Abstract
Avibacterium paragallinarum is the causative agent of infectious coryza, a highly contagious respiratory disease in chickens. Given its fastidious nature, this bacterium is difficult to recover and identify, particularly from locations colonized by normal bacterial flora. Standard PCR methods have been utilized for detection but are labor-intensive and not feasible for high-throughput testing. We evaluated a real-time PCR (rtPCR) method targeting the HPG-2 region of A. paragallinarum, and validated a high-throughput extraction for this assay. Using single-tube extraction, the rtPCR detected 4 A. paragallinarum (ATCC 29545T and 3 clinical) isolates with a limit of detection (LOD) of 10 cfu/mL and a PCR efficiency of 89–111%. Cross-reaction was not detected with 33 non–A. paragallinarum, all close relatives from the family Pasteurellaceae. Real-time PCR testing on extracts of 66 clinical samples (choana, sinus, or trachea) yielded 98.2% (35 of 36 on positives, 30 of 30 on negatives) agreement with conventional PCR. Duplicate samples tested in a 96-well format extraction in parallel with the single-tube method produced equivalent LOD on all A. paragallinarum isolates, and 96.8% agreement on 93 additional clinical samples extracted with both procedures. This A. paragallinarum rtPCR can be utilized for outbreak investigations and routine monitoring of susceptible flocks.
Keywords
Introduction
Infectious coryza (IC) is an acute, severe, respiratory disease of chickens associated with high morbidity in affected flocks, and is caused by Avibacterium paragallinarum.4,10,11 Clinical signs include ocular and nasal discharge, rhinitis, and sinusitis reported as “swollen heads,” as well as anorexia.2,3,9 This highly contagious disease affects chickens worldwide, resulting in poor growth in young birds and reduced egg production in layers.1,9,11 Concurrent respiratory agents, including Mycoplasma synoviae, M. gallisepticum, Pasteurella sp., and infectious bronchitis virus (IBV; species Avian coronavirus), as well as stress factors, can exacerbate disease.10,11
A. paragallinarum is a fastidious gram-negative bacterium in the Pasteurellaceae family.2,5 Given its slow growth rate and a need for specialized laboratory media and conditions, the organism is difficult to detect by culture, particularly from sites colonized by normal flora.1,5 Recovery has also been shown to be impacted by delays in sample processing. 2 If suspect organisms are recovered, conventional PCR is often required to confirm the identity of A. paragallinarum given its limited reactivity in routine biochemical tests.4,5 Outbreaks in California poultry flocks 6 highlight the need for a robust high-throughput method for detecting A. paragallinarum in large-volume submissions, particularly when concurrent pathogens and opportunistic bacteria are present. We evaluated a previously published real-time PCR (rtPCR) assay for use in our laboratory and validated a high-throughput extraction method on swab samples from affected sites that can facilitate rapid diagnosis of IC.
Materials and methods
One strain of A. paragallinarum (ATCC 29545T) and 3 clinical isolates (CI 1 from a California commercial layer flock in 2010, CI 2 from a California backyard flock in 2016, and CI 3 from a California broiler commercial flock in 2009) were utilized to establish the limit of detection (LOD) for the assay (Fig. 1). The 3 clinical A. paragallinarum isolates were identified by nicotinamide adenine dinucleotide (NAD)-dependent growth, biochemical nonreactivity, and a positive result in a conventional PCR specific for A. paragallinarum. 4 Bacterial species used for specificity included clinical isolates of closely related organisms of NAD-dependent A. volantium (n = 6) and A. avium (n = 3), as well as NAD-independent A. gallinarum (n = 12). Additional Avibacterium sp. isolates were used that could only be identified to the genus level and represented both NAD-dependent and -independent forms (n = 15). The isolates used for specificity testing were identified using biochemical reactions, growth conditions, a matrix-assisted laser desorption/ionization time-of-flight identification system (MALDI-TOF MS, Microflex; Bruker Daltonics, Bremen, Germany), a negative result in the conventional A. paragallinarum PCR, 4 and partial 16S ribosomal DNA sequencing. The specificity testing included organisms frequently recovered from diseased respiratory sites, namely Escherichia coli (strain ATCC 25922) and Proteus vulgaris (strain ATCC 6896).
