Abstract
Pets, including reptiles, have been shown to be a source of Salmonella infection in humans. Due to increasing popularity and variety of exotic reptiles as pets in recent years, more human clinical cases of reptile-associated Salmonella infection have been identified. However, limited information is available with regard to serotypes in different reptiles (turtles, snakes, and lizards) and antimicrobial resistance of Salmonella in pet reptiles. The current study was thus conducted to determine the prevalence of Salmonella colonization in pet reptiles. Salmonella organisms were isolated from 30.9% of 476 reptiles investigated. The isolation prevalences were 69.7% (23/33), 62.8% (27/43), and 24.3% (97/400) in snakes, lizards, and turtles, respectively. A total of 44 different Salmonella serovars were identified. Compared with S. Heron, Bredeney, Treforest, and 4,[5],12:i:-, S. Typhimurium isolates were resistant to many antimicrobials tested, and notably 61.1% of the isolates were resistant to cephalothin. The results indicated that raising reptiles as pets could be a possible source of Salmonella infection in humans, particularly zoonotic Salmonella serovars such as S. Typhimurium that may be resistant to antimicrobials.
Introduction
Salmonella are mainly considered to be food-borne pathogens transmitted through consumption of contaminated meat, eggs, vegetables, and fruits. 2,19,26,35,37 Pets, including reptiles, have been shown to be a source of infection in humans. 31,33,36,39,41 The disease usually causes abdominal cramps and self-limiting diarrhea. However, serious complications may occur, especially among children less than 10 years of age, elderly people, and immunocompromised patients. 30
In recent years, reports of salmonellosis from reptile reservoirs have gained more attention, especially in the United States, 1,6,7 where it has been estimated that 1.4 million human cases with Salmonella infections and approximately 600 associated deaths occur annually. 21 Due to the increasing popularity and variety of exotic reptiles as pets in recent years, more human clinical cases with reptile-associated Salmonella infection have been identified. The Centers for Disease Control and Prevention (CDC) estimated that over 50,000 cases of reptile-associated salmonellosis occurred in 1996. 22 The results from a previous study 23 further suggested that reptile and amphibian exposure is associated with about 74,000 Salmonella infections in the United States every year. According to results of Salmonella surveillance in the United States during 1987–1997, 25 7 out of the top 20 serovars were reptile-associated Salmonella serovars, including S. IV 48:g,z51:-(previously S. Marina), S. Flint (now known as S. IV 50:z4,z23:-), S. Kintambo, S. Wassenaar (now known as S. IV 50:g,z51:-), S. Ealing, S. Carrau, and S. Abaetetuba.
Recently, Salmonella infection in humans and animals has become an important issue in Taiwan, and antimicrobial-resistant Salmonella infections have been identified. 8,13,20 According to a previous report, 20 the leading 4 Salmonella serovars recovered from human patients were S. Typhimurium, S. Enteritidis, S. Stanley, and S. Schwarzengrund. Furthermore, multi-antimicrobial–resistant S. Choleraesuis has gained much attention in Taiwan. 8,10,12,13 Although these serovars are mainly food-borne pathogens, pet exposure could be also considered as another way of transmission. Pet reptiles are increasingly popular in Taiwan. However, there have been no epidemiological studies conducted in these animals concerning their colonization and excretion of zoonotic pathogens. In pet reptiles, limited information has been reported with regard to antimicrobial resistance of Salmonella worldwide. A previous study 32 in Trinidad indicated that 100%, 66.7%, 66.7%, and 33.3% of the Salmonella isolates from reptiles were resistant to streptomycin, cephalothin, neomycin, and gentamycin, respectively. The current study was thus conducted to determine the epidemiological patterns of Salmonella colonization in reptiles collected from various sources in Taiwan.
