Abstract
In the present study, the ability of a recently proposed multiplex polymerase chain reaction (mPCR) to determine the serogroups (A, B, and C) of Avibacterium paragallinarum was evaluated. A total of 12 reference strains and 69 field isolates of Av. paragallinarum from Ecuador, Mexico, Panama, and Peru were included in the study. With some exceptions (which were serotyped in the current study), all of the isolates and strains had been previously examined by 2 serotyping schemes (Page and Kume) or were the formal reference strains for the schemes. Three of 6 (50%) reference strains of serogroup A, 2 (100%) of serogroup B, and 1 of 4 (25%) reference strains of serogroup C were correctly serotyped by the mPCR. With the field isolates, the mPCR correctly recognized 16 of the 17 serogroup A isolates, 10 of the 12 serogroup B isolates, and 18 of the 37 serogroup C isolates. Overall, the specificity and sensitivity of the PCR test was as follows: 82.6% and 87.3% (serogroup A), 85.7% and 71.9% (serogroup B), and 46.3% and 100% (serogroup C). The poor performance of the mPCR in terms of recognition of serogroup C isolates (low sensitivity of 46.3%) and the relatively high level of uncertainty about the accuracy of the serogroup A and B results (specificity of 87.3% and 71.9%, respectively) means that the assay cannot be recommended as a replacement for conventional serotyping.
Avibacterium paragallinarum, a member of the family Pasteurellaceae, is the etiological agent of infectious coryza, which is an upper respiratory disease of chickens. 1 Economic loss can be considerable due to reduction in egg production (40%) and increased culling of growing chickens. 4
Isolates of Av. paragallinarum are serotyped by using 2 related schemes. The Page scheme was originally developed with the use of a slide agglutination test to recognize the 3 serovars, A, B, and C. 13 It is now recommended to use a hemagglutination inhibition (HI) test to serotype isolates by the Page scheme. 2 The Kume scheme was based on HI tests that recognized 7 serovars organized into 3 serogroups termed I, II, and III. 10 In further studies, 2 new serovars were recognized into serogroups I and II.3,9 Subsequently, it was proposed to alter the Kume scheme nomenclature on the basis of recognition that the 3 Kume serogroups corresponded to the 3 Page serovars. 9 Thus, the 9 recognized Kume serovars are termed A-1, A-2, A-3, A-4, B-1, C-1, C-2, C-3, and C-4. 9 For the purpose of the current study, the Page types are termed serogroups, in recognition that these Page serogroups match the Kume serogroups. The procedures to serotype Av. paragallinarum isolates by any of these schemes are complicated and require the use of production of hemagglutinating antigens, reference antisera, and the use of chicken erythrocytes fixed with glutaraldehyde. 4
The polymerase chain reaction (PCR) has become a widely used laboratory tool for detection, identification, differentiation, and typing of pathogenic microorganisms in diagnosis of animal diseases. However, there is a need for critical evaluation of any proposed methodology. 14 A PCR for the identification and detection of Av. paragallinarum has been reported7,8 and has been consistently shown to specifically recognize Av. paragallinarum. 11 However, this species-specific PCR does not identify serogroups and/or serovars of this bacterium. A previous study has shown the potential of a multiplex PCR (mPCR) assay to recognize the Page serogroups. 15 This multiplex assay targets the HMTp210 gene, which codes for an outer membrane protein associated with protection against challenge. 17 It is known that a region within the HMTp210 gene (termed region 2), shows, on the basis of a limited data set, sequence diversity between Page serogroups, a diversity that has been exploited in the proposed PCR. 15 In the current study, this proposed PCR alternative to conventional serotyping has been evaluated with the full set of Page and Kume reference strains as well a number of diverse Av. paragallinarum field isolates. In this evaluation, the 2 inter-related conventional serotyping schemes have been accepted as the “gold standard.”
