Abstract
Various PCR-based assays for rabbit viruses have gradually replaced traditional virologic assays, such as virus isolation, because they offer high-throughput analysis, better test sensitivity and specificity, and allow vaccine and wild-type virus strains to be fully typed and differentiated. In addition, PCR is irreplaceable in the detection of uncultivable or fastidious rabbit pathogens or those occurring in low quantity in a tested sample. We provide herein an overview of the current state of the art in the molecular detection of lagomorph viral pathogens along with details of their targeted gene or nucleic acid sequence and recommendations for their application. Apart from the nucleic acids–based methods used for identification and comprehensive typing of rabbit viruses, novel methods such as microarray, next-generation sequencing, and mass spectrometry (MALDI-TOF MS) could also be employed given that they offer greater throughput in sample screening for viral pathogens. Molecular methods should be provided with an appropriate set of controls, including an internal amplification control, to confirm the validity of the results obtained.
Introduction
The rabbit breeding industry has experienced a considerable increase in rabbit meat production, despite the frequent occurrence of infectious diseases in this animal species.16,24,45,48,53,85 Rabbits can suffer from many viral, bacterial, and parasitic diseases (Agnoletti F. Update on rabbit enteric diseases: despite improved diagnostic capacity, where does disease control and prevention stand? Proc 10th World Rabbit Congress; Sep 2012; Sharm El-Sheikh, Egypt. Available from: https://world-rabbit-science.com/WRSA-Proceedings/Congress-2012-Egypt/Papers/07-Pathology/P00-Agnoletti.pdf)5,69,70; however, viral infections seem to have the highest impact on their health (Lavazza A, Capucci L. Viral infection of rabbits. Proc 9thWorld Rabbit Congress; Jun 2008; Verona, Italy. Available from: https://world-rabbit-science.com/WRSA-Proceedings/Congress-2008-Verona/Papers/P0-LavazzaCapucci.pdf).19,55 Viral diseases of rabbits are characterized by sudden onset and a fatal course. Widespread use of vaccines against myxomatosis and rabbit hemorrhagic disease (RHD) has significantly reduced mortality on rabbit farms. Although outbreaks of rabbit viral diseases are only occasionally described in the scientific literature, viral infections leading to animal losses occur worldwide, especially on small-scale rabbit farms. If such infections appear, the use of sensitive and rapid tests is essential for disease recognition and introduction of appropriate control measures limiting virus spread in the herd.
Among several rabbit and hare virus types, the most important are RHD virus (RHDV), European brown hare syndrome virus (EBHSV), and myxoma virus (MYXV). RHDV causes one of the most common fatal diseases of domestic and wild European rabbits.88,117 According to the proposed classification in 2017, all RHDV strains have been allocated to the Lagovirus europaeus genogroup I (GI) within the Caliciviridae family. 68 This genogroup encompasses several genotypes (GI.1–4) represented by classic RHDV and its RHDVa variants (GI.1), 72 nonpathogenic rabbit caliciviruses (RCVs; GI.3–GI.4), 15 and novel, antigenically distinct RHDV-2/b (GI.2) virus strains. 63 Moreover, RHDV classic strains that previously were classified in the G1–G6 genogroups 65 are now placed within the GI.1a–GI.1d clusters of virus variants. Furthermore, EBHSV strains and other EBHSV-related viruses are currently located in Lagovirus genogroup II (GII). 68 RHDV and RHDV-2 have also been detected in hares, suggesting possible cross-species virus transmission.13,75,109 EBHSV is morphologically similar to, and both antigenically and genetically related to, RHDV. Second to RHDV in destructive health impact, MYXV causes myxomatosis, a highly infectious systemic disease of European rabbits. 33 Two clinical forms of this disease have been described, namely nodular (systemic) and atypical (amyxomatous),78,103 with mortality reaching 100% among susceptible animals.9,54
Molecular methods have gradually replaced traditional antigen-detection assays, such as virus isolation, given that they offer better specificity and sensitivity with ~100% accuracy in detection of viral genetic material. They are irreplaceable in the detection of uncultivable or fastidious rabbit pathogens or those occurring in low quantity in a tested sample. In addition, they allow early detection of infection, even before signs of disease occur. Despite many advantages over the classical virus-detection methods, they also have limitations. Namely, molecular assays do not allow identification of unknown pathogens and are prone to error given the possibility of sample contamination with target DNA/RNA from another sample or the PCR product of a previous amplification. 73 Therefore their application for routine testing requires highly qualified personnel and a suitably equipped laboratory. Our aim in this review is to summarize existing information on the molecular methods used in the detection of rabbit and hare viruses and the diseases that they cause.
