Abstract
Rabbit hemorrhagic disease virus 2 (RHDV2), a virulent and contagious viral pathogen that affects wild and domestic lagomorph populations, was identified in Wyoming, USA in December 2020. A surveillance program was developed involving full-carcass submission and liver analysis, although carcass quality as a result of predation and decomposition impeded analysis. To increase the number of submissions and provide flexibility to field staff, we evaluated 2 sample types: 77 dried blood on filter paper samples, 66 ear punch samples. At initial sampling, test specificity and sensitivity of the RT-rtPCR utilizing dried blood on filter paper and ear punch samples were both 100% compared to liver. Filter paper results were consistent over time; sensitivity stayed >96% through weeks 2, 4, and 6, with a maximum mean difference of 6.0 Ct from baseline liver Ct values (95% CI: 5.0–7.3) at 6 wk. Test sensitivity of the ear punch sample at 1, 3, 5, and 7 wk post-sampling remained at 100%, with a maximum mean difference of 5.6 Ct from baseline liver Ct values (95% CI: 4.3–6.9) at 5 wk. Filter paper and ear punch samples were suitable alternatives to liver for RHDV2 surveillance in wild lagomorph populations. Alternative sampling options provide more flexibility to surveillance programs, increase testable submissions, and decrease exposure of field personnel to zoonotic disease agents.
Keywords
Rabbit hemorrhagic disease (RHD) is an acute and fatal viral hepatitis of European rabbits (Oryctolagus cuniculus), caused by a nonenveloped, single-stranded, positive-sense RNA virus (RHDV; Caliciviridae, Lagovirus).2,10,22 Two main genotypes have been identified, GI.1 (RHDV) and GI.2 (RHDV2), which include several variants 3 ; RHDV2 GI.2 was first reported in France in 2010, and, unlike RHDV GI.1, this variant can cause infections in both younger and older animals and has been shown to cause morbidity and mortality in native North American lagomorphs.7,16,17 RHDV2 was first detected in southeastern Wyoming, USA, December 2020, in a wild cottontail rabbit. 31 In Wyoming, the list of potentially susceptible wildlife includes hares of the genus Lepus (snowshoe hare, L. americanus; black-tailed jackrabbit, L. californicus; white-tailed jackrabbit, L. townsendii), North American cottontails of the genus Sylvilagus (eastern cottontail, S. floridanus; mountain cottontail, S. nuttallii; desert cottontail, S. audubonii), pygmy rabbits (Brachylagus idahoensis), and the American pika (Ochotona princeps). 27 Although the current risk to these populations is not well defined, Wyoming has a strong interest in closely monitoring the outbreak given the important economic and ecologic status of these species, particularly species of concern, such as the pygmy rabbit.
Wildlife disease monitoring and surveillance is essential for understanding the epizootiology of an infectious disease within wildlife populations. 20 Some of the limitations of wildlife disease monitoring and surveillance strategies are related to case acquisition, lack of properly validated diagnostic tests, and environmental characteristics within the natural systems wherein samples are procured. 26 Limitations in prior RHD surveillance programs have included the inability of obtaining fresh carcasses, which limited studies to syndromic surveillance only. 5 Having flexibility in validated sample types could overcome some of these constraints by increasing sample submissions. When RHDV2 is suspected, the “gold standard” tissue for testing is liver. 22 Some pathogens may be more difficult to detect outside of the gold standard sample types or may be dependent on the duration of time between exposure and sampling.5,29
In most cases, it is preferable to submit a whole carcass; however, obtaining a specific tissue (i.e., liver) or an intact carcass can be challenging. Small prey species disappear from the landscape quickly as a result of predation or scavenging. 5 Given the difficulty of locating fresh carcasses on the landscape (e.g., large land mass, sparsely populated area), the carcasses, when found, are often desiccated and degraded.20,25,26 Finally, human safety and biosecurity can play a role when there is a potential zoonotic disease risk (e.g., plague [Yersinia pestis] and tularemia [Francisella tularensis]) to personnel or the public when collecting and/or transporting tissues. 28 Moving carcasses from one area to another could also result in the unintentional spread of pathogens and/or disease vectors. 12
Providing diagnostic sampling flexibility to a surveillance or monitoring program requires test validation for each sample. We evaluated 2 sample types: blood filter paper and ear punch samples. Filter paper has been validated as a suitable sample type for disease detection (e.g., avian influenza, Chagas disease),11,14 and allows for sample collection in the field with little training, equipment, and without the need for climate-controlled transport. 6 Samples must be eluted from filter paper before analysis, leading to lower sensitivity in some assays.1,21 The quality of RNA in blood stored on filter paper diminishes over time, which affects test sensitivity and sometimes completely precludes the detection of viral RNA. 21 Ear punch samples taken in the field have also been used successfully (e.g., bovine viral diarrhea virus, African swine fever virus).15,24 Collecting whole ears requires limited equipment, and ears tend to be available on the landscape longer compared to organs that may have been lost to predators, scavengers, or decay. 23 However, there may be limited utility of using dermal tissue biopsies for pathogen detection, attributed to pathogen predilection for specific tissue or fluid types, normally reserved for gold standard testing. 30
Therefore, our main objective was to validate the use of filter paper and ear punch samples, using reverse-transcription real-time PCR (RT-rtPCR), for the detection of RHDV2 in free-ranging native lagomorph species. Additionally, we investigated the ability to detect RHDV2 from alternative samples long-term, attempting to mimic environmental conditions that can affect typical tissue or carcass submissions collected in semi-arid, sagebrush steppe, or similar habitat.