A total of 11 reference strains, representing the 3 Page serogroups and the 9 Kume serovars of Av. paragallinarum were used in the study. Also, strain NCTC (National Collection of Type Cultures) 11296T (CCUG [Culture Collection, University of Göteborg, Sweden]12835T; ATCC [American Type Culture Collection] 29545T) of Av. paragallinarum was used. Furthermore, 69 field isolates of Av. paragallinarum were included in the current study (Table 1). The isolates were from collections held by the Center for Advanced Investigations and Studies on Animal Health (Mexico) and FARVET (Peru) laboratories, and were obtained from infectious coryza outbreaks on different farms with no epidemiological connections. Of these 69 field isolates, a total of 65 (26 from Ecuador, 37 from Mexico, and 2 from Panama) had been previously serotyped by the Page and Kume schemes.5,6,12 Strain NCTC 11296T and the Peruvian isolates were serotyped, in the current study, by the Page and Kume schemes using methods reported elsewhere.3,16
Origin, Page serogroup, Kume serovar, and multiplex polymerase chain reaction (mPCR) results for the Avibacterium paragallinarum strains and isolates included in the current study.*
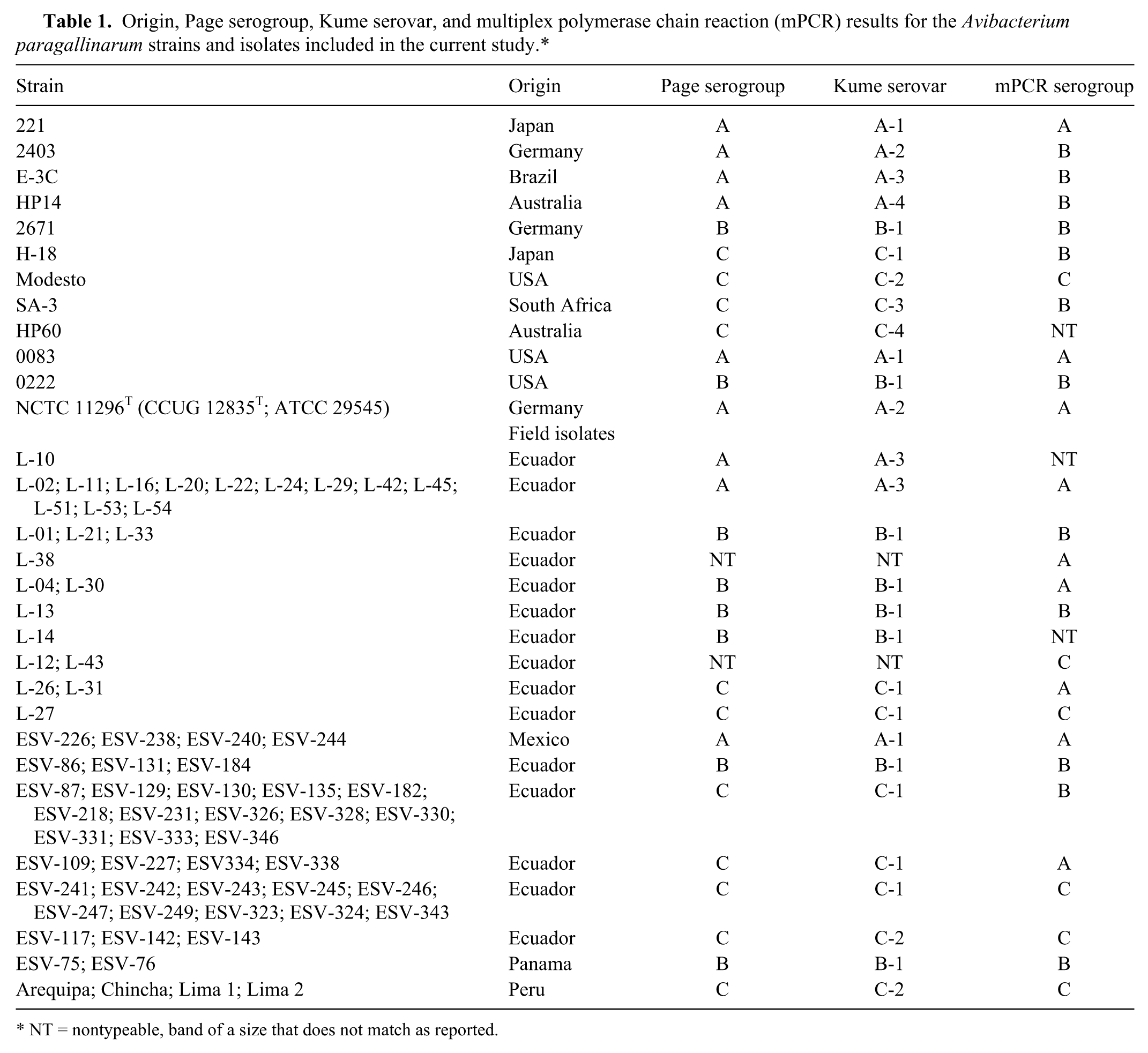
NT = nontypeable, band of a size that does not match as reported.