Materials and methods
Sample collection
From September 2005 through April 2006, a total of 476 samples were collected by convenience sampling, including 418 cloacal swabs obtained from smuggled pet reptiles kept in Taipei Zoo (Taipei, Taiwan), 20 fecal samples of pet reptiles from pet shops, and 38 fecal samples of reptiles presented to the Veterinary Teaching Hospital, National Chung Hsing University (Taichung, Taiwan). At the time of the sampling, information with regard to health status, whether the animals showed signs of diarrhea, was obtained through the records offered by the Zoo, by veterinarians in the hospital, or by direct inspection in pet shops. Clinical manifestations reported for ill animals included nasal discharge, ocular discharge, anorexia, hepatitis, or parasite infection. According to diet patterns, the animals were categorized as carnivorous, omnivorous, and herbivorous. In the study area, not all pet shops could be investigated, and only pet reptiles from pet shops where owners provided permission were sampled and further investigated.
Cultivation, identification, and serotyping of Salmonella isolates
Cultivation of bacteria from the cloacal swabs or fecal samples was performed according to procedures by International Organization for Standardization 6579 (http://www.iso.org/iso/catalogue_detail.htm?csnumber=40377) with minor modifications. Each cloacal swab was first soaked in 4.5 ml of buffered peptone water a for transportation. The swabs were then incubated at 37°C for 18–24 hr with 3.5 relative centrifugal force (rcf) mild shaking. Five hundred microliters of the incubated product were transferred from each tube into 4.5 ml of Rappaport–Vassiliadis R10 (RV) broth a and incubated for 18–24 hr at 42°C. One loopful of the incubated broth was streaked onto duplicate plates of xylose–lysine–deoxycholate (XLD) agar a and Salmonella–Shigella (SS) agar. a The plates were incubated at 37°C for 18–24 hr. Simultaneously, 30 μl of incubated RV broth were inoculated onto the surface of the modified semi-solid RV (MSRV) medium. a The plates were incubated at 42°C for 18–24 hr and examined for growth and development of a migration zone. When a migration zone was observed, specimens sampled from the rim of the migration zone in the MSRV plates were subcultured onto XLD and SS agars and then incubated at 37°C for 18–24 hr. For the identification of suspected Salmonella isolates, subculture from a single colony pick was used for the agglutination reaction using polyvalent somatic (O) antisera b for Salmonella. 32 Only isolates with a positive result by agglutination test were tentatively confirmed to be Salmonella. The Salmonella isolates were stored at −80°C after subculture on XLD agar for further serotyping and testing of antimicrobial resistance. Salmonella isolates were serotyped with specific O- and H-antisera, b , c and serovar identification was carried out according to the principles of the Kauffmann–White scheme. 4,29
Antimicrobial susceptibility test
All the Salmonella isolates were tested for antimicrobial susceptibility to a panel of 12 antimicrobials by agar disc diffusion test on Mueller–Hinton II agar, d following the Clinical and Laboratory Standards Institute (CLSI) guidelines. The following antibiotic discs e were used in this study: ampicillin (AMP), 10 μg; cephalothin (KF), 30 μg; chlor-amphenicol (C), 30 μg; ciprofloxacin (CIP), 5 μg; enrofloxacin (ENR), 5 μg; gentamicin (CN), 10 μg; nalidixic acid (NA), 30 μg; nitrofurantoin (F), 300 μg; norfloxacin (NOR), 10 μg; streptomycin (S), 10 μg; trimethoprim–sulfamethoxazole (SXT), 25 μg; and tetracycline (TE), 30 μg. The isolates were classified as susceptible (S), intermediate (I), or resistant (R) following the zone diameter standards by the National Committee for Clinical Laboratory Standards. 38 Escherichia coli ATCC15922 was utilized for internal quality control.
Statistical analysis
For each categorical group, percentage of Salmonella isolation or antimicrobial resistance was calculated. To test if percentages among different categorical groups were statistically different, chi-square (χ 2 ) test or Fisher's exact test (when the expected number of observations was less than 5) wasperformed using SAS version 9.1 for Windows. f A P-value less than 0.05 was considered to be statistically significant.