Materials and methods
Sample collection
Intact carcasses of native free-ranging lagomorphs (hereafter, cottontail(s) and jackrabbit(s)) were collected from across the state of Wyoming, USA from March 2021 to June 2021. Carcasses for our study were gathered by field personnel opportunistically and were submitted to the Wildlife Health Laboratory, Wyoming Game and Fish Department (Laramie, WY, USA). Cottontail and jackrabbit species were determined morphologically. All 3 of Wyoming’s native cottontail species share very similar morphologic characteristics, and for the purposes of our study, no genetic evaluation was performed to differentiate species. Blood samples absorbed onto filter paper (Nobuto blood sampling paper type 1; Advantec), liver, lung, spleen, and ears were collected during routine postmortem examination of submitted carcasses. Pooled liver, lung, and spleen was submitted to the Wyoming State Veterinary Laboratory (WSVL; Laramie, WY, USA) for screening of plague and tularemia using a fluorescent antibody test (FAT). All samples were kept at 4°C throughout the screening for plague and tularemia (<84 h). If not extracted immediately after screening, liver and ears were kept at either −20°C or −70°C until RNA extraction occurred (<72 h); filter paper was kept at room temperature.
Sample processing and RNA extraction
All samples were extracted using a commercial kit (E.Z.N.A total RNA kit 1; Omega Bio-tek). Primers and probes specific for RHDV2 9 (RHDV2-F: 5′-TGGAACTTGGCTT GAGTGTTGA-3′; RHDV2-R: 5′-ACAAGCGTGCTTGT GGACGG-3′; RHDV2 probe: 5′-6-FAM-TGTCAGAAC-ZEN-TTGTTGACATCCGCCC-IABkFQ-3′) were used in a 25-µL reaction with primers and probe at a final concentration of 1 µM and 0.2 µM, respectively, and utilizing 5 µL of RNA template. Samples were analyzed using an adapted optimized RT-rtPCR (CFX96 Touch; Bio-Rad) thermocycling protocol 9 provided by the USDA Animal and Plant Health Inspection Services, Foreign Animal Disease Diagnostic Laboratory (Greenport, NY, USA).
Fifty mg (±2 mg) of liver were placed into a 2.0-mL screw-top tube containing one 4-mm glass bead and 500 µL of chilled ultra-pure PBS (PBS 1 × sterile; VWR). Samples were kept on ice, then homogenized (Bead Ruptor Elite 24; Omni International; 0.8 speed × 45 s, hold 15 s, 0.8 speed × 45 s), and centrifuged at 5,870 × g for 2 min. To begin the extraction, 10 µL of supernatant was added to 350 µL of TRK lysis buffer with 2% 2-mercaptoethanol (β-ME). All samples were then extracted according to kit instructions and eluted in 70 µL of nuclease-free water.
The left ear (right ear if left was not intact) was removed from the carcass and prepared by removing the fur from the base of the medial aspect, using a standard single-edge razor blade. A 6-mm punch (MedBlades) was used to remove a full-thickness sample, from the rostral aspect at the base of the ear over the central auricular artery (Fig. 1). The ear punch sample was minced and placed into a 2.0-mL screw-top tube with one 4-mm glass bead and 700 µL of TRK lysis buffer with 2% β-ME, homogenized (Bead Ruptor Elite 24; 0.8 speed × 45 s, hold 15 s, 0.8 speed × 45 s), and placed into a 55°C heat block for 120 min. After incubation, the ear sample was centrifuged at 5,870 × g for 2 min, and 350 µL of supernatant were used for RNA extraction, as described above. An initial ear punch sample was analyzed within 1 wk of postmortem examination and retained at −20°C thereafter. A subset of 20 of the ears retrieved from carcasses was suspended individually with 12-ga steel wire through the initial ear punch site, within 3 cage traps (Original series live trap, model 103; Tomahawk), ensuring that no ears touched. To prevent scavenging, the Tomahawk traps containing ears were suspended inside 3 secondary wire cages (41 × 76 × 76 cm), ~91 cm above a concrete pad, and placed outside from August 2, 2021, to September 20, 2021. Average maximum daily temperature and precipitation during the 7-wk period were 26.4°C and 0.08 cm, respectively.18,19 Ears were retrieved at 1, 3, 5, and 7 wk to remove subsequent ear punch samples; these samples were extracted and analyzed immediately following collection.
