Abstract
Enteric disease in pigs is usually of multifactorial etiology, including infectious and non-infectious factors. In many cases of endemic diarrhea in weaner-to-finisher pigs, the combination of 2 or more microorganisms leads to aggravation of intestinal lesions and, consequently, clinical signs. We autopsied a 4-mo-old fattening pig with diarrhea and diagnosed severe fibrinonecrotizing typhlocolitis. Numerous spiral-shaped bacteria and amoeba-like PAS-positive protozoa were observed in the cecal and colonic mucosa and submucosa. Brachyspira hyodysenteriae was detected by PCR from colonic content. By in situ hybridization, large numbers of Entamoeba polecki were found within the lamina propria and submucosa; moderate numbers of Blastocystis sp. and scattered trichomonads were present in intestinal content. In addition, Entamoeba polecki, Balantidium spp., Blastocystis sp., and Trichomonas sp. were also detected by PCR.
Enteric disorders in swine are usually of multifactorial origin, including combinations of microorganisms plus the concurrence of different non-infectious risk factors. 4 Coinfection with 2 or more agents often causes enhanced mucosal inflammation. In addition, damage to the intestinal epithelial barrier may allow the uncontrolled proliferation of other organisms that would be harmless under healthy conditions.
Swine dysentery (SD) is one of the most severe enteric diseases of pigs. Brachyspira hyodysenteriae, the etiologic agent of SD, is a beta-hemolytic spirochete able to cause significant large intestinal lesions, without the need for other coinfecting agents. 5 However, several coinfecting agents have been described in pigs affected by SD, including Campylobacter spp., Fusobacterium necrophorum, and Bacteroides vulgatus.2,6,20 Protozoan overload is also a usual finding in enteric lesions, including in cases of SD.1,16,19
We describe herein the coinfection of B. hyodysenteriae and Entamoeba polecki leading to severe necrotizing lesions in colon and cecum in a diarrheic pig. A 4-mo-old, crossbred pig, from a fattening unit (site 3) on an indoor pig production farm, with ongoing problems of diarrhea died and was submitted to the Servei de Diagnòstic de Patologia Veterinària of the Veterinary Faculty of the Universitat Autònoma de Barcelona (Spain) for diagnostic purposes. The farm of origin was a 1,350 sow farm that was positive-stable for porcine respiratory and reproductive syndrome virus (PRRSV), seropositive to porcine circovirus 2 (PCV-2) and Mycoplasma hyopneumoniae, and negative to pseudorabies virus. Sows and piglets were vaccinated with a modified-live PRRSV vaccine. The affected pig belonged to a batch of 450 grower–finishers allocated to pens with a complete slatted floor; animals received a conventional finishing feed. Clinical problems were observed only in fatteners (14–16-wk-old), with ~10% of animals with mucoid-red diarrhea starting 1 mo after entering the fattening unit. The field veterinarian established a differential diagnostic list including porcine proliferative enteropathy, SD, and colibacillosis.
At autopsy, the pig was emaciated and pale. Severe diffuse fibrinonecrotizing hemorrhagic typhlocolitis was observed, with abundant mucous exudate and bloody fluid in the lumen (Fig. 1). Nasal turbinates had severe bilateral atrophy. No other lesions were observed grossly.