Detection of lagomorph caliciviruses
PCR
Although several methods such as ELISA, hemagglutination (HA), and electron microscopy are commonly employed for RHDV and/or EBHSV detection, such methods are gradually being replaced by modern PCR-based amplification techniques (Tables 1, 2). PCR primers are usually designed in conserved regions of the VP60 capsid protein gene of the known rabbit caliciviruses encompassing RHDV,41,44,98 RHDVa,2,14,36 RHDV-2, 23 and EBHSV, as well as RCV strains.8,64,105 Reverse-transcription PCR (RT-PCR) has been used successfully for the detection of RHDV in a vast range of tissues, blood, and nasal secretions of RHDV-infected rabbits. 118 The method was characterized by high specificity and 10,000 times greater sensitivity than HA, and was able to detect both HA-negative and -positive RHDV strains. Another one-step RT-PCR method employing a universal primer pair targeting the VP60 gene of the RHDV and RHDVa strains has been described (Pugliese N, et al. A RT-PCR based strategy to identify rabbit hemorrhagic disease virus. Proc 10th World Rabbit Congress; Sep 2012; Sharm El-Sheikh, Egypt. Available from: https://world-rabbit-science.com/WRSA-Proceedings/Congress-2012-Egypt/Papers/07-Pathology/P-Pugliese.pdf). This method could widen the efficiency of the existing molecular techniques given that it permits simultaneous detection of RHDV variants and differentiation between RHDV and EBHSV strains. The protocol could be easily adapted to the real-time RT-PCR (RT-rtPCR) format given the generated amplicon size and melting temperatures of the primers required for this method. Recently, a SYBR Green RT-qPCR based on the viral VP60 gene was developed for detection of all GI rabbit lagoviruses circulating in Australia. 42 It was used solely for initial screening of animals for RHDV infections; virus detection was performed using the more sensitive multiplex RT-PCR. The assay was able to detect ≥10 capsid copies per reaction. In addition to the RHDV-specific primers, internal control primers targeting the rabbit 12S mitochondrial rRNA gene were used to monitor the isolation of viral RNA. 84
Characteristics of PCR-based methods used for detection of European brown hare syndrome virus (EBHSV) and differentiation of rabbit hemorrhagic disease virus (RHDV) from EBHSV strains.
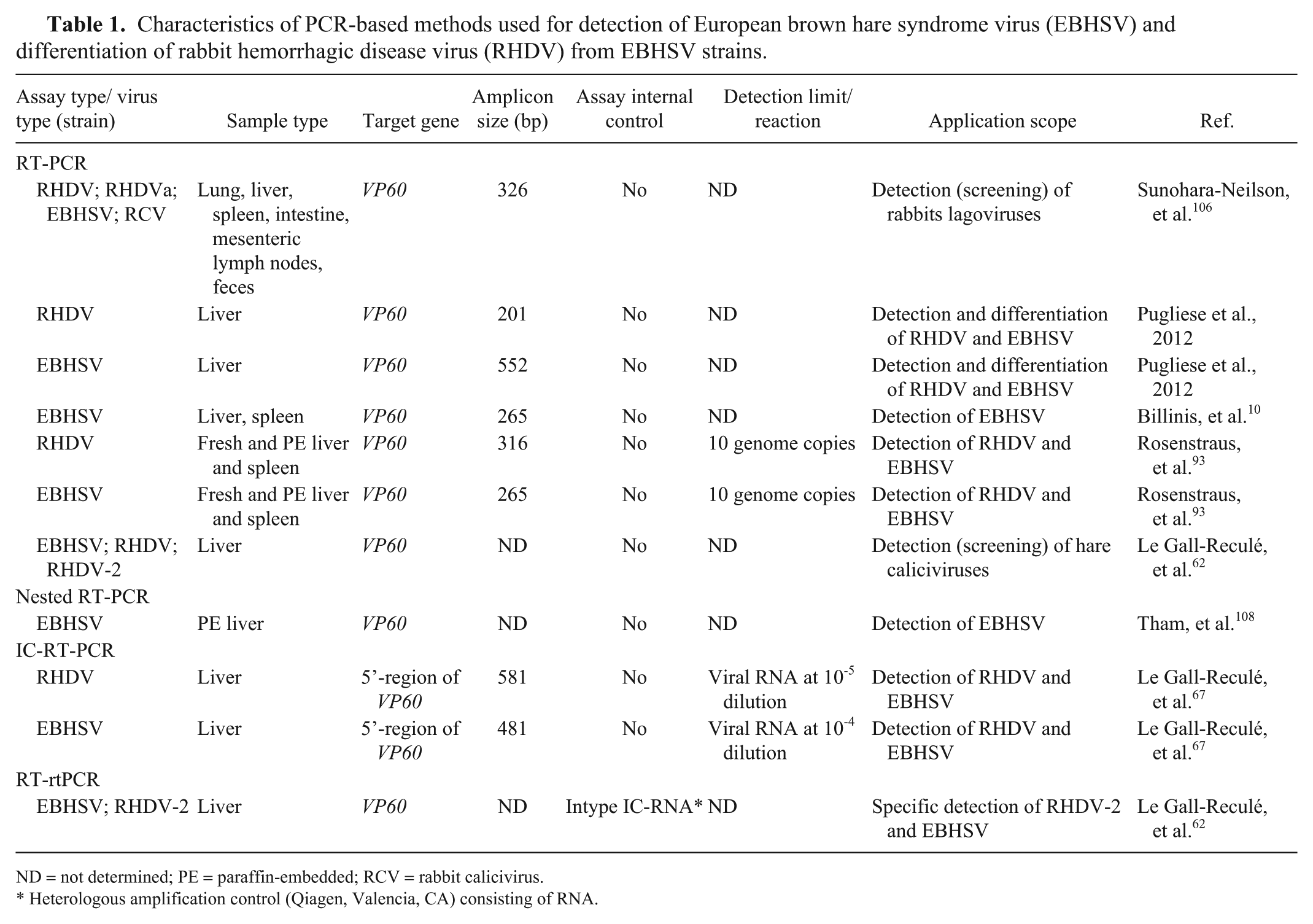
ND = not determined; PE = paraffin-embedded; RCV = rabbit calicivirus.
