, 10 of 45 (22%), and/or 13 of 45 (29%) by rtPCR. Among rtPCR-positive results, cloacal swabs (12 of 45 [27%]) resulted in a higher detection than body-feet wipes (4 of 45 [9%]; p = 0.01). Among culture-positive results, shedding was most commonly detected after additional incubation at room temperature when testing cloacal swabs (9 of 45 [20%]). However, there was significant disagreement between sampling methods (cloacal vs. body-feet; p = 0.03). No samples were positive by LFI. In general, cloacal swabs yielded the highest test-positive rates, irrespective of testing method. Our study highlights the importance of using detection methods optimized for the sample being tested.
In the United States, ~1.4 million sporadic human Salmonella enterica infections occur annually, with an estimated 6% attributable to reptile or amphibian exposure.12,16 Of particular concern are outbreaks of Salmonella-associated disease commonly reported among children < 5 y of age who have had contact with small turtles (carapace length of < 10 cm)—accounting for an estimated 27% of Salmonella-associated disease within this age group in the UK. 13 Consequently, public health authorities, including the Centers for Disease Control and Prevention (CDC), recommend that all of these animals be considered carriers, with contaminated housing environments, until proven otherwise. 17
The Colorado State University Veterinary Teaching Hospital (CSU-VTH; Fort Collins, CO) conducts routine environmental surveillance for S. enterica as part of ongoing infection control efforts. A high prevalence of S. enterica has been reported for pet reptiles, estimated to be 48–50% in lizards, 7–10% in chelonians, and 51–83% in snakes. 14 However, on average, only 3.6% of environmental samples collected annually in the Avian, Exotic and Zoological (AEZ) Medicine Service (CSU-VTH; ~36 samples collected over 12 mo) are positive when using a previously reported enriched culture method. 5 This figure is in contrast to previous reports of 54 of 452 (12%) environmental-positive samples in a 3-mo period from the large and small animal hospital areas of the CSU-VTH. 5 As well, the figure differs from that detected in the environment of the reptile department at the Antwerp Zoo when using comparable methods (47 of 100 samples collected over a 4-mo period). 2 If infection rates in pet reptiles presented to the CSU-VTH are similar to those reported by others, we might expect a higher rate of environmental contamination than we have detected. Although this may indicate an effective infection control program or a lack of animal shedding, one must consider whether the detection methodology may be ineffective—the implications of which are important for the hospital, its personnel and patients, and patient owners.
Ensuring accurate detection of S. enterica among reptilian patients would allow for tailoring prevention efforts, both in-hospital and in the home environment. However, there are multiple challenges in detecting Salmonella in samples from these patients including obtaining appropriate samples and limitations with currently available laboratory methods. Of particular concern among reptilian patients is the challenge of obtaining fecal samples or cloacal swabs from small patients. As well, the culture of swab samples is often less sensitive than other sample types (e.g., 1-g or 10-g fecal samples). 10 It has been shown consistently that environmental sampling (Swiffer; Procter & Gamble, Cincinnati, OH) combined with multi-stage enrichment greatly enhances the sensitivity of Salmonella detection.11,15 As such, we hypothesize that wiping the body and feet from reptilian patients and culturing these samples in an effort to identify shedding may be an easy alternative to fecal samples or cloacal swabs with similar rates of detection.
Additionally, there are limitations in time and cost among commonly used methods, such as enriched culture and PCR, which, in our experience, typically require 48–96 and 24–72 h from submission to reporting, respectively, for the detection of S. enterica. 6 Lateral flow immunoassays (LFIs), marketed for use in food safety applications, offer a rapid, inexpensive, alternative detection method that has been shown to be effective in other veterinary relevant samples (e.g., equine feces and hospital environmental samples).6,7 By detecting shedding more rapidly in reptilian patients, it would be possible to implement prevention strategies sooner, thereby allowing earlier mitigation of the risk for transmission. We evaluated sampling methods and currently available detection methods for S. enterica in reptilian patients at a veterinary teaching hospital.