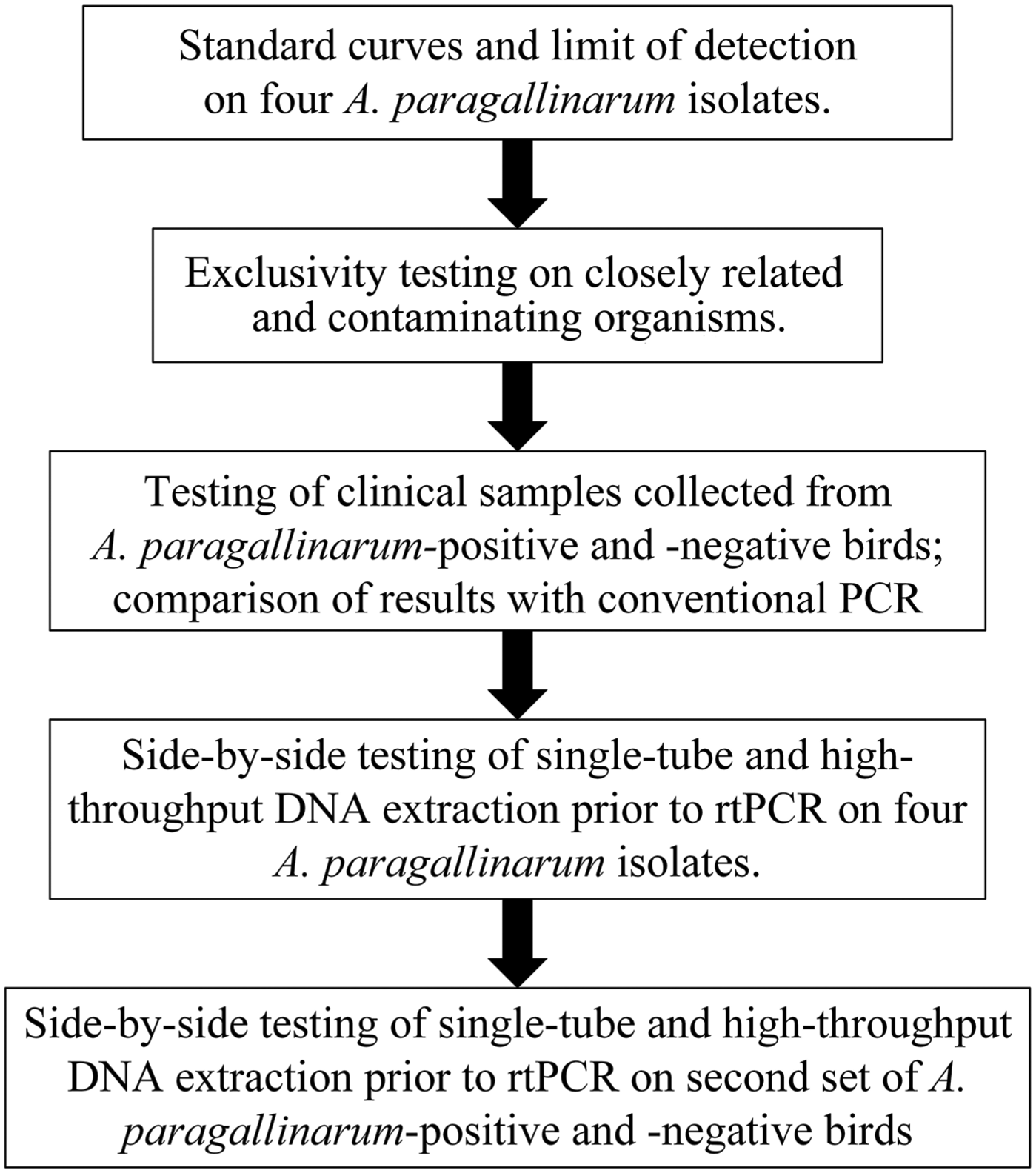
Description of the workflow used to assess a real-time PCR assay for Avibacterium paragallinarum and validate a high-throughput extraction method.