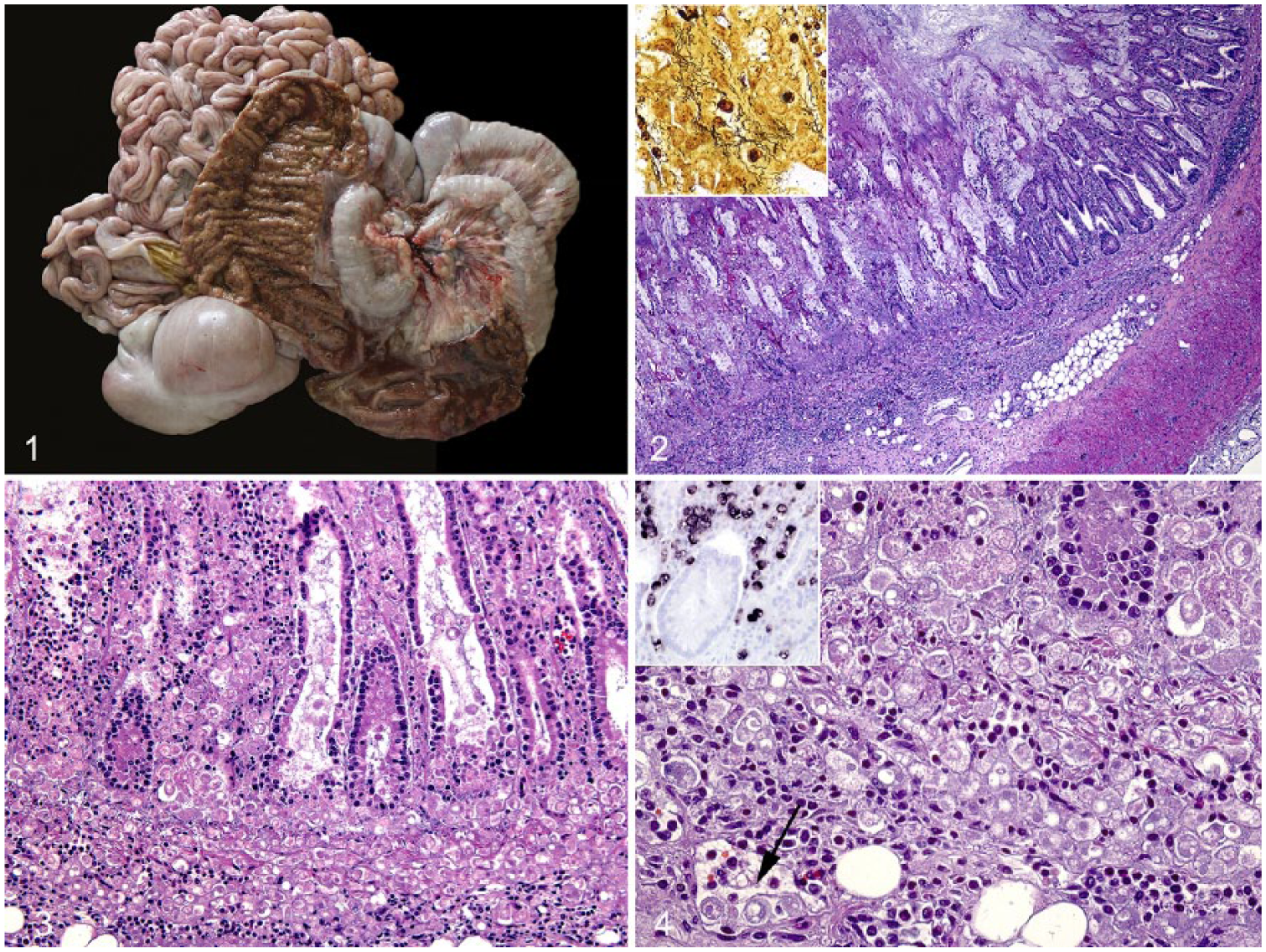
Macroscopic and microscopic lesions of colon and cecum of a pig coinfected with Brachyspira hyodysenteriae and Entamoeba polecki.
Samples of lung, spleen, liver, kidney, mandibular and superficial inguinal lymph nodes, skeletal muscle, heart, stomach, ileum, colon, cecum, and brain were collected and fixed by immersion in 10% buffered formalin at room temperature for 48 h before routine processing; slides were stained with hematoxylin and eosin. Immunohistochemical staining to detect PRRSV 11 and PCV-2 18 was performed on lung and lymphoid tissues (tonsil, lymph nodes, and spleen) as part of the investigation. In both cases, samples were negative.