A cross-sectional study was undertaken from August 2014 through September 2015 to evaluate 2 patient sampling methods (cloacal swabs and body-feet wipes using electrostatic cloths) and 3 detection methods—enriched culture, LFI, and real-time PCR (rtPCR)—for S. enterica among reptile patients at the CSU-VTH. Our study was approved by the Colorado State University Institutional Animal Care and Use Committee (IACUC Protocol 14-1586A).
All reptilian patients (i.e., lizards, chelonians, and snakes, weighing >30 g) that were admitted to the CSU-VTH AEZ Medicine Service as inpatients or outpatients, during this 13-mo period, were eligible for study participation. Patient enrollment was restricted to times when members of the research team were overseeing clinical cases (MS Johnston) and available for sample collection (AC Fagre). On average, during the study period, the AEZ Medicine Service treated 19 reptiles per month (271 in 15 mo).
All enrolled patients were sampled by cloacal swab and body-feet wipe on the day of admission. Cloacal swabs were collected using clean examination gloves and a sterile swab (Puritan Medical, Guilford, ME), placed into a sterile conical tube. In addition, after donning clean examination gloves, a commercial electrostatic cloth (Swiffer; Procter & Gamble) was used to wipe the body and feet (where applicable), covering at least 80% of the body surface. After sampling, each cloth was placed in a sterile plastic bag with a gloved hand. Personnel changed gloves between sample collections. Patient samples were stored at 4°C until transported to the laboratory for processing, within 12 h of sample collection.
Our culture methods were based on standard methods used by the CSU Veterinary Diagnostic Laboratory (VDL; Fort Collins, CO) as part of the CSU-VTH Infection Control Program. Cloacal swabs were incubated overnight at 43°C in 9 mL of tetrathionate broth (TET; Becton Dickinson, Cockeysville, MD) containing iodine and brilliant green (MilliporeSigma, St. Louis, MO), vortexed, and streaked for isolation on xylose–lysine–tergitol 4 agar plates (XLT4; Hardy Diagnostics, Santa Maria, CA), and incubated overnight at 43°C. Cloths used to wipe the body and feet of patients were incubated overnight at 43°C in 90 mL of buffered peptone water (BPW; Becton Dickinson). The incubated BPW was then vortexed, and 1 mL added to 9 mL of TET broth containing iodine and brilliant green, and incubated for 24 h at 43°C. The TET broth was then vortexed, streaked for isolation on XLT4 agar, and incubated overnight at 43°C. The XLT4 plates from all sample types were evaluated after 18 h of incubation to identify suspect colonies, and again after an additional 24 h at room temperature. Suspect colonies were subcultured on trypticase soy agar containing 5% sheep blood (TSA; BD Diagnostic Systems, Sparks, MD) and incubated overnight at 43°C. The isolates were tested for agglutination using commercial polyvalent and O group–specific antisera (BD Diagnostic Systems). Agglutination with antisera was considered confirmatory for S. enterica identification. Final serovar determination was performed by the U.S. Department of Agriculture (USDA) National Veterinary Services Laboratories (Ames, IA). Antimicrobial susceptibility testing was performed on all S. enterica isolates by the Kirby–Bauer disc diffusion method against a panel of antimicrobial drugs (amikacin, ampicillin–clavulanate, ampicillin, ceftiofur, cephalothin, chloramphenicol, enrofloxacin, gentamicin, streptomycin, sulfonamides, and trimethoprim–sulfamethoxazole) following Clinical and Laboratory Standards Institute guidelines 8 at the CSU-VDL.
For all samples included in our study (regardless of culture results), a 2-mL aliquot of the TET broth (post-incubation) was frozen at −80°C and subsequently tested using rtPCR. After thawing the archived samples, DNA was extracted (MagMAX total nucleic acid isolation kit; Thermo Fisher Scientific, Waltham, MA), and rtPCR was conducted (MicroSEQ Salmonella spp. detection kit; Thermo Fisher Scientific). The primers for this rtPCR target a conserved region of the S. enterica genome and thus were designed to detect all Salmonella serovars. 1 All rtPCR testing was performed by personnel without knowledge of the culture results. A cycle threshold (Ct) value < 35 was used to classify test results as indicative of a positive test.