The A. paragallinarum strain and isolates were cultured on chocolate agar (CHOC; Remel, Thermo Fisher Scientific, Waltham, MA) for 48 h at 37°C with 5–7% CO2. Colonies were suspended in sterile saline (pH 7.4, Thermo Fisher Scientific) to a concentration of 1 × 107 cfu/mL and serially diluted in sterile saline to obtain 101–106 cfu/mL suspensions. Colony counts on the 1 × 107 cfu/mL suspension were performed on CHOC for each A. paragallinarum isolate or strain.
Specificity testing included clinical Avibacterium sp. isolates grown on CHOC, and P. vulgaris ATCC 6896 and E. coli ATCC 25922 grown on 5% sheep blood agar (Remel, Thermo Fisher Scientific) for 24 h at 37°C with 5–7% CO2. Colonies were diluted in sterile saline to a concentration of 1 × 105 cfu/mL for PCR testing to mimic the high concentrations of competing organisms seen in clinical disease. Colony counts on the 1 × 105 cfu/mL suspension were performed for each non–A. paragallinarum isolate or strain.
Clinical samples (n = 94) were collected from chickens with (50) or without (44) respiratory disease submitted to the California Animal Health and Food Safety Laboratory System (CAHFS) between June and September 2017. Samples included swabs from 35 birds received as part of an A. paragallinarum challenge study conducted to assess disease in susceptible chickens, and 59 birds submitted for autopsy and further testing. Sampling sites evaluated were choana, infraorbital sinus, and trachea based on diagnostician preference and locations frequently used for A. paragallinarum culture. Swabs were placed into 1-mL vials of sterile phosphate-buffered saline (PBS; Thermo Fisher Scientific) at pH 7.4 and vortexed for 15 s; the swabs were then discarded. Forty-two birds also had swabs submitted for A. paragallinarum culture that were plated onto CHOC as described for the LOD isolates.
DNA was extracted in single-tube format (DNeasy blood and tissue kit; Qiagen, Germantown, MD) based on the published method. 5 Briefly, each 1-mL sample was centrifuged for 10 min at 16,000 × g, and the pellet was resuspended in the kit-supplied ATL buffer. Extraction was performed per manufacturer’s instructions (animal tissue protocol), and final DNA elution was 200 µL of kit-supplied AE buffer.
The extract from each 1 mL of A. paragallinarum 10-fold dilution was PCR tested in triplicate, and cycle threshold (Ct) values for each were plotted to obtain standard curves, slopes, and R2 values for each A. paragallinarum isolate or strain. A 1-mL sample containing 1 × 105 cfu/mL for each of the specificity isolates was extracted and tested with the same protocol.
PCR was performed using a previously published assay 5 targeting the HPG-2 (Haemophilus paragallinarum, the historical name for this bacterium) region of A. paragallinarum. Repeatability experiments were performed between the published protocol and the following alterations to ensure modifications provided consistent A. paragallinarum LOD: a low ROX (5-carboxy-X-rhodamine) probe was utilized (Takyon low Rox probe master mix; Eurogentec, Liege, Belgium); internal amplification control (IAC) DNA (TaqMan exogenous internal positive control DNA; Life Technologies, Thermo Fisher Scientific) was added to each reaction to control for possible PCR inhibition; total reaction volume was 22 µL (20 µL of master mix and 2 µL of template); and PCR was performed on the ABI 7500 (Applied Biosystems, Thermo Fisher Scientific) with cycling parameters as follows: 50°C for 2 min, 95°C for 10 min, 40 cycles of 95°C for 15 s, followed by 60°C for 1 min. Extracted DNA from A. paragallinarum ATCC 29545 was used as the positive PCR control. Positive and negative extraction and amplification controls were included with each run. Clinical samples were considered positive at Ct ≤38.0 based on consistent detection of 10 cfu/mL at this value. Although no known function for this target has been identified to date,8,12 evaluation of the primer sequences using the Basic Local Alignment Search Tool (BLAST) of GenBank identified that the closest non–A. paragallinarum bacterial species match to this primer pair was Haemophilus influenzae at 85% coverage, confirming the specificity of this target.