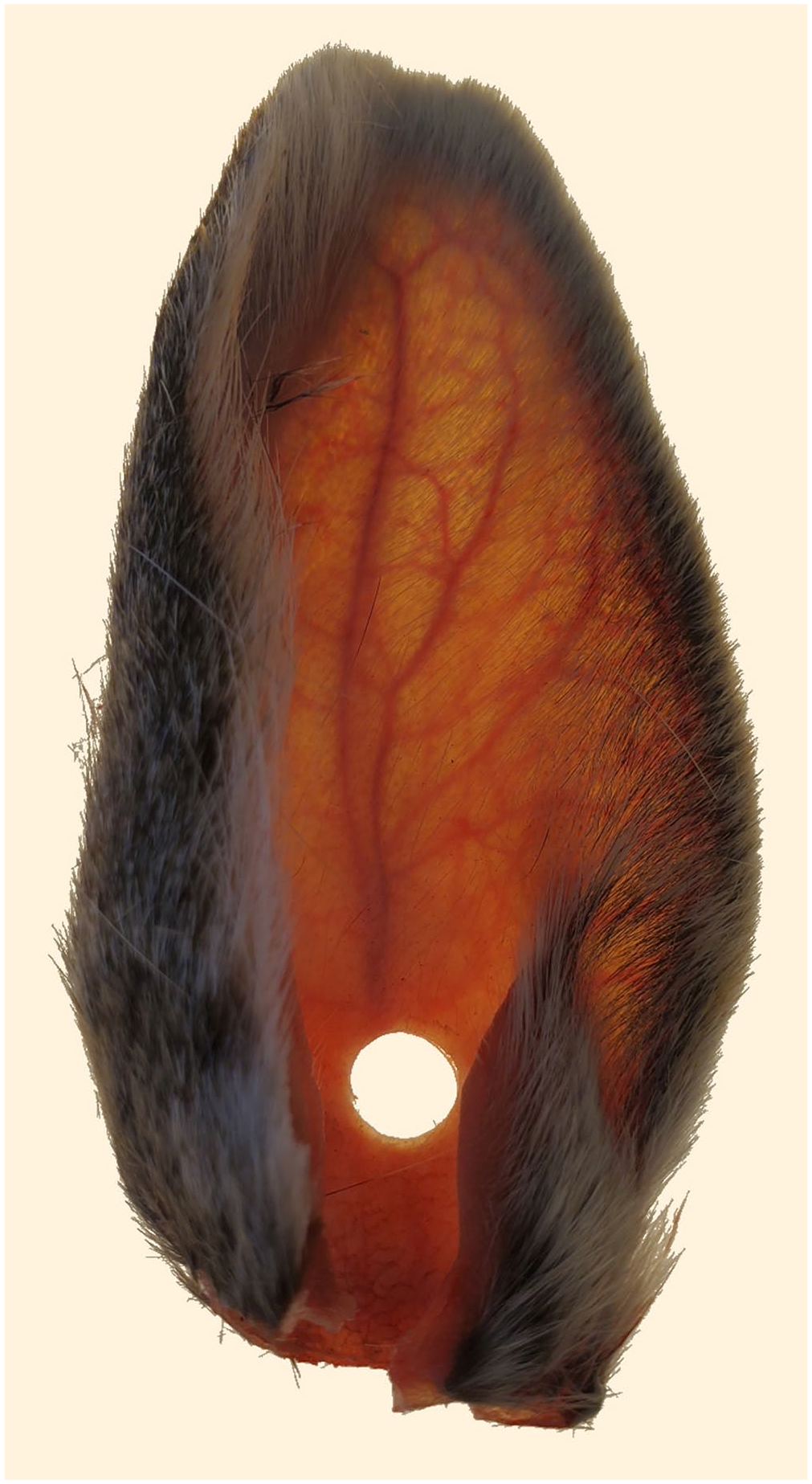
Sylvilagus spp. ear with 6-mm punch sample removed from anterior aspect at the base of the ear over the central auricular artery.
Four filter paper strips per animal were immersed in abdominal blood during postmortem carcass examination allowing contact of the blood with the absorption area of the paper (equivalent to 100 µL of whole blood per strip). 8 All 4 filter paper strips were immediately placed into a paper envelope to dry completely. The filter paper was 5 mm wide, and a sample was removed with a 6-mm punch (MedBlades) by targeting an area 1 mm from the end of the filter paper strip, retaining that extra 1 mm for ~28.3 mm2 of blood (Fig. 2). This punch was placed into a 2.0-mL microcentrifuge tube (Globe Scientific) and eluted with 200 µL of TE buffer (TE 1×, pH 7.4; Quality Biological) for a minimum of 10 min; 140 µL of this elute was added to 350 µL of TRK lysis buffer with 2% β-ME to begin RNA extraction. 1 All samples were extracted immediately after elution. One filter paper strip was processed within 1 wk of the postmortem examination; subsequent filter papers were processed at 2, 4, and 6 wk after postmortem examination. Filter paper samples that tested negative at week 1 from animals whose liver sample also tested negative were not subsequently processed at weeks 2, 4, or 6.
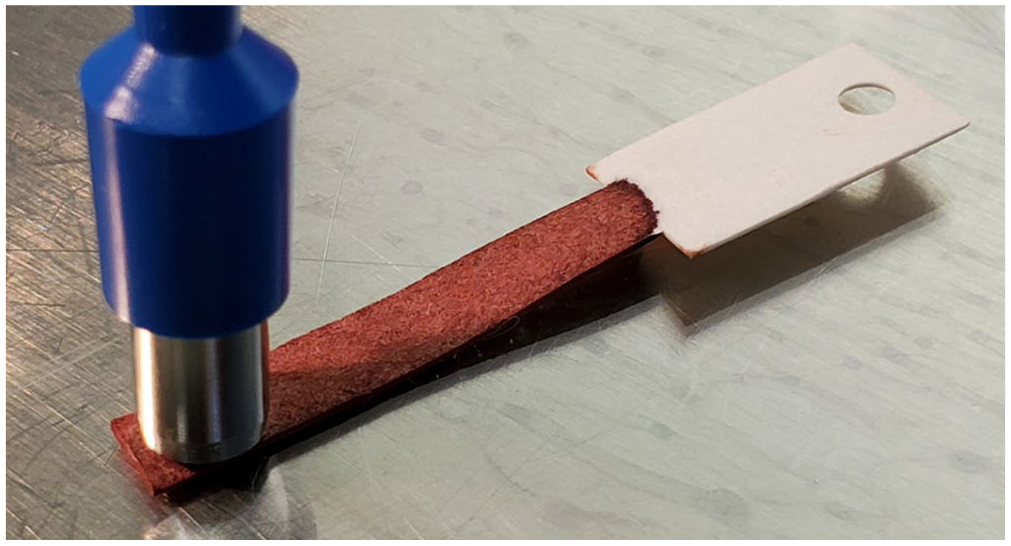

Six-mm punch equivalent taken from the end of the absorption area of the filter paper.