Bacteria were maintained and propagated in brain-heart infusion broth and agar, both supplemented with 1% (w/v) sodium chloride, 0.0025% (w/v) reduced nicotinamide adenine dinucleotide (NADH), and 1% (v/v) filter-sterilized, heat-inactivated horse serum. 12 All reference strains and field isolates included in the study were examined by a serotyping mPCR as reported. 15 Briefly, the genomic DNA was extracted by using a commercial kit, a according to the manufacturer’s instructions. The PCR reactions were performed in a total volume of 25 μl containing 2.5 μl of 10× PCR buffer included in the kit, b 0.2 mM of each deoxyribonucleotide triphosphate, 0.2 μM of each Page serovar–specific primer: ABC forward 5′-GGCTCACAGCTTTATGCAACGAA-3′; A reverse 5′-CGCGGGATTGTTGATTTTGTT-3′; B reverse 5′-GGTGAATTTCACCACACCAC-3′; and C reverse 5′-TAATTTTCTTATTCCCAGCATCAATACCAT-3′. 15 A total of 1.25 units of Taq DNA polymerase b and 2 μl of extracted DNA were used. The amplification steps were as reported previously: 98°C for 1 min; 30 cycles of 98°C for 10 sec, 56°C for 10 sec and 72°C for 2 min, and a final step at 72°C for 7 min. 15 The PCR products were subjected to agarose gel electrophoresis using 2% agarose, visualized, and photographed under ultraviolet light. Multiplex PCR serotyping of all isolates and reference strains included in the study were performed in both testing laboratories (Mexico and Peru).
The results of the multiplex serotyping were the same for all reference strains and field isolates in both testing laboratories (Table 1). Reference strains 221, 0083, 0222, and Modesto were correctly identified as serogroup A, A, B, and C, respectively, as previously reported. 15 Reference strain 2671 (serogroup B) was identified as serogroup B. However, reference strains 2403, E-3C, and HP14 of serogroup A were misidentified as serogroup B. Similarly, reference strains H-18 and SA-3 of serogroup C were misidentified as B. Reference strain HP60 (serovar C-4) was nontypeable (Fig. 1). Strain NCTC 11296T of Av. paragallinarum from Germany was serotyped for the first time and was recognized as serovar A-2, with the PCR correctly assigning the strain to serogroup A. Three of the field isolates that were nontypeable by the HI test were identified as serogroup A or serogroup C. Of the 17 field isolates of serogroup A, 16 were correctly assigned to serogroup A by the mPCR. Of the 12 serogroup B field isolates, 10 were correctly assigned to serogroup B. Of the 37 serogroup C field isolates, only 17 isolates were correctly recognized as serogroup C by the mPCR.