To assess the clinical robustness of our real-time assay, extracted DNA from a subset of clinical samples including known positive and negative samples (n = 66) was sent for testing using a validated conventional PCR assay performed at an AAVLD-accredited laboratory.
Next, the utility of this assay for high-throughput testing was evaluated. DNA extraction was performed in parallel on additional 10-fold dilutions of A. paragallinarum isolates (ATCC 29545T, CI 1–CI 3) using the single-tube method and the MagMax-96 viral RNA isolation kit (Life Technologies, Thermo Fisher Scientific) for total nucleic acid extraction in a 96-well plate format. Duplicate 1-mL suspensions for each bacterial concentration were centrifuged for 10 min at 16,000 × g, and the pellet was resuspended in 200 µL of PBS. Extraction and PCR were performed as described above for the single-tube format. High-throughput testing used a 50-µL aliquot following the manufacturer’s instructions with DNA eluted in the supplied elution buffer for the 96-well plate format and PCR was optimized using 5 µL of template DNA (added to 20 µL of master mix for a 25-µL total reaction volume). Serial dilutions were tested in triplicate, and Ct values for each were plotted to obtain standard curves, PCR efficiencies, and R2 values for each isolate or strain.
A second set of clinical samples (n = 93) was collected from chickens submitted to CAHFS from December 2017 through April 2018. Two swabs were collected from each bird, placed into 2 mL of PBS, and vortexed for 15 s; the swabs were then discarded. The 2-mL tube was vortexed and divided into two 1-mL aliquots, one for extraction with single-tube and one for extraction in plate format.
Results
Using single-tube extraction for A. paragallinarum PCR with IAC DNA on serial dilutions of A. paragallinarum ATCC 29545T, R2 was 0.982 and PCR efficiency was 110%, and for clinical isolates, R2 was 0.976–0.999 with a PCR efficiency of 89–94% (Fig. 2). The 10-cfu/mL samples produced a mean Ct of 36.9 (range: 35.6–38.0), which resulted in a consistent 10 cfu/mL LOD for this assay across all A. paragallinarum tested. None of the non–A. paragallinarum samples at 105 cfu/mL yielded positive results.
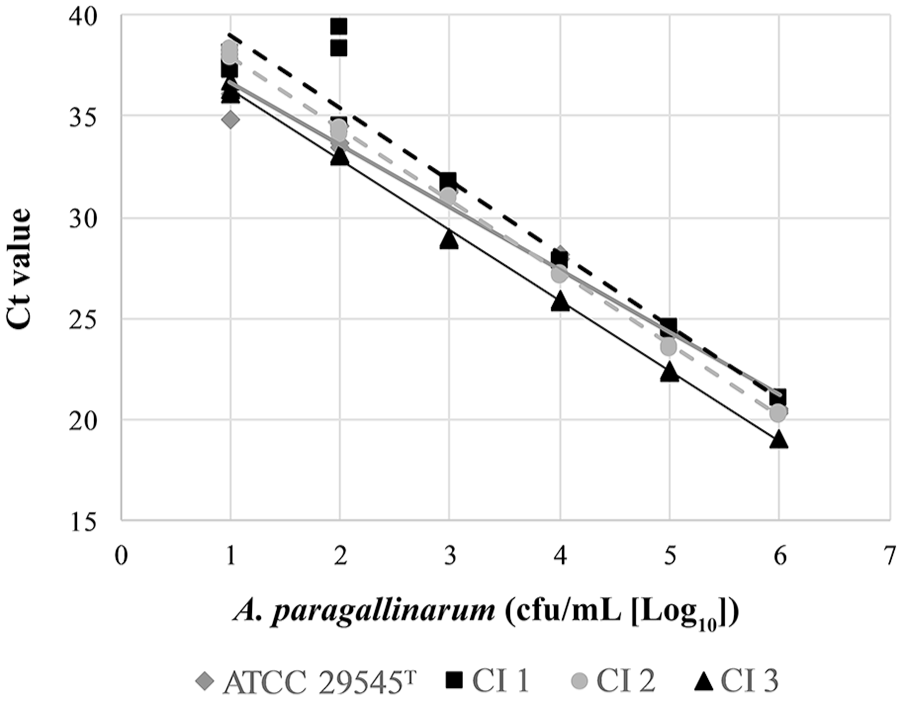
Performance of a real-time PCR assay utilizing the single-tube extraction method on serial dilutions of Avibacterium paragallinarum ATCC 29545T and 3 A. paragallinarum clinical isolates (CI 1–CI 3).