Results
Isolation
Overall, Salmonella organisms were isolated from 147 (30.9%) of 476 reptiles investigated, and a total of 358 isolates were obtained. The use of both XLD agar and SS agar plates for Salmonella isolation resulted in the recovery of more than 1 isolate from some reptile samples. The isolation rates were 69.7% (23/33), 62.8% (27/43), and 24.3% (97/400) in snakes, lizards, and turtles, respectively. The prevalence in turtles was significantly lower than that of snakes (odds ratio: 0.14, 95% confidence interval [CI]: 0.06–0.32) or lizards (odds ratio: 0.19, 95% CI: 0.09–0.38).
Serotyping
The most frequently isolated Salmonella was serogroup B (24.0%). Other serogroups of Salmonella included serogroup I (11.5%), C2 (8.1%), and C1 (5.9%). Although serovars could not be determined for 87 isolates due to the limited spectrum of specific O- and H-antisera, a total of 44 different Salmonella serovars were identified in reptiles (Table 1). Moreover, 15.0% (22/147) of reptiles were co-colonized with different Salmonella serovars. Overall, the most common Salmonella serovars were S. Heron (8.9%), S. Bredeney (7.8%), S. Typhimurium (5.0%), and S. Treforest (5.0%). It was further observed that the distribution of various Salmonella serovars was different in snakes, lizards, and turtles (Table 1).
The comparison of percentages (number of isolates) of Salmonella serovars in turtles, lizards, and snakes in Taiwan (%). *