Heterologous amplification control (Qiagen, Valencia, CA) consisting of RNA.
Characteristics of PCR-based methods used for detection of classic rabbit hemorrhagic disease virus (RHDV) and its variants.
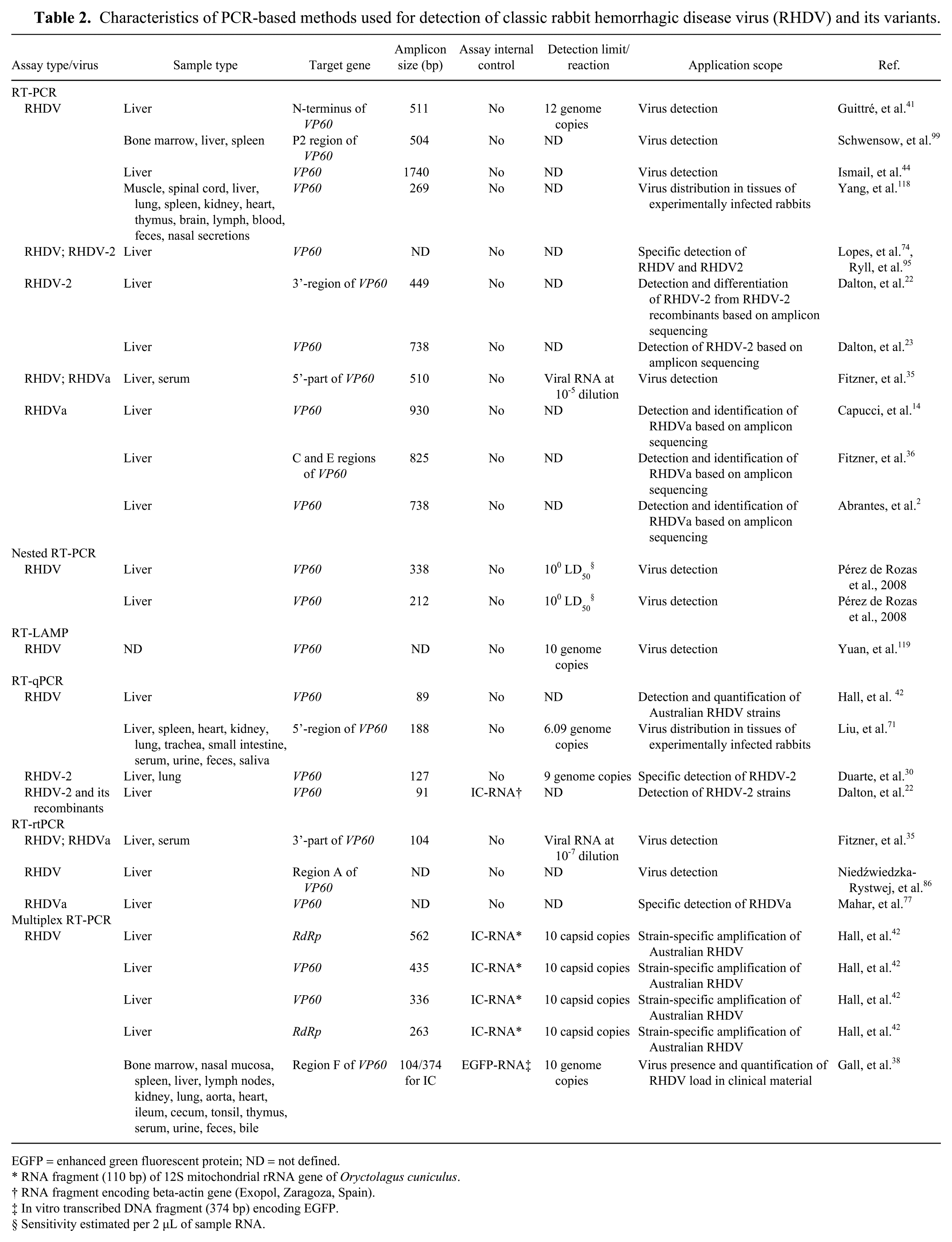
EGFP = enhanced green fluorescent protein; ND = not defined.