The LFI that we used (Reveal 2.0 for Salmonella test system; Neogen, Lansing, MI) was designed for point-of-use testing of food products to rapidly detect S. enterica contamination, 3 and has been evaluated for use with fecal and environmental samples collected from veterinary settings.6,7 This immunoassay uses antibodies specific for surface antigens of S. enterica and colloidal gold antibody conjugates incorporated into a lateral flow test strip. The TET broths were tested after 24 h of incubation using this system, a timing that matches the manufacturer’s directions.
Animal characteristics were summarized from patient histories and existing medical records. Data were entered into a computer spreadsheet, and entries were validated. Descriptive statistics were calculated, and comparisons were made using chi-squared test or Fisher exact test. Agreement was assessed descriptively among sampling methods and testing methods, and the McNemar test was used to evaluate agreement statistically. All statistical analyses were performed with SAS v.9.4 (SAS Institute, Cary, NC).
During the study period, 45 reptilian patients (20 lizards, 12 snakes, and 13 chelonians) were enrolled in the study on 37 different days (representing ~17% of all reptiles treated during the study period) and were sampled as per investigation protocols. The majority of patients were female, were obtained from pet stores, and were presented most commonly for a wellness evaluation or gastrointestinal disease (e.g., diarrhea, anorexia, stomatitis; Table 1).
Patient characteristics of study participants.
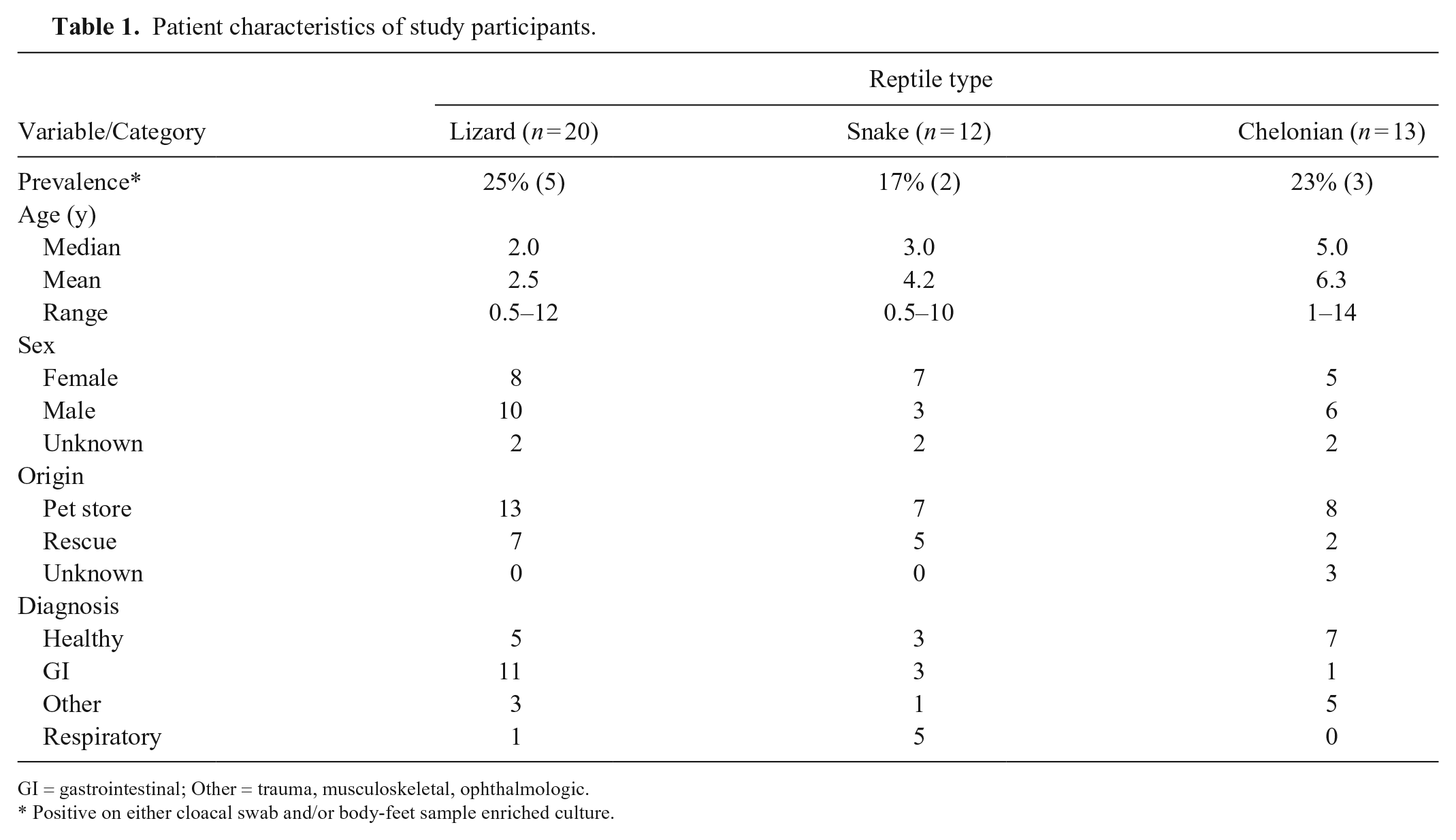
GI = gastrointestinal; Other = trauma, musculoskeletal, ophthalmologic.
Positive on either cloacal swab and/or body-feet sample enriched culture.
Overall, 22% (95% confidence interval [CI]: 12, 36) of these patients were culture-positive for S. enterica on at least one sample type (e.g., cloacal swab or body-feet wipe), and 29% (95% CI: 18, 43) were rtPCR-positive on at least one sample type (Table 2). In general, there was not a significant difference in the likelihood of being test-positive using either of these tests (culture or rtPCR; p = 0.26). The LFI that we used did not detect S. enterica in any patient sample tested.
Prevalence of Salmonella shedding among reptilian patients using multiple detection strategies.
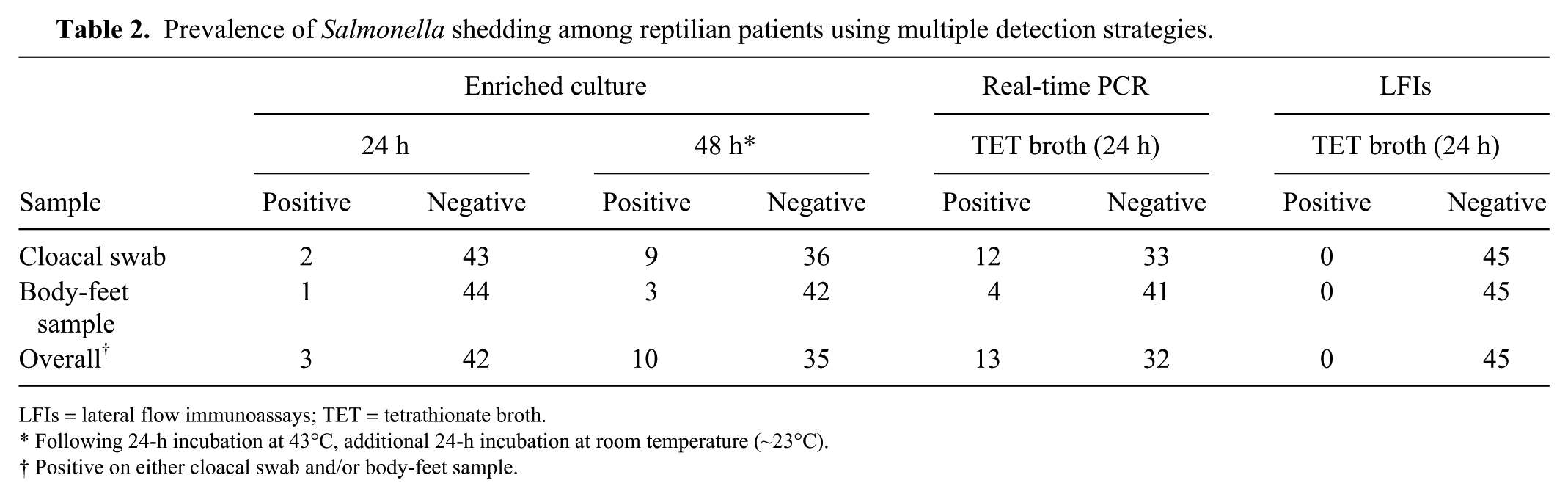
LFIs = lateral flow immunoassays; TET = tetrathionate broth.