Test sensitivity and specificity of filter paper and ear punch samples at baseline (<1 wk postmortem) and over time
The validity of ear punch samples and filter paper samples were assessed with test sensitivity and specificity using baseline (<1 wk postmortem sampling) liver test results as the gold standard and a Ct value cutoff of 30, with anything ≤30 considered a positive test result. To examine the performance of filter paper and ear punch methods over time compared to the liver test results, test sensitivity was also calculated at longitudinal times following sample collection (negative samples were not re-tested; thus, test specificity was not assessed). For both sets of analyses, an overall assessment was performed for all rabbit samples together, as well as for the subset of jackrabbit samples, to assess any differences by species.
Statistical analysis of viral retention at baseline and over time
A 2-sided Wilcoxon sign rank test was used to test for significant differences between Ct values for each novel sampling method at various times paired with the corresponding liver baseline Ct value. Only positive samples according to liver results were included, because negative samples were not re-tested over time for either novel method. For both filter paper and ear punch methods, a separate assessment was performed for all cottontail samples (excluding jackrabbit samples) along with an overall assessment (all species), to determine any differences by species. All tests were 2-tailed, and a Bonferroni correction was applied to account for multiple testing; the overall significance level α = 0.05 was divided by the number of time intervals assessed for each method (n = 5 for the ear punch method; n = 4 for the filter paper method). 4
Results
We collected 70 paired liver and filter paper samples from free-ranging cottontails and 7 from black-tailed jackrabbits. Additionally, ear punch samples were collected from 60 cottontails and 3 black-tailed jackrabbits with paired liver and filter paper samples. All samples submitted to the WSVL from our study tested negative for both plague and tularemia.
Test sensitivity and specificity of filter paper and ear punch samples at baseline and over time
Sensitivity and specificity of the RT-rtPCR, utilizing both the filter paper and ear punch from all animals, were 100% at baseline sampling. In the subset of jackrabbit samples, test sensitivity and specificity were 100% when analyzing the filter paper samples at baseline. Ear punches taken from jackrabbits at baseline also elicited a value of 100% test sensitivity; test specificity calculation was not possible given the lack of negative animals (Table 1). Longitudinal test sensitivity for filter paper (all samples) was >96% at 2, 4, and 6 wk. Longitudinal test sensitivity for filter paper (jackrabbits only) was 83.3% at 2 wk, 100% at 4 wk, and 80% at 6 wk. Longitudinal test sensitivity of ear punches over time was 100%.
Baseline (<1 wk postmortem sampling) test sensitivity and specificity of filter paper (Nobuto) and ear punch methods for the detection of rabbit hemorrhagic disease virus 2 via reverse-transcription real-time PCR* compared to paired liver samples.
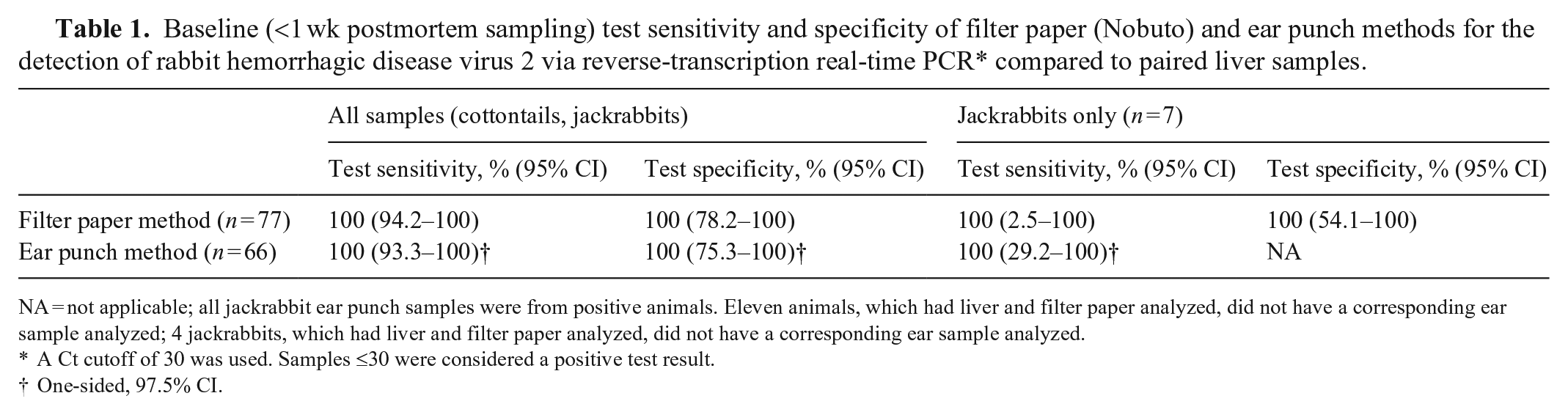
NA = not applicable; all jackrabbit ear punch samples were from positive animals. Eleven animals, which had liver and filter paper analyzed, did not have a corresponding ear sample analyzed; 4 jackrabbits, which had liver and filter paper analyzed, did not have a corresponding ear sample analyzed.