Histologically, severe diffuse necrosis of the apical two-thirds of the colon and cecal mucosa, or complete mucosal necrosis, was observed in the colon and cecum (Fig. 2). Necrotic and sloughed epithelial cells were present in the lumen admixed with abundant mucus, degenerate neutrophils, fibrin, and myriad rod- and spiral-shaped bacteria. The latter finding was confirmed by Warthin–Starry stain, in which numerous spiral-shaped bacteria were observed within the crypt lumina (Fig. 2 inset). Numerous protozoan structures suggestive of amoeba trophozoites were observed free in the necrotic debris, the lamina propria, submucosa, and within lymphatic vessels (Figs. 3, 4). These structures were round, 10–15 μm diameter, with a single nucleus and intracytoplasmic vacuoles. Amoeba-like structures were periodic acid–Schiff (PAS)-positive and Grocott-negative. No fungal structures were observed in the Grocott stain. Scattered Balantidium coli were also seen throughout the intestinal lumen. Based on these results, further microbiologic and molecular investigations were pursued to identify the lesion-associated bacteria and protozoa.
Routine bacterial cultures were attempted for Escherichia coli (blood agar and MacConkey agar) and Salmonella spp. (brain-heart infusion– and Rappaport-Vassiliadis–enriched broths) on samples of ileum and colon, which yielded growth of non-hemolytic E. coli colonies and no growth of Salmonella spp. DNA was extracted from 200 mg of intestinal content (QIAamp DNA stool mini kit, Qiagen, Vienna, Austria). B. hyodysenteriae, B. pilosicoli, and Lawsonia intracellularis DNA were tested by specific PCR methods8,9 on samples of colon contents. B. hyodysenteriae was detected in colon, but no PCR products for B. pilosicoli or L. intracellularis were obtained by PCR.
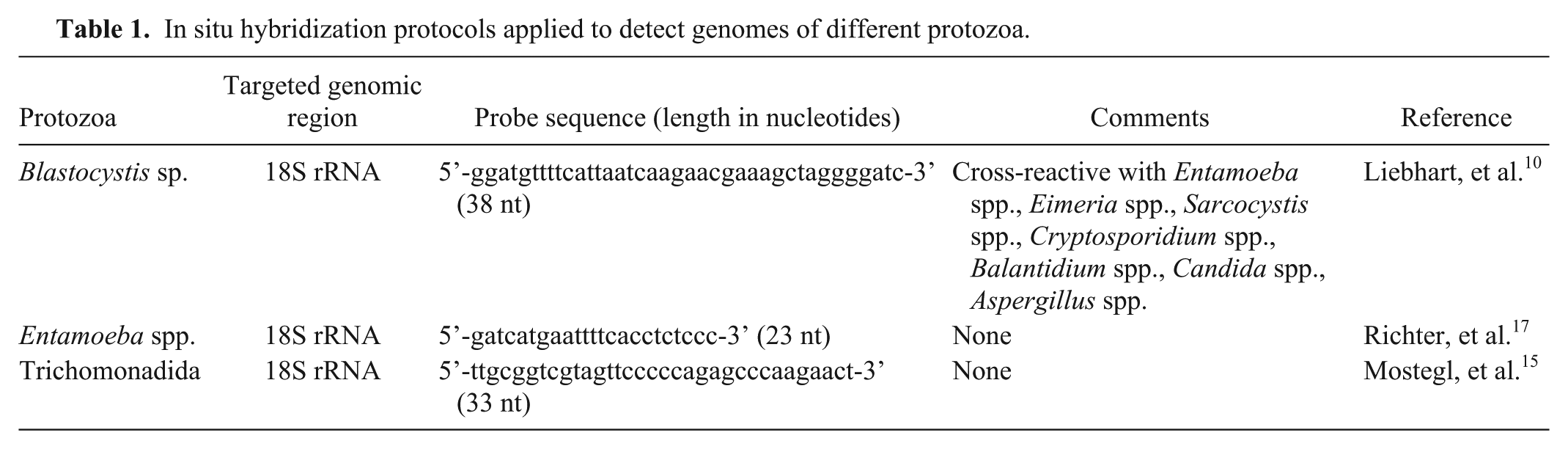