Following 24-h incubation at 43°C, additional 24-h incubation at room temperature (~23°C).
Positive on either cloacal swab and/or body-feet sample.
Among culture-positive patients, shedding was most commonly detected using cloacal swab samples, particularly after an additional 24 h of sample incubation on XLT4 at room temperature (9 of 45 [20%]). Body-feet wipes had a significantly lower likelihood of being culture positive (3 of 45 [7%]; p = 0.03). Salmonella was isolated in both cloacal and body-feet samples from only 2 patients.
Among rtPCR-positive patients, shedding was most commonly detected when testing cloacal swabs (12 of 45 [27%]). Although the use of PCR increased the detection rate for both sampling methods (cloacal swab vs. body-feet wipe; Table 2), there was still significant disagreement between these methods as a result of the lower detection prevalence of body-feet wipe samples (4 of 45 [9%]; p = 0.01). Of the 4 patients with rtPCR-positive body-feet wipe samples, 3 were also rtPCR-positive by cloacal swab.
Various serovars were detected among positive patients (Table 3) and included some previously associated with zoonotic infections—specifically serovars Muenchen, IV 45:g,z51:-, and IV 50:g,z51:- (https://www.cdc.gov/salmonella/cotham-04-14/index.html; https://www.cdc.gov/salmonella/muenchen-05-15/index.html). Of the 10 isolates tested for antimicrobial susceptibility, only 1 was resistant to any of the antimicrobials tested (2 isolates were no longer viable for characterization at the time of susceptibility testing).
Characterization of Salmonella isolates from reptilian patients at the Colorado State University Veterinary Teaching Hospital (n = 10 isolates).
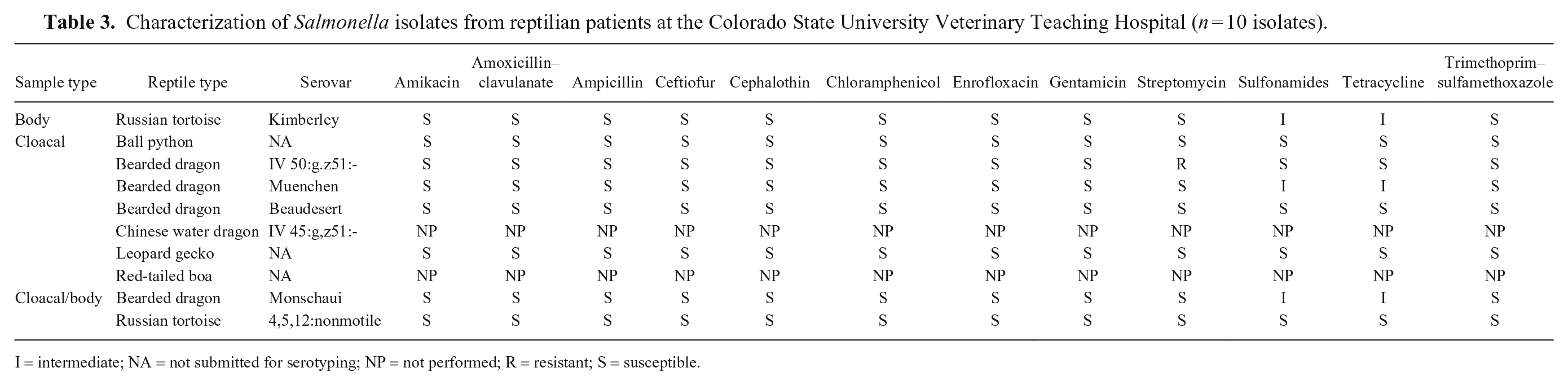
I = intermediate; NA = not submitted for serotyping; NP = not performed; R = resistant; S = susceptible.