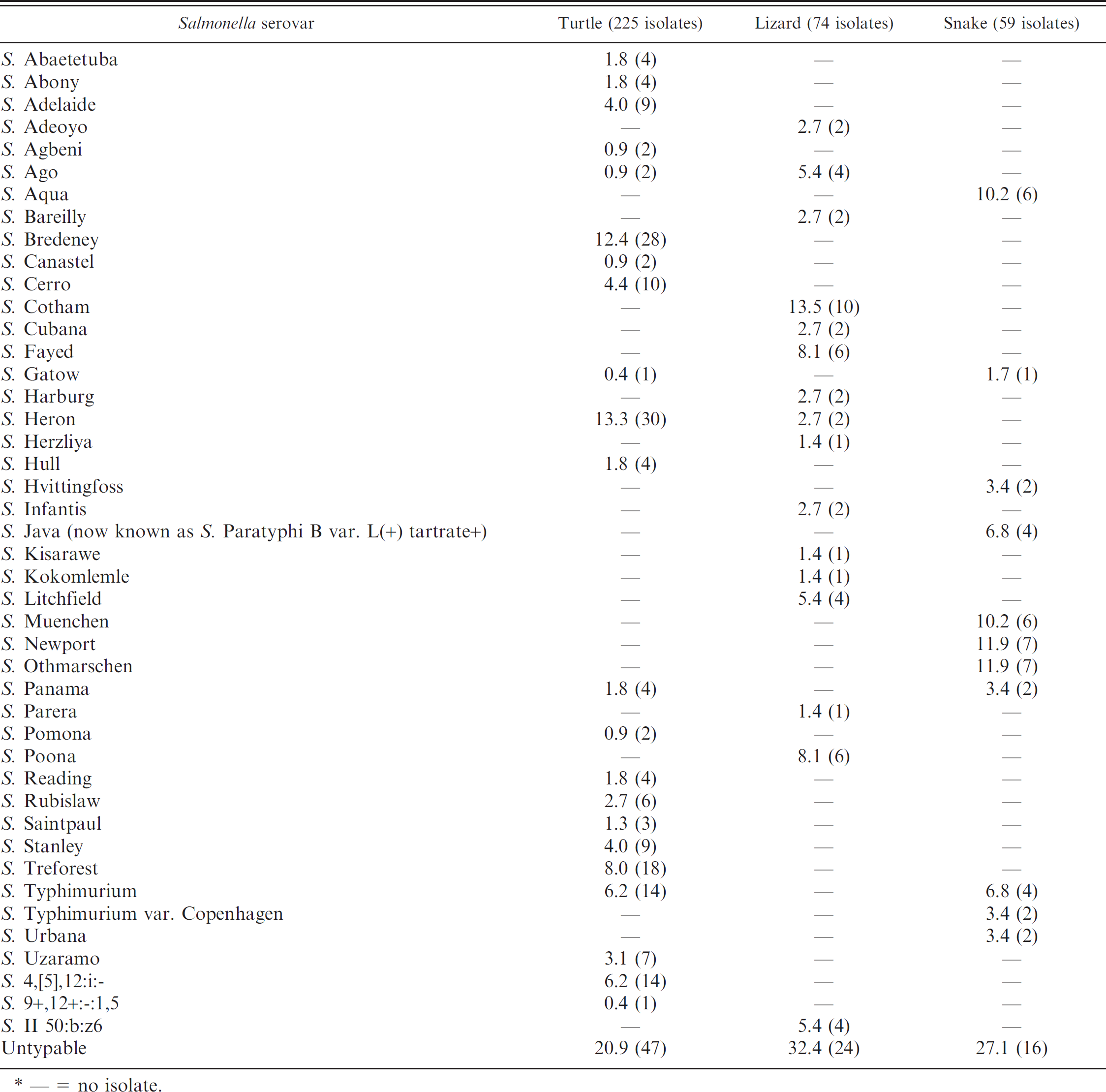
— = no isolate.
Analysis of factors associated with Salmonella infection
The results are listed in Table 2. The isolation prevalences were 66.0% (31/47), 42.9% (42/98), and 21.7% (68/313) in the carnivorous, omnivorous, and herbivorous reptiles, respectively. Such a difference was statistically significant (P < 0.05). In carnivorous animals, the most common serovars were S. Newport (8.9%) and S. Othmarschen (8.9%), followed by S. Muenchen (7.6%), S. Aqua (7.6%), S. Stanley (5.1%), S. Java (now known as S. Paratyphi B var. L(+) tartrate+; 5.1%), S. Poona (5.1%), and S. Ago (5.1%). In omnivorous animals, the most frequent serovar was Salmonella 4,[5],12:i:- (15.6%), and other common serovars included S. Typhimurium (13.3%) and S. Cotham (11.1%). In herbivorous animals, the most frequent serovar was S. Heron (17.5%), followed by S. Bredeney (14.0%) and S. Treforest (10.5%). The prevalence in turtles was significantly higher than that in tortoises (40.3% vs. 20.8%, odds ratio = 2.57, P < 0.05). There was no significant difference of prevalence of Salmonella colonization between animals with and without obvious clinical manifestations. Among sick animals, individuals with diarrhea were more likely to excrete Salmonella organisms in feces than animals without diarrhea (55.6% vs. 31.9%, odds ratio: 2.67, 95% CI: 0.57–12.96, P > 0.05).
Risk factors associated with Salmonella infection.