In situ hybridization (ISH) was used to probe for several protozoa (Table 1) on paraffin-embedded intestinal tissue (colon) based on a previously described protocol. 3 Briefly, proteolysis with proteinase K (2.5 μg/mL; Roche, Basel, Switzerland) in Tris-buffered saline was carried out for 30 min at 37°C. For hybridization, slides were incubated overnight at 40°C with hybridization mixture and a final probe concentration of 20 ng/mL for the labeling of Blastocystis sp. and trichomonads, and 10 ng/ml for Entamoeba spp. (Microsynth, Balgach, Switzerland). Digoxigenin-labeled hybrids were labeled with anti-digoxigenin–alkaline phosphatase Fab fragments (1:200; Roche) for 1 h at room temperature. The detection reaction was carried out using the color substrates 5-bromo-4-chloro-3-inodyl phosphate and 4-nitro blue tetrazolium chloride (Roche). Slides were evaluated by light microscopy using semiquantitative scoring. ISH yielded positive signals for all 3 tested protozoa. Large numbers of Entamoeba spp. were predominantly present within the lamina propria and submucosa (Fig. 4, inset), whereas moderate numbers of Blastocystis sp. were exclusively located in superficial necrotic debris and intestinal contents. Scattered trichomonads were confined to crypt lumina.
In situ hybridization protocols applied to detect genomes of different protozoa.

To support the ISH results, PCR to detect Balantidium spp., Blastocystis sp., Entamoeba spp., and Trichomonas spp. was used (Table 2). The PCR reaction master mixture consisted of 12.5 µL of KAPA2G Fast HotStart ready mix with dye (Sigma-Aldrich, Vienna, Austria), 0.4 µM of each primer, 2 µL of template DNA, and distilled water to a total volume of 25 µL per reaction. An aliquot of 10 μL of each PCR product was analyzed by gel electrophoresis using 2% Tris acetate–EDTA–agarose gel. The agarose gel was stained (GelRed nucleic acid gel stain; VWR, Vienna, Austria), and bands were detected (BioSens gel imaging system software; GenXpress, Wiener Neudorf, Austria). PCR products of the expected sizes (Table 2) were evaluated positively. Finally, PCR products were extracted (MinElute PCR purification kit; Qiagen) and were submitted for Sanger DNA sequencing (Microsynth). Nucleotide sequences were analyzed using a BLAST search of the GenBank database.