RNA fragment (110 bp) of 12S mitochondrial rRNA gene of Oryctolagus cuniculus.
RNA fragment encoding beta-actin gene (Exopol, Zaragoza, Spain).
In vitro transcribed DNA fragment (374 bp) encoding EGFP.
Sensitivity estimated per 2 µL of sample RNA.
In addition to RT-PCR for RHDV detection, conventional and nested RT-PCRs (nRT-PCR) employing EBHSV VP60 gene primers were used in epidemiologic studies of viral prevalences in European brown hares in Greece and free-living mountain hares in Finland.10,107 Primers were designed to amplify EBHSV exclusively and to detect its genetically distant variants. Cross-reactivity of the EBHSV primers with RHDV strains has not been shown, in spite of RHDV and EBHSV being closely related caliciviruses. The assay was capable of detecting as few as 10 genome copies per reaction. It was optimized to detect viral RNA not only in fresh clinical material but also in formalin-fixed, paraffin-embedded tissue specimens. 92 Nested RT-PCR appeared to be a useful tool and able to confirm a histologic diagnosis of EBHSV infection even if autolyzed tissues were used for testing. A second round of amplification with internal primers increased the sensitivity of the assay and reduced background caused by nonspecific amplification.92,107 The VP60 gene was also chosen in another study 62 as the target for PCR assays (i.e., RT-PCR) used as a general screening tool for rabbit and hare caliciviruses, and as the target for a RT-rtPCR method for the detection of infections caused by EBHSV and RHDV/RHDV-2 in hares. In addition to virus-specific protocols, the VP60 gene was also used for the development of a universal lagovirus RT-PCR with primers spanning a conserved region within the RHDV, EBHSV, and RCV genomes. 105 Contrary to these encouraging results, limited usefulness has been reported for some primers annealing to the N-terminal part of VP60 and the C-terminus of RNA-dependent RNA polymerase (RdRp) RHDV genes when used for molecular detection of RHDV infections in farmed rabbits. 34 The lack of amplification can to some extent be explained by primer mismatches to target sequences within the viral genome; nucleotide mutations in the amplified fragments were revealed. It is also noteworthy that RHDV strains are gradually replaced by RHDV-2 and their recombinants.22,74,94,100 As a result, the tests used in molecular detection of RHDV infections may require regular review, and updates of primer sequences and negative results should be verified by another test. 34
Nested RT-PCR
In comparison to a single-round PCR assay, nRT-PCR is considered a more sensitive and specific molecular method for use in the detection of rabbit viruses (Pérez de Rozas AM, et al. Standardization of nested-PCR for the detection of Pasteurella multocida, Staphylococcus aureus, myxomatosis virus, and rabbit hemorrhagic disease virus. Proc 9th World Rabbit Congress; Jun 2008; Verona, Italy. Available from: https://world-rabbit-science.com/WRSA-Proceedings/Congress-2008-Verona/Papers/P-PerezdeRozas1.pdf); application of 2 primer sets greatly improved the method’s sensitivity in the detection of MYXV and RHDV in biological material containing a low virus load.40,92 Furthermore, nRT-PCR was able to reduce the effect of reaction inhibition and eliminate nonspecific amplification when rabbit and hare tissues were tested for RHDV and EBHSV. 92 Although nPCR increases the specificity of DNA amplification by reducing background amplification of nonspecific DNA, it requires more detailed information on the target sequence as well as lengthening the analysis compared to a single-round PCR. In addition, this technique is susceptible to carryover contamination with PCR products from a preceding amplification. Moreover, gel electrophoresis of PCR amplicons with their subsequent visualization under ultraviolet light is required.
RT-LAMP
One-step reverse-transcription loop-mediated isothermal amplification (RT-LAMP) is a rapid and cost-effective method not requiring the use of specialized equipment. This method has been employed for in vitro amplification of RHDV RNA. 119 A set of 4 specific primers was used under isothermal conditions to amplify the RHDV VP60 gene fragment. The RT-LAMP products were detected by DNA electrophoresis or SYBR Green I staining. Method sensitivity was established at 10 copies of viral RNA per reaction, which was comparable to the sensitivity of RT-rtPCR, and 100-fold higher than conventional RT-PCR. However, the application of the LAMP assay was only confirmed for detection of Chinese RHDV isolates and not for RHDVa strain variants. Therefore, further optimization of primer sequences targeting more conserved regions of the RHDV genome is required to make a universal virus detection tool.
Immunocapture RT-PCR
To simplify the time-consuming procedure of sample preparation for molecular analyses, an immunocapture RT-PCR (IC-RT-PCR) has been developed for the detection of RHDV and EBHSV in liver specimens of domestic and wild rabbits and hares.66,67 In this method, viral particles present in the sample are captured by specific antibodies immobilized on a microtitration plate. After enzymatic disruption of virus-antibody complexes, viral RNA is released and subjected to a reverse transcription step coupled with PCR. Compared to ELISA, this method showed at least 50 times higher sensitivity (Chrobocińska M. Charakterystyka fenotypowa i molekularna krajowych szczepów wirusów krwotocznej choroby zajęcy (EBHSV) i krwotocznej choroby królików (RHDV) [Phenotypic and molecular characterization of Polish strains of the European brown hare syndrome virus (EBHSV) and rabbit hemorrhagic disease virus (RHDV)] [habilitation dissertation]. Puławy, Poland: National Veterinary Research Institute, 2007. Summary available from: http://www.piwet.pulawy.pl/piwet7/newslet/2009-01/prace_dr_hab/Praca_habil_2.pdf).