A Ct cutoff of 30 was used. Samples ≤30 were considered a positive test result.
One-sided, 97.5% CI.
Viral retention at baseline and over time
Based on raw data, there appeared to be a correlation between liver Ct values and those obtained using ear punch and filter paper methods, although the correlation was more apparent for the ear punch method (Fig. 3). The filter paper method appears to be consistent over time; qualitatively, the relationship between the Ct values derived from the filter paper versus the baseline liver sample was similar between weeks 2, 4, and 6 (Fig. 3A). Raw data from the ear punch method showed the same correlational pattern from baseline until 3 wk, but weeks 5 and 7 revealed a different trend, wherein the slope of the segmented linear regression appears to level off or decrease for higher liver Ct values (Fig. 3B).
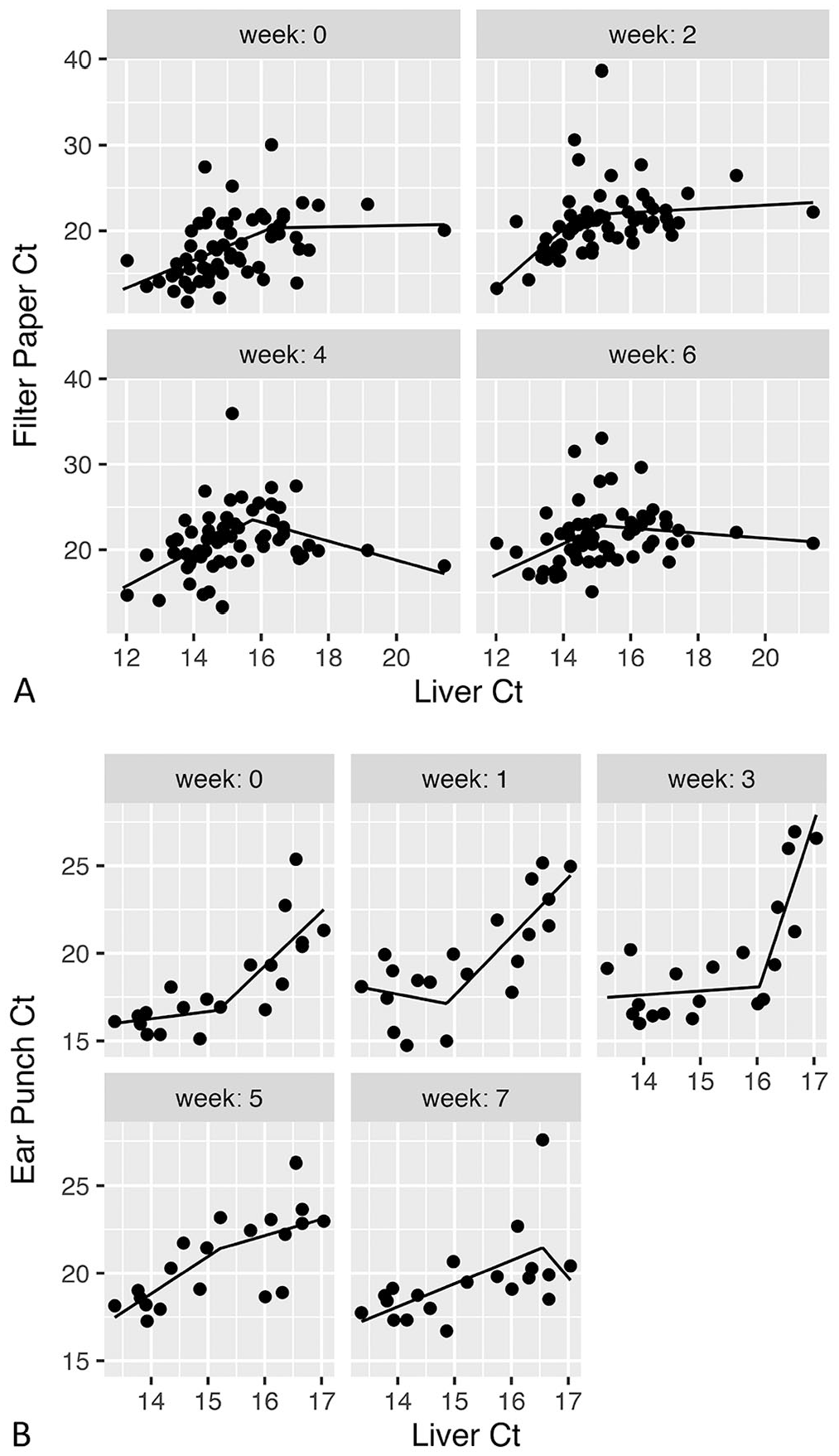
Segmented linear regression of Ct values for rabbit hemorrhagic disease virus 2 over bi-weekly intervals (starting with <1 wk postmortem sampling (week 0) for
Both filter paper and ear punch Ct values were higher initially than liver Ct values and tended to increase over time (although Ct values for ear punch samples decreased in certain weeks). The average difference in Ct values between filter paper and liver samples was lowest at baseline, increased at 2 wk, and remained relatively stable until 6 wk; the average difference for these longitudinal time points was consistently 5–6 Ct values higher than liver samples (Fig. 4A; Table 2). Similar to filter paper, the average difference in Ct values between paired ear punch and liver samples was lowest at baseline, increased at 1 wk and fluctuated between 1 wk and 7 wk. The average difference in Ct values for these longitudinal time points was 4–5.6 Ct values higher than liver samples (Fig. 4B; Table 3). The maximum mean difference in Ct value from liver baseline samples was 6.0 (95% CI: 5.0–7.3) in filter paper samples at 6 wk, and 5.6 (95% CI: 4.3–6.9) in ear punch samples at 5 wk (Tables 2, 3). There did not appear to be any major difference in mean Ct differences after exclusion of the subset of jackrabbit samples (Fig. 4).