Results from our study birds demonstrated that 18 of 18 challenged chickens were PCR-positive and 17 of 17 unchallenged control chickens were PCR-negative with our assay. Of the 59 birds submitted for autopsy, 18 of 59 tested positive for A. paragallinarum; 14 of 18 of these birds had clinical respiratory disease reported on the submission form and/or noted during autopsy. Only 10 of 36 of the PCR-positive chickens had A. paragallinarum recovered by culture; agar plates from all challenged birds were overgrown with Proteus sp. None of the cultures from PCR-negative birds (6) had A. paragallinarum recovered.
Overall agreement between the rtPCR and conventional PCR testing performed at an AAVLD laboratory was 98.5%, including 35 of 36 rtPCR-positive and 30 of 30 rtPCR-negative samples producing matching results. The discordant sample had a Ct of 37.9 and was negative on the conventional PCR assay.
Side-by-side testing of the 10-fold dilutions for the single-tube and 96-well plate extractions yielded a R2 of 0.981–0.989 (efficiency: 93–111%) for A. paragallinarum ATCC 29545T and 0.957–0.989 (efficiency: 89–94%) for the clinical isolates (Table 1). Side-by-side testing on the second set of clinical samples with the 2 extraction methods produced 95.7% agreement on 37 positive and 52 negative swabs from clinical cases. The remaining 4 samples had Ct values >37.6 from the single-tube (1) or high-throughput (3) methods.
Results of linearity (R2) and efficiency of real-time PCR testing on Avibacterium paragallinarum.
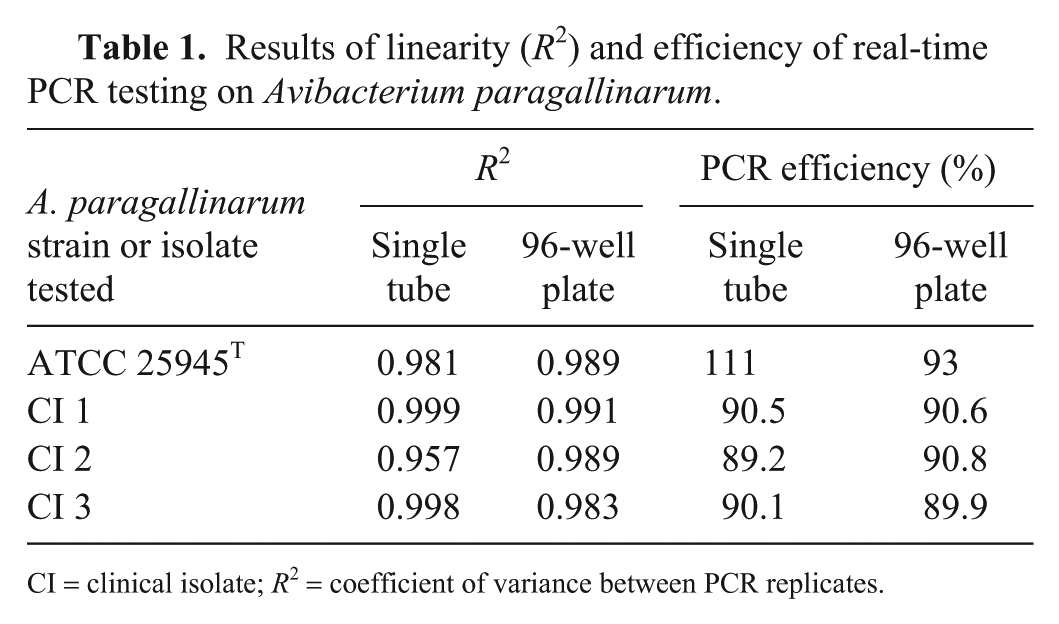
CI = clinical isolate; R2 = coefficient of variance between PCR replicates.