RT-rtPCR
Conventional PCR assays are steadily being replaced by rtPCR assays. This technique is known to be more efficient than conventional PCR in the detection and quantification of small numbers of viral particles.59,111 It has been employed successfully for detection of RHDV strains circulating in rabbit populations in various European countries1,16,22,30,35,38,86,87 and Australia 77 or for studying virus distribution in tissues of experimentally infected rabbits. 71 Broadly reactive primers within the conserved 3’-region of the VP60 RHDV gene were designed and used in the TaqMan 35 and SYBR Green71,86 RT-rtPCR methods. When compared with conventional RT-PCR, RT-rtPCR was 100 times more sensitive, 35 with its detection level established at 6.09 genome copies per reaction. 71
The emergence of new RHDV-2 strains in rabbits initiated research toward the development of strain-specific TaqMan RT-rtPCR methods.16,22,30 The primers and probes targeted a highly conserved fragment of the RHDV-2 VP60 gene, with only a few nucleotide mismatches compared to RHDV genes as well as to non- and low-pathogenic RCVs. Although these methods are able to detect RHDV-2 strains, full sequence analysis of the complete VP60 gene is necessary when differentiation between vaccine and field virus strains is needed.16,30
An internally controlled multiplex one-step quantitative RT-PCR (RT-qPCR) has been developed with primers and a TaqMan probe designed in the conserved F region of the virus capsid protein gene, which enabled detection of RHDV RNA in sera, tissues, and excretions of experimentally infected rabbits. 38 This assay allowed simultaneous amplification of viral RNA and an in vitro transcribed DNA fragment encoding enhanced green fluorescent protein (EGFP-RNA), which served as the internal amplification control (IAC). The IAC was detected using an independent primer–probe system. It was designed to control the efficiency of RNA isolation and the RT-PCR detection step. The application of an IAC-specific, primer–probe mix in the same reaction tube independently from the target RNA present did not affect the assay’s sensitivity. The internally controlled duplex one-step RT-rtPCR assay has also been elaborated to study RCV-A1 distribution, and determination of the virus load in various rabbit organs. 105 The IAC consisting of EGFP-RNA efficiently monitored the reaction inhibition during molecular detection of viral RNA extracted from rabbit tissues. Molecular methods should be equipped with an appropriate set of controls to confirm the validity of the results obtained. An IAC that is simultaneously co-amplified with target viral DNA or RNA using the same (competitive IAC) or different (noncompetitive IAC) primer pairs seems to be the most useful.27,91,93 The application of IAC is recommended in microbiologic testing, and in some research areas, its use is obligatory during molecular analysis of the sample. Another approach is the use of an endogenous amplification control consisting of the beta-actin gene, which has been incorporated into the RT-rtPCR designed for RHDVb detection. 22 The endogenous amplification control confirmed the correct extraction and amplification performance of the assay.
Hybridization techniques
In situ hybridization (ISH) is another molecular method for early detection of RHDV infection. Although the method is highly sensitive and specific, its use in veterinary laboratories is not common given the high costs of analysis and complicated methodology compared to other virologic tests. ISH has been successfully employed in studies on RHD pathogenesis. 39 Virus distribution was analyzed in organs of infected animals, indicating the liver as a major site of viral replication. RHDV was detected in rabbit tissues using a digoxigenin-labeled probe hybridizing to sequences within a structural protein gene of the RHDV genome. To increase sensitivity and specificity of PCR-based methods for RHDV detection in rabbit tissues, southern blot hybridization has been applied (Chrobocińska M, 2007).15,41,108 This method combines the transfer of electrophoretically separated fragments of viral DNA to a filter membrane and their subsequent detection by specific digoxigenin- or radio-labeled probes.15,41,108 A combination of several virus-detection techniques (ELISA, PCR, and hybridization) in one PCR-ELISA method for detection of RHDV in tissues, blood, and excretions of infected rabbits was proposed. 89 The oligonucleotide primers were derived from the conserved part of the capsid region of the RHDV genome, although the primers were also effective in amplification of some EBHSV isolates. This alternative procedure of analysis of the digoxigenin-labeled PCR products gave a yield several times higher than traditional gel electrophoresis.
Detection of myxoma virus
A diagnosis of rabbit myxomatosis is usually based on clinical signs; however, identification of the amyxomatous form of the disease requires laboratory testing. For years, the diagnosis of myxomatosis has relied on isolation of the virus in cell culture, electron microscopy, and immunologic methods such as agar gel immunodiffusion test and indirect fluorescent antibody test. 113 Such methods are gradually being replaced by molecular techniques (Table 3).