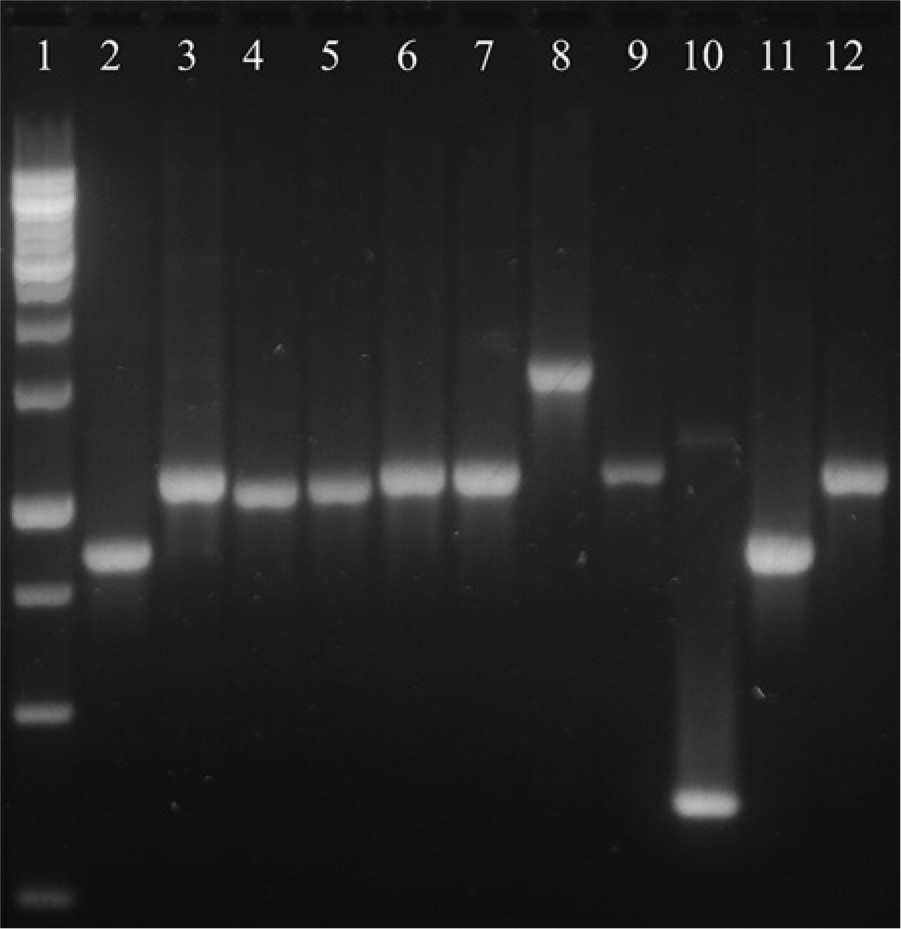
Agarose gel electrophoresis of multiplex polymerase chain reaction products from 11 Avibacterium paragallinarum reference strains included in the current study. Lane 1: molecular weight marker; lane 2: strain 221; lane 3: strain 2403; lane 4: strain E3-C; lane 5: HP14; lane 6: strain 2671; lane 7: H-18; lane 8: strain Modesto; lane 9: strain SA-3; lane 10: strain HP60; lane 11: strain 0083; lane 12: strain 0222.
To fully evaluate the performance of the mPCR, the complete results for all the reference strains and the field isolates were assembled and the sensitivity, specificity, positive predictive value (PPV), and negative predictive value (NPV) were calculated (Table 2). Overall, the sensitivity of the PCR assay ranged from 46.3% (serogroup C) to a high of 85.7% (serogroup B). In terms of specificity, the assays varied from 71.9% (serogroup B) to 100% (serogroup C). The PPV varied from 40% (serogroup B) to 100% (serogroup C) while the NPV varied from 62.7% (serogroup A) to 95.8% (serogroup B). Based on these results, field isolates that were nontypeable by the HI test may be misidentified by this mPCR.
Summarized results of the multiplex polymerase chain reaction (mPCR) assay as compared with conventional serotyping.*
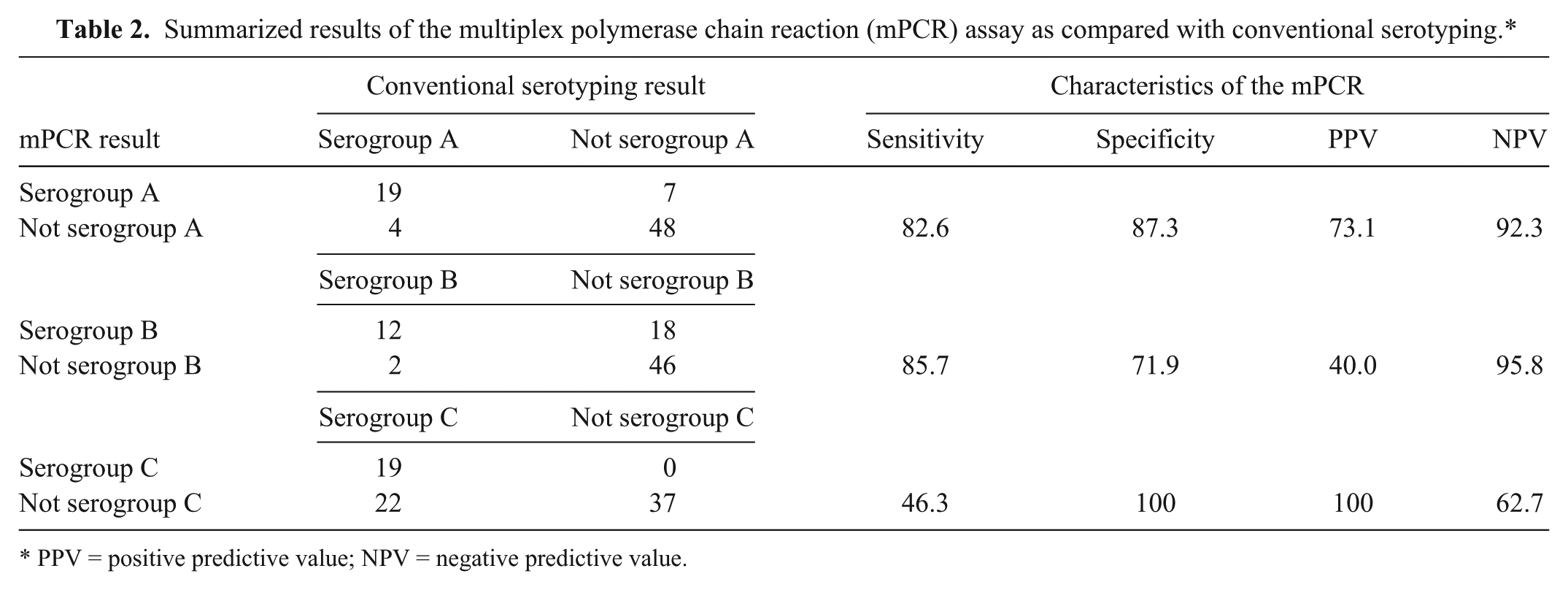

PPV = positive predictive value; NPV = negative predictive value.
The present study has confirmed the results of the initial evaluation of this mPCR, 15 a study that was performed on a limited set of the reference strains and which indicated the possible potential of this assay. However, when evaluated on a more complete set of reference strains and a diverse set of field isolates, the limitations of the mPCR are clear. The results of the current study clearly show that the apparent link between sequence diversity in region 2 and Page serogroups only holds true for the small set of strains examined in the initial study that proposed the methodology. 15 Indeed the evidence of greater diversity in the region has been shown in an earlier study using recombinant proteins based on region 2. 17 This earlier study found that the HI antibodies elicited by a recombinant protein based on this region could be strain specific and not react with other strains of the same serovar. 17 It is clear that there is no simple, direct connection between the regions targeted by the mPCR and Page serogroups. A more extensive study looking at the sequence of region 2 in a much larger and more diverse set of strains and isolates than that used in the original study 15 may be able to identify alternative PCR assays that show a much closer alignment with Page serogroups.