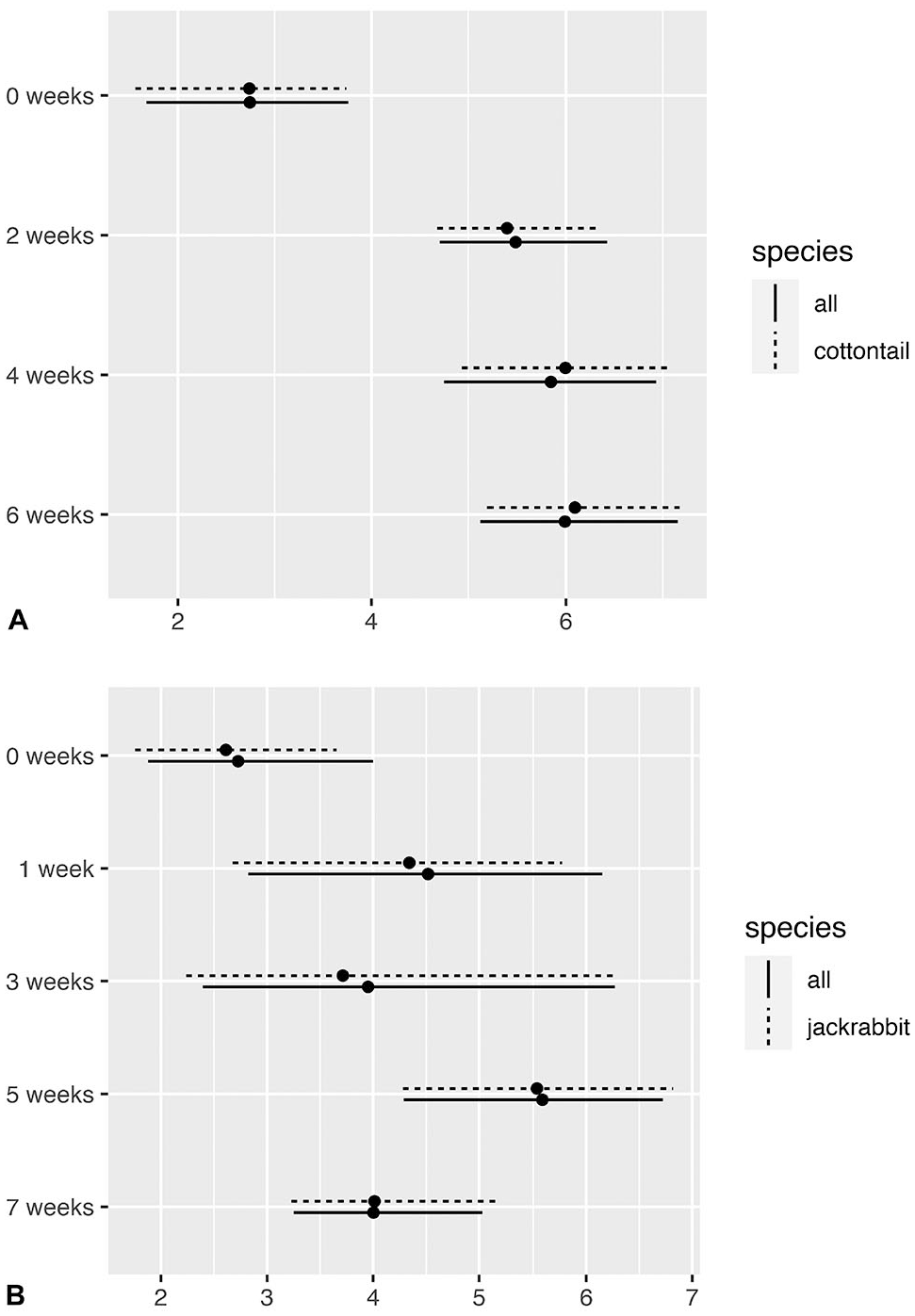
Viral retention of rabbit hemorrhagic disease virus 2 over time (starting with <1 wk from postmortem exam sampling (week 0)) for
Mean difference* between Ct values, via rabbit hemorrhagic disease virus 2–specific reverse-transcription real-time PCR, using filter paper (Nobuto) at <1, 2, 4, and 6 wk postmortem sampling compared to liver at baseline (<1 wk postmortem sampling).