Characteristics of PCR-based methods used for detection of myxoma virus.
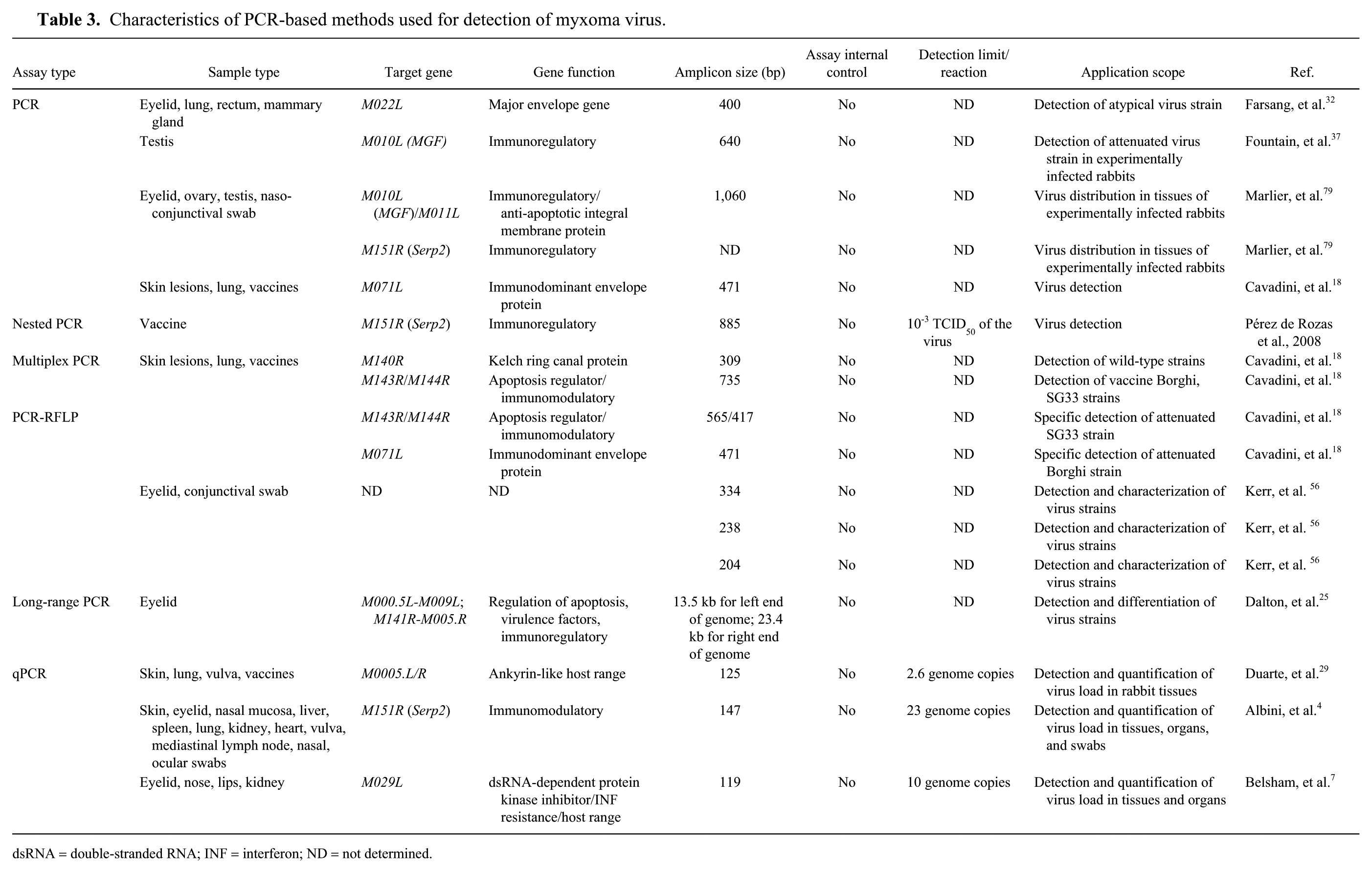
dsRNA = double-stranded RNA; INF = interferon; ND = not determined.
PCR assays
Given the appearance of an atypical form of myxomatosis associated with respiratory distress and only limited or absent skin changes, recognition of the disease based only on clinical signs is difficult. Therefore, PCR-based methods were designed and subsequently used for virus detection in various tissues.29,32,113 PCR also showed its usefulness in clinical studies of virus distribution in tissues of infected animals.37,79 Typically the highly conserved regions of the MYXV genome are chosen for primer selection during elaboration of PCR-based assays, thus allowing detection of circulating virus strains. For example, the M071L, M140R, and M142R/M144R MYXV genes were selected to design specific primers for PCR and PCR–restriction fragment length polymorphism (PCR-RFLP) protocols able to discriminate between vaccinated and naturally infected animals rapidly, and to detect mixed infections caused by wild-type and vaccine MYXV strains. 18 The sequence differences in the M143R, M144R, and M071L genes between the attenuated vaccine and reference MYXV strains result in different DNA restriction profiles.