Discussion
Additional infectious agents from PCR-positive chickens in our study included IBV, infectious laryngotracheitis virus (species Gallid alphaherpesvirus 1), Mycoplasma synoviae, M. gallisepticum, Gallibacterium anatis, adenovirus, E. coli, and Aspergillus sp., indicating that the assay was not affected by the presence of other bacterial, viral, or fungal pathogens. G. anatis, a normal resident of the chicken upper respiratory tract, produces mild or no clinical respiratory disease on its own, but enhances the pathogenicity of A. paragallinarum during coinfection and increases severity of IC signs. 11 Ornithobacterium rhinotracheale can produce a range of respiratory signs depending on the virulence factors present in the infecting isolate and presence of other infectious agents. 10 Both G. anatis and O. rhinotracheale produced more severe respiratory disease in chickens coinfected with A. paragallinarum than when inoculated alone.10,11 Clinical coinfections of these pathogens with A. paragallinarum may be missed given difficulties with recovery of this organism when other bacterial agents are present.
Although PCR efficiency was different between the plate extraction method (93%) and the single-tube method (111%) for A. paragallinarum ATCC 29545T, both values are within the acceptable range for a rtPCR assay 13 (Applied Biosystems application note PC1503-PJ9169-CO019879, https://www.thermofisher.com/us/en/home/life-science/pcr/real-time-pcr/real-time-pcr-learning-center/real-time-pcr-basics/real-time-pcr-understanding-ct.html) and were very consistent (89–91%) for the 3 clinical isolates tested. Inclusion of IAC DNA is essential when testing clinical materials that may contain a variety of PCR inhibitors. However, IAC DNA will compete for nucleotides and DNA polymerase in the reaction, which may lower PCR efficiency of the target reaction. 7 We optimized the assay to ensure detection of low numbers of A. paragallinarum (10 cfu/mL) while confirming that IAC Ct values were within a target range to prevent false-negative results as a result of PCR inhibition.
Several limitations are present in our study. The genetic relatedness and serovar identities of isolates used for validation have not been determined. Although tested samples represented commercial layer flocks that did employ A. paragallinarum vaccination programs, as well as broiler and backyard operations that did not, clinical materials were from a restricted geographic area and may not represent the molecular and serologic heterogeneity present in A. paragallinarum.3,9 Work is ongoing to assess performance of this assay on identified serovars and on birds from different areas to ensure that this target detects diverse A. paragallinarum isolates.
Given the difficulties with detection and confirmation by culture, it is reasonable to assume that A. paragallinarum is under-reported and that the asymptomatic birds detected by our assay may reflect variations in virulence. 14 This rtPCR assay provides a high-throughput detection method for chickens with respiratory disease and may provide more accurate and rapid detection needed for the characterization of the epidemiology of IC in chicken populations. It can also serve as a screening method for birds prior to entry into a new flock as well as allow producers to initiate interventions, such as quarantine and vaccination, to limit the spread of disease.
Footnotes
Acknowledgements
We thank Dr. Tomy Joseph from the Animal Health Centre, British Columbia Ministry of Agriculture for providing technical assistance with this study.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