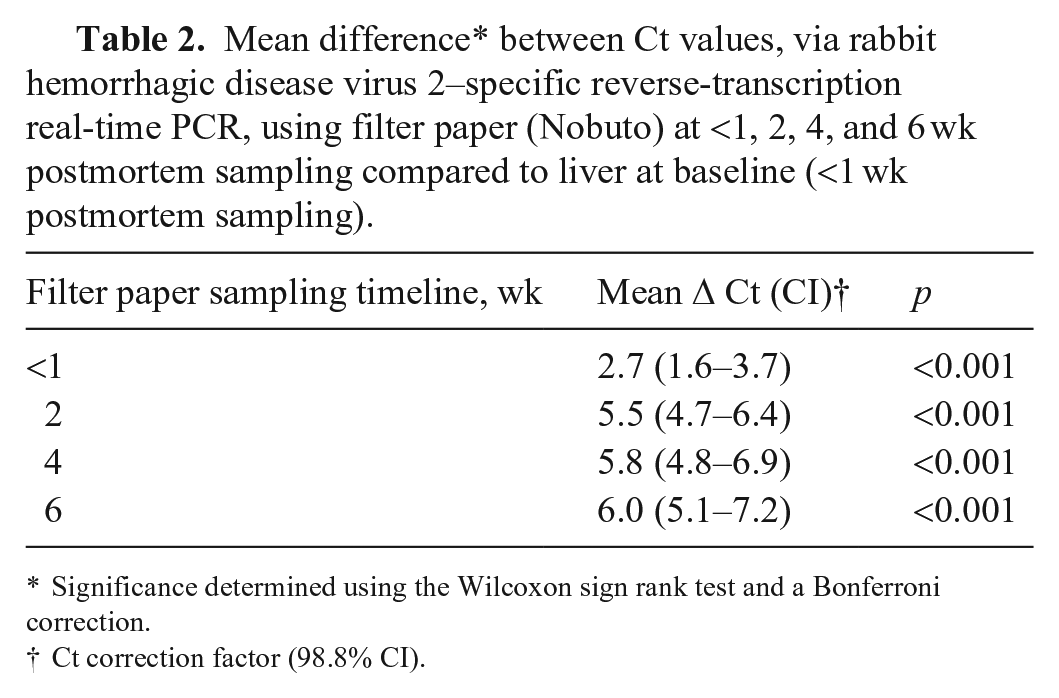

Significance determined using the Wilcoxon sign rank test and a Bonferroni correction.
Ct correction factor (98.8% CI).
Mean difference* between Ct values, via rabbit hemorrhagic disease virus 2–specific reverse-transcription real-time PCR, determined using ear punch samples at <1 wk postmortem exam sampling, and after 1, 3, 5, and 7 wk outside compared to liver Ct values at baseline (<1 wk postmortem sampling).
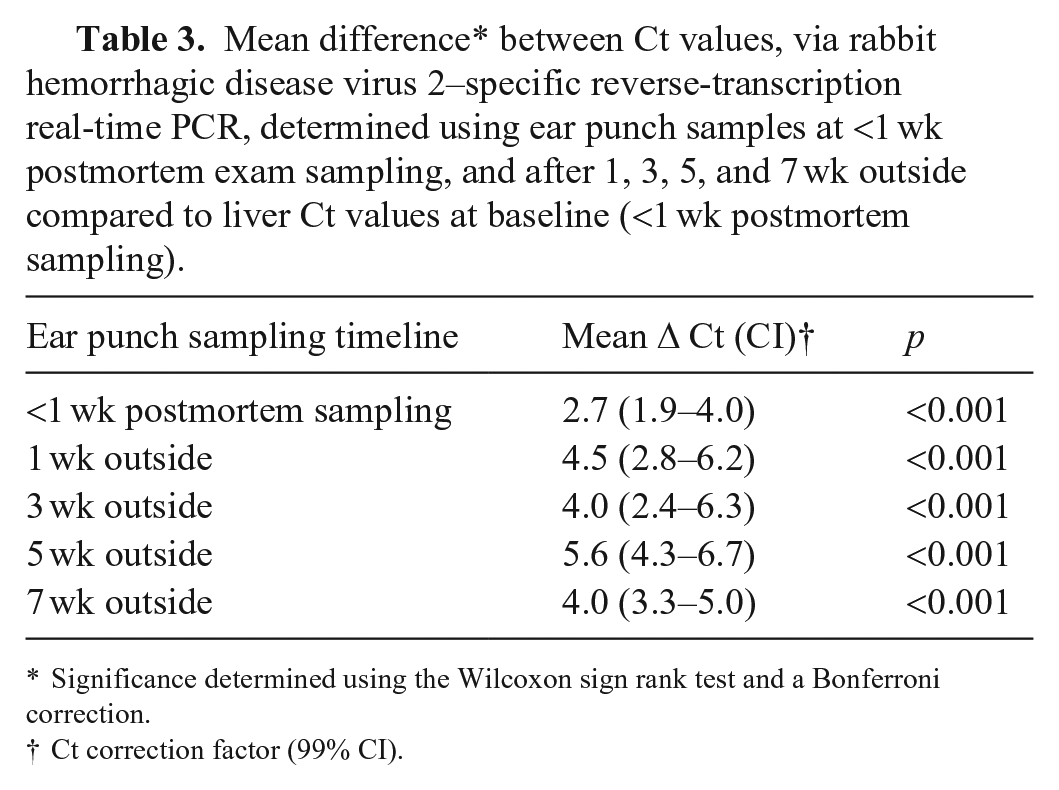
Significance determined using the Wilcoxon sign rank test and a Bonferroni correction.