Enzymatic digestion of the entire virus genome or its fragments was also employed for identification of differences and similarities in genomic DNA between field, standard laboratory strains (SLS), or Lausanne strains.56,57,60,90,96 However, because of the large size of the MYXV genome (161.8 kb), these analyses are laborious and suffer from a lack of appropriate controls, which limits their routine application.
A long-range PCR-RFLP method directed toward amplification of genomic MYXV DNA from the left and right terminal inverted repeat regions (TIRs) with subsequent RFLP differentiation of virus strains has been proposed. 25 Two sets of primers covering the entire TIRs and their flanking sequences (M009L gene and genome regions from M141R to M156R) were designed. The long-range PCR-RFLP proved to be a simple and highly efficient method for identifying mutations in the amplified genome fragments of MYXV strains, with potential application in phylogenetic studies of variable regions within the virus genome. This method can simplify detection and characterization of virus strains because most nucleotide changes in the MYXV genome have been recognized in the TIRs. 96
rtPCR
In addition to qualitative data, quantitative determination of viral copy number in the tested sample can be performed by rtPCR. This technique offers better sensitivity than other molecular methods.31,76 It has been used for detection of MYXV DNA in fresh and paraffin-embedded rabbit tissues as well as in nasal, ocular, and genital swabs collected from pet, farmed, and wild rabbits. 4 Primers and probe were designed to amplify a 147-bp fragment of the serpin (Serp2 gene. The test was validated, and its analytical specificity and sensitivity were confirmed. Analytical sensitivity was determined using an internal Serp2 oligo standard yielding the detection limit of 23 genome copies of MYXV DNA per reaction. 4
Another TaqMan qPCR targeting a M029L gene of MYXV was elaborated and subsequently used for detection and confirmation of suspected cases of myxomatosis. 7 In this assay, to eliminate amplicon carryover, an additional reaction step involving sample incubation with uracil-N-glycosylase (UNG) was included. UNG excises uracil residues from PCR amplicons from previous amplifications containing incorporated dUTP. The assay efficiently detected MYXV DNA but did not produce amplification signals for other tested poxviruses. It could detect as few as 10 copies of the virus sequence per reaction.
A highly sensitive qPCR with primers complementary to the nucleotide sequences within the M000.5L/R virus gene was also established. 29 Interestingly, this gene has 2 copies in the MYXV genome, in the right and left TIR, respectively. Therefore, in comparison to other PCR protocols targeting virus genes present in a single copy, its sensitivity was significantly increased and reached 2.6 genome copies of MYXV DNA per reaction. In addition, M000.5L/R is a unique gene in the Leporipoxvirus genome, 112 and its use and amplification also increases the specificity of this PCR-based assay. 29
Molecular detection of other rabbit viruses
Rotavirus
Epizootic rabbit enteropathy is in the group of economically significant rabbit disorders. It is caused by bacterial and viral pathogens, with lapine rotavirus (LRV) mainly detected in outbreaks of rabbit enteritis.5,19,26,70,97 Virus detection can be performed by the use of ELISA, electron microscopy, and RT-PCR (Agnoletti F, 2012; Lavazza A, Capucci L, 2008). A molecular survey of rotavirus prevalence in farmed rabbits in Canada was conducted; 116 virus occurrence was estimated based on detection of the LRV VP6 gene. Other RT-PCR methods targeting VP7 or VP4 genes have been used for LRV typing (Lavazza A, et al. Rotavirus in diarrheic rabbits: prevalence and characterization of strains in Italian farms. Proc 9th World Rabbit Congress; Jun 2008; Verona, Italy. Available from: https://world-rabbit-science.com/WRSA-Proceedings/Congress-2008-Verona/Papers/P-Lavazza.pdf; Martella V, et al. Exploring the epidemiology of lapine rotaviruses: evidence for spreading of rotaviruses displaying the newly-recognized P(22) VP4 gene allele in Italy. Proc 8th World Rabbit Congress; Sep 2004; Puebla, Mexico. Available from: https://world-rabbit-science.com/WRSA-Proceedings/Congress-2004-Puebla/Papers/Pathology/P-Martella.pdf).6,80,81
Coronavirus
Rabbit coronavirus (RbCoV) is another virus involved in rabbit enteritis (Lavazza A, Capucci L, 2008), although it was also detected in healthy animals. 61 Viral load in fecal samples was determined using two methods: a universal coronavirus RT-PCR with primers amplifying the RdRp gene of known CoVs, and a virus-specific RT-qPCR targeting the same virus gene. 61
Vesivirus
A new rabbit vesivirus (RaV), a member of the Caliciviridae family, was detected in diarrheic rabbits. 83 Microscopy and biochemical and virologic methods were used to characterize the virus isolate. One-step RT-PCR with primers amplifying the full-length of the virus genome was employed in order to identify this putative rabbit calicivirus. Although rabbit caliciviruses share common features of genome organization and a high degree of sequence similarity, 83 it is unlikely that PCR methods developed for the specific detection of lagomorph viruses from different genera of the Caliciviridae family could also be used for detection of rabbit vesiviruses.