The performance of the mPCR was poor with both the definitive internationally recognized reference strains as well as the field isolates. This poor level of performance with standard strains and field isolates strongly supports the conclusion that it is the PCR results that are wrong and not the serotyping results. If the PCR had performed well with the reference strains (which by definition have to be correctly serotyped) and poorly with the field isolates, this would be an argument that the PCR was simply performing better than serotyping when examining field isolates. However, the poor performance of the PCR was found with both reference strains and field isolates.
The PCR did assign some isolates that were nontypeable by the conventional assay to a serovar. However, given the overall poor performance of the PCR assay, it is not possible to conclude if the PCR results for these nontypeable isolates are actually correct.
The original study on the mPCR also described an alternative technology—an assay in which a PCR using a different set of primers is followed up by a restriction fragment length polymorphism analysis to generate serogroup-specific patterns. 15 This second technology has also only been evaluated with a very limited set of strains. Hence, this alternative technology also needs a thorough evaluation as it may also have the same limitations as have been found for the mPCR in the current study. The additional step of restriction digestion is a complication that many diagnostic laboratories may regard as too challenging for routine use. In conclusion, the mPCR cannot be recommended as an alternative to conventional serotyping of Av. paragallinarum.
Footnotes
Acknowledgements
The authors gratefully acknowledge Dr. Arturo Cabrera, LAVETEC CIA LTDA., Ecuador, for providing the Ecuadorian isolates of Av. paragallinarum.
a.
DNeasy blood & tissue kit, Qiagen, Hilden, Germany.
b.
Taq DNA polymerase recombinant, Invitrogen, Sao Paulo, Brazil.
Declaration of conflict interests
The author(s) declare no potential conflicts of interest with respect the research, authorship, and/or publication of this article.
Funding
This work was supported by Universidad Autónoma del Estado de México, project 3102/2011. Vladimir Morales-Erasto held a scholarship from the Consejo Nacional de Ciencia y Tecnología (CONACYT), Mexico.