Ct correction factor (99% CI).
Discussion
Based on our analyses, both the filter paper and ear punch methods have potential utility in the field for surveillance of RHDV2. The performance of both tests is highly congruent with liver sampling, with 100% test sensitivity and specificity at initial sampling. Although test sensitivity of the filter paper method declines slightly over time, it is unknown whether the filter paper samples from 2 positive animals that were missed at longitudinal sampling points would have undergone a similar decline in detectable viral RNA. Bearing in mind that test performance over time is a product of viral retention in tissues and blood over time, our assessment of test performance at baseline is perhaps the most valid comparison. Our longitudinal comparison of Ct values using filter paper compared to liver baseline values should be interpreted as a guide for sample storage time and conditions when delivery to the laboratory cannot be immediate. The longitudinal comparison of Ct values using ear punch methods compared to liver at baseline should be interpreted as a guide for viable sample collection in the field from a decayed carcass and storage time before delivery to the laboratory.
On average, Ct values measured using filter paper and ear punch methods were higher (i.e., less detectable RNA) than those measured from liver samples. The average Ct value of all positive livers analyzed in our study (all species) was 15.1 (95% CI: 14.7–15.5). Although Ct values of the alternative methods were significantly different, and higher than liver, the maximum mean change in Ct value for both sample types initially and over time was small (filter paper 6.0, ear punch 5.6). For both filter paper and ear punch samples, the maximum mean increase in Ct value would still be considered positive with the PCR assay that was used, and therefore both sample types are considered appropriate alternatives for RHDV2 surveillance in wild lagomorph populations. Based on mean Ct differences from baseline liver samples, the filter paper method demonstrated greater consistency in viral retention over time compared to the ear punch method, based on narrower CIs, and less variation in Ct mean differences at longitudinal times. In contrast, the ear punch method demonstrated slightly increased viral retention over time, although this method was not as consistent over time as the filter paper method.
The lower consistency of Ct values over time in ears could be attributed to continued decomposition or desiccation over time; these processes could also have been inconsistent across ears. 23 It is also important to consider the variation in sample location 13 on the ear as a result of removal of preferred positions for punches during previous sampling. The differences in accuracy and consistency between these sample types have been included to help guide future validations of these testing methods in experimental and field settings, particularly if longer sampling intervals are assessed.
Plague and tularemia screening presents a challenge when utilizing alternative RHDV2 sample types. To date, these sample types are not validated for the FAT or rtPCR. Ears of 2 tularemia-positive rabbits were analyzed via a Francisella tularensis/Yersinia pestis multiplex rtPCR (PCR primers and probes are considered proprietary to the Centers for Disease Control and Prevention, Laboratory Response Network), and although the Ct values of the ears were higher compared to the pooled tissues, Francisella was still detected in both ears. Further research is required to validate ear punches and filter paper for the plague/tularemia rtPCR. Validating ear punches would allow for continued surveillance of these diseases in RHDV2 suspected animals as well as provide a safer sampling method for these zoonotic diseases.
The results of our study are encouraging, potentially leading to further research of this sample type. Since validating ear punch sampling, we have been able to document RHDV2 by utilizing ears in 2 cases, one in which a whole carcass submission was not possible because of scavenging.
Validating alternative sample types for established tests provides flexibility to wildlife surveillance and monitoring programs; however, our work also carries inherent limitations and biases. In both alternative sample types, Ct values were higher initially than liver and increased over time. This could result in a false-negative PCR result in samples with a low viral load, initially, and over time. In our study, virus was still detectable in filter paper at 6 wk and in ears at 7 wk. Performing analyses as soon as possible would maximize RNA detection, decreasing the risk of a false-negative PCR result, but based on our results, that risk appears to be low. Utilizing ears may decrease risk of exposure to zoonotic pathogens by limiting exposure to, and movement of, the carcass. Utilizing filter paper, however, could increase the risk of exposure to the sampler by requiring the carcass to be opened in the field, but would decrease chances of unintentional spread of the disease with movement. Receiving full carcass submissions could also limit environmental contamination of samples, given that the sample would be contained internally until being extracted aseptically in the laboratory.
Footnotes
Acknowledgements
We thank members of the field staff of the Wyoming Game and Fish Department’s Casper, Cody, Lander, Laramie, Green River, and Sheridan regions, and especially Parker Everhart. We also thank the USDA Foreign Animal Disease Diagnostic Laboratory for sharing their standard operating protocols for the RHDV2 RT-rtPCR. Additionally, thank you to Dr. Kerry Sondgeroth, Hally Killion, and Madison Vance of the Wyoming State Veterinary Laboratory for providing plague and tularemia analysis. We are also grateful to the public of Wyoming for their diligence in lagomorph carcass submissions. Finally, we thank the JVDI editor-in-chief and reviewers for their thoughtful comments and suggestions.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Funding for this work was provided by the Wyoming Game and Fish Department.