Astrovirus
RT-rtPCR was employed for astrovirus (AstV) detection in asymptomatically infected animals and in rabbits with enterocolitis. 82 Use of the broadly reactive AstV primers allowed detection of viral RNA in fecal samples with a method detection limit of 10 genomic equivalents (GEs) per reaction. Another approach that improves detection of rabbit enteric viruses is a Virochip microarray used in conjunction with PCR and sequencing. Its application showed the presence of AstVs in an outbreak of rabbit enterocolitis associated with high animal mortality. 104 The same PCR primers, as used in a combined PCR and microarray protocol targeting the RdRp AstV gene, were applied in studies assessing the prevalence of AstV infections in commercial rabbit farms in Canada. 116 The virus was detected in both ill and healthy rabbits, with infections having no clinical significance for their health.
Herpesvirus
A new virus called leporid herpesvirus 4 (LHV-4; species Leporid alphaherpesvirus 4) was detected in pet and farmed rabbits in Alaska by the use of a universal alphaherpesvirus PCR. 47 The highly conserved region that encodes a large subunit of the ribonucleotide reductase (RR1) gene of the human herpesvirus was used to design degenerate primers. In order to determine the extent of LHV-4 infection, tissues of experimentally infected rabbits were tested using an LHV-4–specific PCR protocol. The RR1 or entire LHV-4 RR gene was chosen in other PCRs aiming at virus detection and evaluation of distribution in rabbit tissues.11,106
Hepatitis E virus
Hepatitis E virus (HEV; species Orthohepevirus A) has been detected in farmed, wild, and pet rabbits worldwide,3,12,17,21,43,46,49,95,115,116,122 with evidence also existing for virus circulation in European brown hares. 43 Although rabbit HEV strains belong to a cluster of the rabbit HEV genotype, they are genetically and antigenically closely related to other HEV strains detected in pigs and wild boars, which are known to infect humans.28,50,120 Therefore, HEV detection in rabbits is conducted using PCR primers targeting the open reading frame (ORF)1, 122 ORF2, 21 or ORF346,51 genome sequences of swine, human, or rabbit HEV strains.
Novel methods
Currently applied molecular methods for the detection of rabbit viruses have limited potential for multiplexing; therefore, the number of pathogens detected in a single assay is relatively small. A new approach allowing simultaneous screening of the sample for several viral pathogens is a Virochip microarray. 110 The assay is based on a microarray technique with a panel of oligonucleotide probes present on a chip. The Virochip method proved its usefulness in identification of AstV infection in an outbreak of gastroenteritis in rabbits. 104 Although Virochip offers comparable sensitivity and specificity to pathogen-specific PCR methods,20,58 its routine use is limited because of high costs and a burdensome workload.
An xTAG assay (Luminex, Austin, TX) has been developed for simultaneous detection of RHDV, LRV, and Sendai virus (SV; species Murine respirovirus) in swabs, feces, and rabbit tissues. 114 This method involves PCR amplification of VP60 (RHDV), VP4 (LRV), and N (SV) virus gene fragments using sets of biotinylated primers containing a unique “TAG” sequence coupled with a multiplexed bead-based suspension array detection system. The xTAG assay was rapid, labor-saving, and sensitive, with a detection limit of 100 virus genome copies per µL of the tested sample.
Next-generation sequencing (NGS) is becoming a popular and frequently used tool in molecular virology. For instance, Illumina sequencing technology (Illumina, San Diego, CA) was employed for the detection of the first cases of RHDVa infection in domestic and wild rabbits in Australia. 77 Analysis of the complete viral genome sequences revealed that all detected RHDVa strains were recombinants between RHDVa and nonpathogenic RCV-A1–like viruses. NGS has also been used for the analysis of virus genes associated with rabbit resistance to natural RHDV infection. The use of a “genotyping by sequencing” technique identified biologically interesting genes that might be contributing to increased rates of wild rabbit survival in Australia. 99
Apart from nucleic acid–based methods used for identification and characterization of rabbit viruses, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) could be employed. MALDI-TOF MS is a powerful tool in the analysis of nucleic acids 52 and provides insights into the structure of viral particles. 121 For example, it has been used in studies aimed at the characterization of the major capsid protein of MYXV. 121 The application of MALDI-TOF MS in virology is less advanced than the detection of bacterial pathogens in clinical microbiology, possibly because of the low protein content and higher molecular weight of viral proteins. 101 The sensitivity and detection limits of human DNA viruses by MALDI-TOF MS were high and comparable to oligonucleotide microarrays and multiplex PCR. 102
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was funded by the KNOW (Leading National Research Centre) Scientific Consortium “Healthy Animal–Safe Food”, under the Ministry of Science and Higher Education, Poland (decision 05-1/KNOW2/2015).