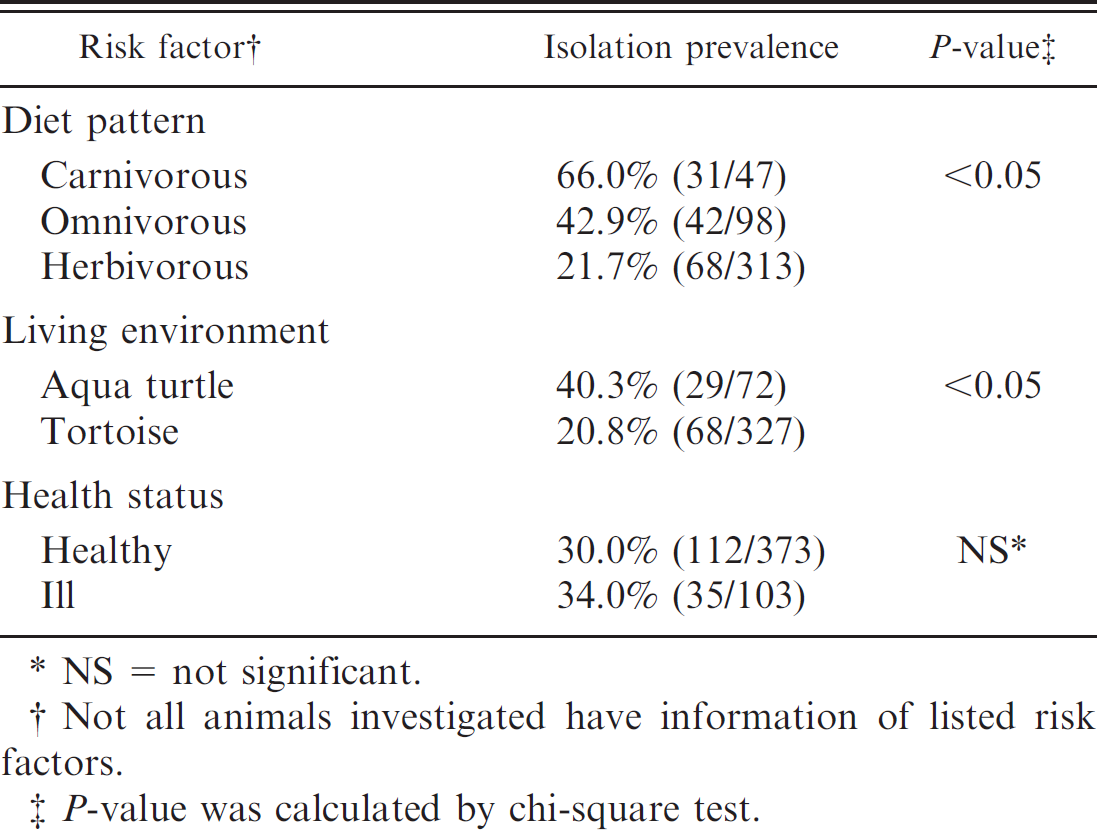
NS = not significant.
Not all animals investigated have information of listed risk factors.
P-value was calculated by chi-square test.
Antimicrobial susceptibility testing
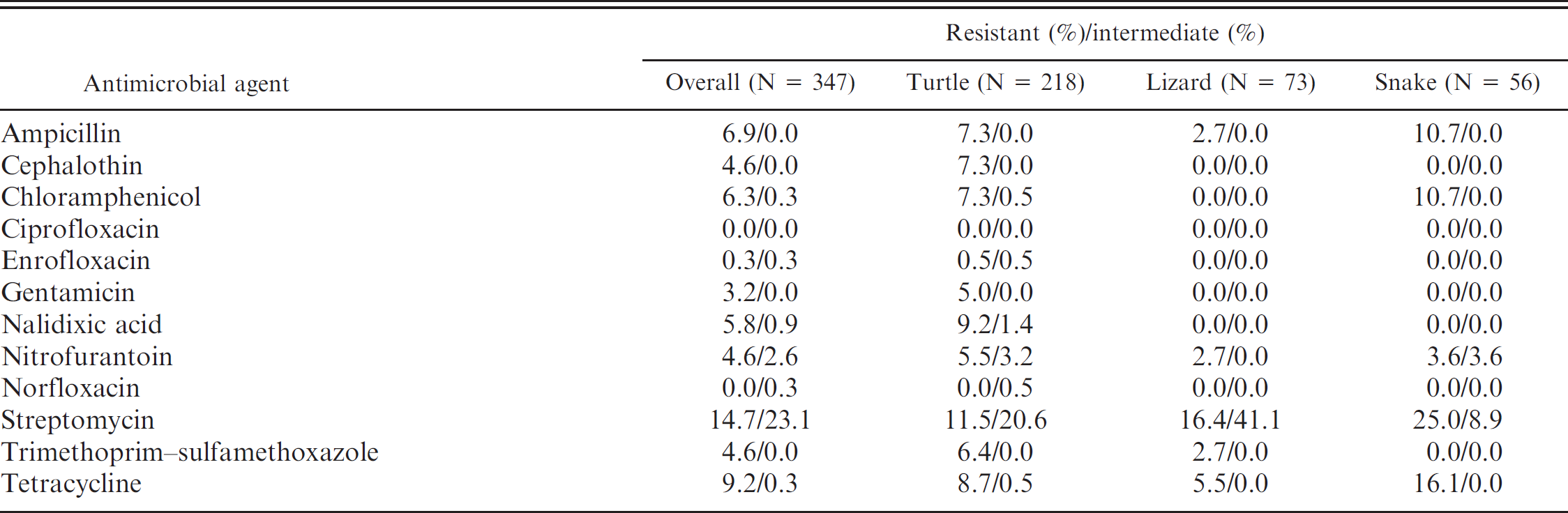
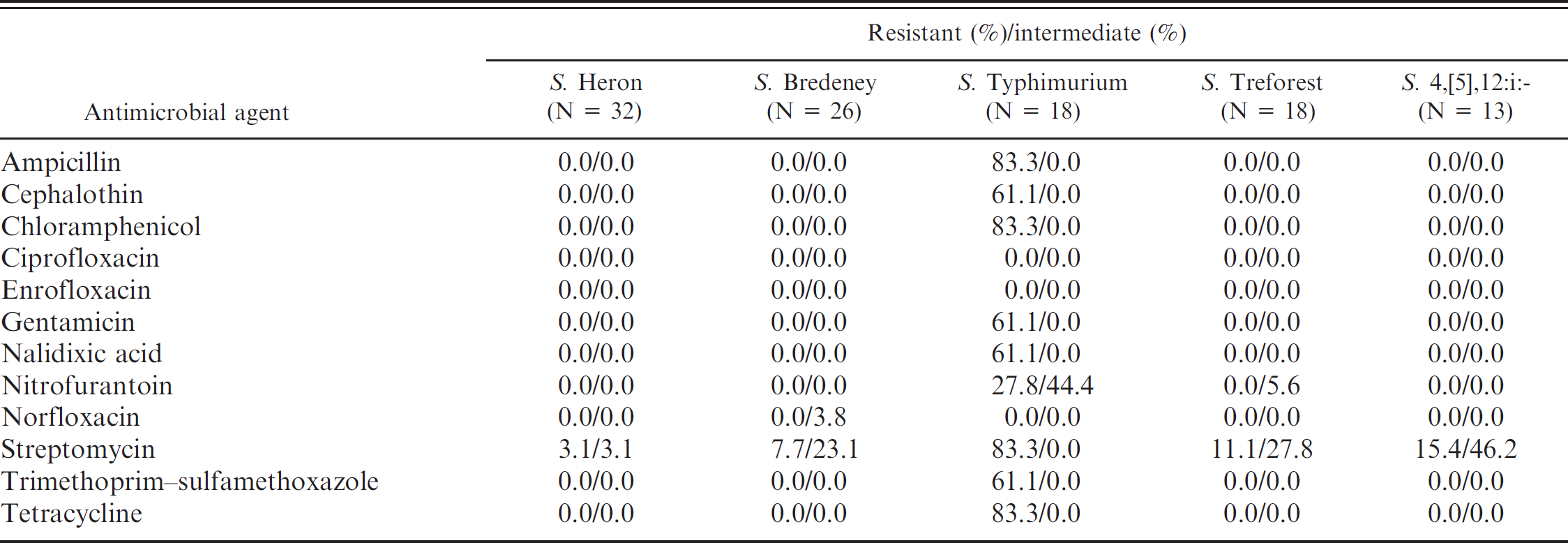
Except for 11 Salmonella isolates that contained major contamination problems, antimicrobial susceptibility testing by disk diffusion test was conducted on 347 isolates. As shown in Table 3, although the overall resistance percentages were not very high, it was found that 24 (6.9%) isolates were resistant to 4 or more different antimicrobial agents tested in the current study. The highest resistance percentages were related to streptomycin (14.7%, 51/347) and tetracycline (9.2%, 32/347). After comparing the antimicrobial resistance percentages in different animal orders, the percentage of streptomycin resistance remained the highest in turtles, lizards, and snakes (Table 3). More kinds of antimicrobial resistance were observed in isolates from turtles, as compared with those from lizards and snakes (Table 3). Furthermore, 7.3% of the Salmonella isolates from turtles were resistant to cephalothin. Nalidixic acid resistance was present only in isolates from turtles (9.2%). Among the Salmonella serovars recovered from reptiles that are related to human infections in Taiwan, it was observed that S. Typhimurium isolates were resistant to a wide range of antimicrobials tested, as compared with resistance