The shedding prevalence among patients sampled in our study varied by species, sample type, and detection method used. Although shedding prevalence tended to be lower than in the literature (~50%),9,14 it still represents a significant risk to other patients and personnel. In general, cloacal swabs yielded the highest test-positive rates, irrespective of testing method, and are therefore recommended as a primary sample for the detection of S. enterica in reptilians, size permitting. However, body-feet wipe samples using an electrostatic cloth may be a viable alternative for very small patients. We did not evaluate recovery from fecal samples but requesting that pet owners provide fecal samples from the home environment may provide superior results to the use of body-feet wipes in these small patients.
Although there was good agreement between enriched aerobic cultures and rtPCR, testing using rtPCR yielded more positives, and results were generally available 24–48 h before culture results, which could have implications for management of Salmonella-positive patients. This difference in positive detection rate between enriched culture and rtPCR may be a result of the low numbers of organisms shed or may indicate the presence of strains that grow poorly under the conditions used in our study. It is also possible that the rtPCR test provided some false-positive test results, although a conservative Ct cutoff value was used when interpreting test results. Irrespective of the test used, it is generally considered important to obtain an isolate to allow for phenotypic characterization and further our understanding of the epidemiology of Salmonella in reptiles.
Interestingly, the culture-positive rate increased from 6% (3 of 45) to 22% (10 of 45) when cultures were allowed to stand for 24 h at room temperature (~23°C) after initial overnight incubation at 43°C. This observation is perhaps not surprising given that these isolates were derived from reptiles. We hypothesize that this may be the result of variation among Salmonella isolates recovered from ectothermic animals and their preferred thermal niche, which is lower than that used for incubation in our study. 4 This suggests that methods for the detection of Salmonella should be optimized for the types of samples that are being tested.
The LFI that we used was designed for point-of-use testing of food products to rapidly detect Salmonella enterica serovars commonly associated with foodborne disease, 3 which generally differ from those serovars commonly detected among reptiles. Although this is a limitation of this particular test, and this assay did not prove useful for samples obtained from reptiles, it has been shown to be effective in other veterinary relevant samples (i.e., equine fecal samples and hospital environmental samples).6,7
It is important to note that our study was based on convenience samples and thus may not be representative of all reptilian patients. Each patient was only sampled a single time. If reptiles shed intermittently, as is seen in other species, then this will decrease the ability to find a positive (i.e., decreased sensitivity) resulting in lower estimates of prevalence. Additionally, only one colony was selected from each sample for further characterization, precluding the ability to detect coinfection and potentially underestimating the diversity of Salmonella serovars present in a single patient. Finally, the observation that many positives were detected after an additional 24 h incubation at room temperature suggests that some strains may preferentially grow at lower temperatures and that our methods may be suboptimal for the detection of S. enterica in this particular patient group.
Although ours was a relatively small cross-sectional study, it provides the CSU-VTH with critical information with respect to managing this risk. We established a crude estimate of baseline prevalence among reptilian patients. Importantly, our study also highlights the importance of using detection methods optimized for the intended sample to be tested, and the potential impact this may have on effectively managing this risk among personnel, patients, and the facility.
Footnotes
Acknowledgements
We acknowledge the technical assistance of Denise Bolte, Christina Weller, Michael Richards, and Dr. Enrique Doster; and Drs. Sarah Higgins and Terry Campbell for assistance with sample collection.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported in part by the Colorado State University Center for Companion Animal Studies’ Young Investigator Grant. The funding body had no role in the design of the study, collection, analysis, and interpretation of data, or writing the manuscript.