exhibited by S. Heron, S. Bredeney, S. Treforest, and S. 4,[5],12:i:- isolates (Table 4). The percentage of streptomycin resistance present in S. Typhimurium isolates reached 83%, and 61% of S. Typhimurium isolates were resistant to cephalothin.
Antimicrobial resistance prevalence of 347 Salmonella isolates obtained from 476 animals.
Discussion
To the authors′ knowledge, this is the first investigation of prevalence of Salmonella colonization in pet reptiles in Taiwan. The results indicated high prevalence (30.9%) of Salmonella colonization in these animals, especially in snakes (69.7%) and lizards (62.8%). This observation was in accordance with previous findings. 14–16,18,24,34 The prevalence of Salmonella infection in turtles varies in different studies, ranging from 2.6% to 72.2%. 18,24,27 In the current study, Salmonella was recovered from 24.3% of the turtles tested. Seasonal behaviors, such as hibernation, have been speculated to result in a low isolation rate of Salmonella in turtles. 18 Nevertheless, a previous study 14 observed that low isolation rate of Salmonella in turtles (10.3%) could also be obtained in summer. It was considered that season of sampling could be one of the factors associated with the prevalence of Salmonella colonization in turtles.
Antimicrobial resistance of the top 5 Salmonella serovars identified from 347 Salmonella isolates in 476 animals.
The majority of the study subjects were convenience samples collected from the smuggled pet reptiles housed in the Taipei Zoo. Therefore, information with regard to imported countries was unavailable. As different prevalence of Salmonella colonization can be seen in animals across different geographical locations, serovars identified in the present study could not be reasonably compared with data from other countries. For example, a U.S. study 25 identified that the top 7 reptile-associated Salmonella serovars in the United States were Salmonella IV 48:g,z51:-, S. Flint (now known as S. IV 50:z4,z23:-), S. Kintambo, S. Wassenaar (now known as S. IV 50:g,z51:-), S. Ealing, S. Carrau, and S. Abaetetuba, which did not correlate with the top 4 serovars found in the present study.
In the current investigation, a total of 44 different Salmonella serovars were identified. Of major importance, S. Heron, S. Bredeney, S. Typhimurium, and S. Treforest, which are recovered from human cases with Salmonella infections in Taiwan, were the top 4 Salmonella serovars identified in reptiles. Similar to the previous findings, the present study indicated that 15.0% (22/147) of reptiles were co-colonized with different Salmonella serovars. 18,39 According to the co-infection prevalence and using combinations of culture media for Salmonella isolation in the present study, it may imply that 1 serovar dominates most of the animals. Nevertheless, it will be of major interest to understand whether reptiles with co-infection are more likely to excrete Salmonella in the environment, and to threaten human health.
In the present study, except for 4 isolates, the majority of the salmonellae belonged to Salmonella enterica subspecies II; the remaining ones belonged to S. enterica subsp. enterica. These results differed from observations reported in other studies, which found mainly S. enterica subsp. II, IIIa, IIIb, IV, VI, and S. bongori associated with cold-blooded animals and their living environment. 5 Although the reason for such a discrepancy is still unclear, S. enterica subsp. enterica was found to be common in reptiles in several studies. 15,18,24,28 The present research findings may imply that reptiles could play a role as important vectors for Salmonella species commonly associated with human disease, and further research needs to be conducted to determine if water treatment and food handling could be associated with this observation.
It has been suggested that the distribution of Salmonella in reptiles may reflect their living environments. 24 In the present study, the isolation rate of Salmonella in aqua turtles was significantly higher than that in tortoises. Salmonella has high survival rates in aquatic environments and is frequently isolated from water sources. 3,9,11,40 A previous study further indicated that Salmonella can persist in living environments for up to 10 months. 17 Therefore, if the water is contaminated with Salmonella, it could be a source for Salmonella transmission in aquatic animals, and eventually to humans.
Although the overall resistance prevalence was not very high (0–14.7%), S. Typhimurium was the serovar with the widest range of antimicrobial resistance. High resistance percentages (61.1–83.3%) in S. Typhimurium were observed to ampicillin, chloramphenicol, gentamicin, streptomycin, trimethoprim–sulfamethoxazole, and tetracycline, which are all traditional antimicrobial agents used clinically in humans. Cephalothin is frequently used for the treatment of bacterial infection with multidrug resistance; however, 61.1% of the S. Typhimurium isolates were resistant to cephalothin. This result is similar to a previous report, 32 which showed 66.7% of Salmonella isolates resistant to cephalothin in reptiles. Although none of the S. Typhimurium isolates in reptiles were resistant to fluoroquinolones, such as norfloxacin, enrofloxacin, and ciprofloxacin, 61.1% of them were resistant to nalidixic acid. It is of major importance that nalidixic acid resistance was present only in turtles. As the majority of reptile-associated Salmonella cases have been from contact with turtles in the United States, such resistance could impact the treatment of human cases associated with turtle exposure in Taiwan.
In the current study, serovars for 24.3% of the isolates could not be determined. Although the authors′ laboratory has the capacity to identify most known Salmonella serovars, the result may imply the limited spectrum of specific O- and H-antisera in the authors′ laboratory for identification of all serotypes. It is reasonable to consider that unrecognized serovars can be identified in reptiles. These untypable isolates are likely to be uncommon serovars, and will need to be further identified. According to a study conducted in Japan, 24 although biochemical confirmation was done for Salmonella isolates in reptiles, 58 of the 112 Salmonella isolates tested were untypable with commercial antisera. These reasons could explain the percentage of untypable isolates in the present study.
Based on these observations, it was concluded that raising reptiles as pets could be a possible source for Salmonella infections in humans and may eventually threaten human health due to antimicrobial-resistant Salmonella. The results offer valuable information to educate people who are raising or are considering raising reptiles as pets and can be applied in future risk assessment of Salmonella infection in humans.
Acknowledgements
This project was supported by grant NSC 94-2313-B-005-047 from the National Science Council, Taiwan. The authors greatly appreciate the major assistance from the Taipei Zoo, Taipei, Taiwan, for collecting reptile specimens.
Footnotes
a.
Difco™, BD, Franklin Lakes, NJ.
b.
Serotest®, S&A Reagents Lab Ltd., Bangkok, Thailand.
c.
Denka Seiken Co. Ltd., Tokyo, Japan.
d.
BBL™, BD, Franklin Lakes, NJ.
e.
OXOID Ltd., Cambridge, UK.
f.
SAS Institute Inc., Cary, NC