Primer sequences and temperature conditions of the PCR methods used to detect different protozoan genomes.
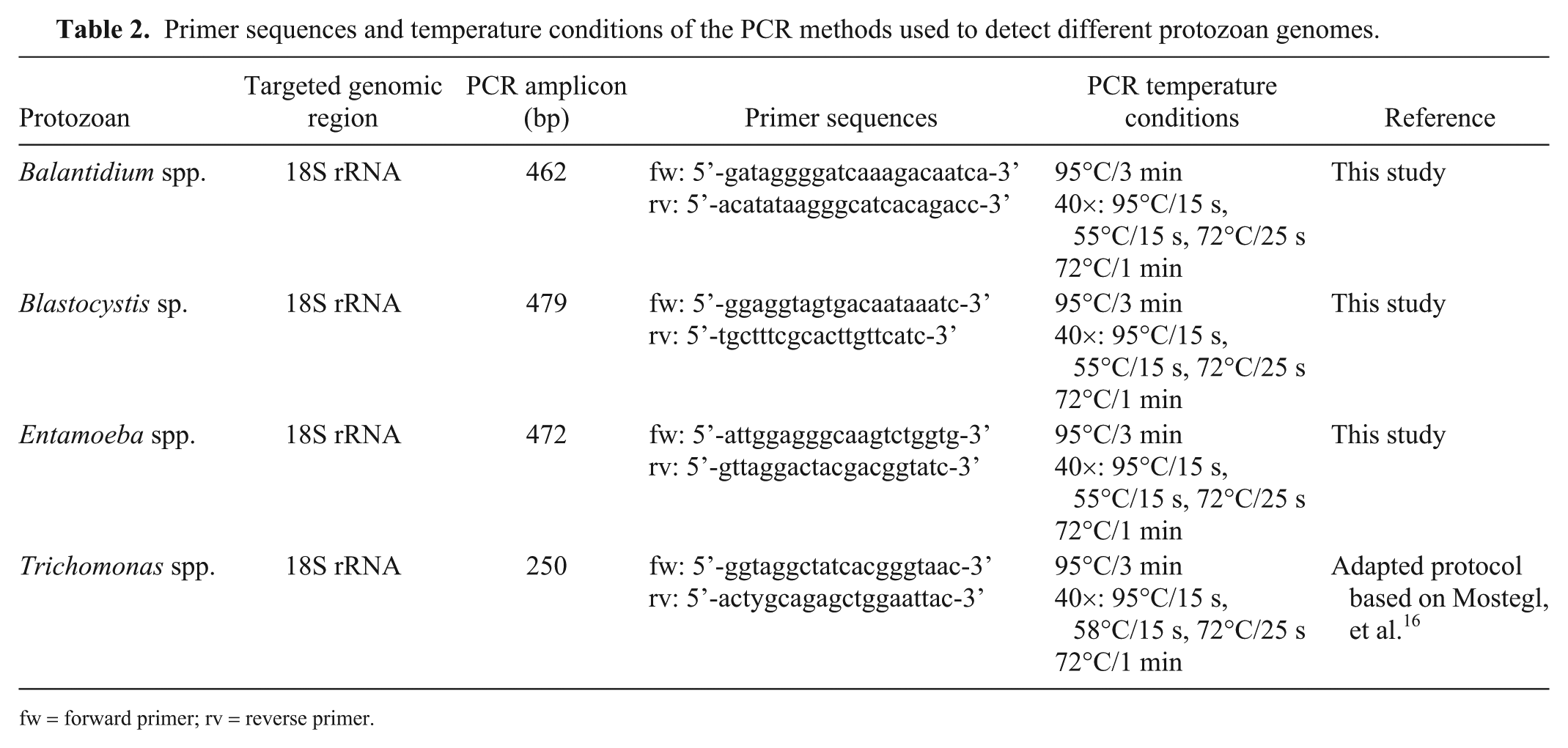
fw = forward primer; rv = reverse primer.
The intestinal content was PCR-positive for Entamoeba spp., Balantidium spp., Blastocystis sp., and Trichomonas spp. Sanger DNA sequencing of the Trichomonas sp. PCR product had 100% identity to the 18S rRNA gene sequence in GenBank (accession JF742057), a sequence of porcine origin with 96–97% similarity to Trichomitus batrachorum. 16 Furthermore, the PCR products had 100% identity to the 18S rRNA gene sequences of Balantidium coli (accession GQ903678), Blastocystis sp. subtype 5 (accession KF410605), and E. polecki (accession MG747668).
To our knowledge, B. hyodysenteriae coinfection with E. polecki associated with fibrinonecrotizing typhlocolitis has not been described previously in the domestic pig. Although other protozoa were found by PCR (Balantidium coli, Trichomonas spp., and Blastocystis sp.), no tissue damage was associated with the presence of trophozoites of these agents. This was in contrast with E. polecki, in which trophozoites were in mucosa, submucosa, and lymphatic vessels of the colon wall, and were associated with severe fibrinonecrotizing inflammation. Importantly, B. hyodysenteriae can also produce necrotizing lesions in large intestine by itself. It is possible that the severity of macroscopic and microscopic lesions observed was the result of the interaction between B. hyodysenteriae and E. polecki. E. polecki was found in the lamina propria and submucosa only in areas of erosion and ulceration, suggesting that it can be an opportunistic pathogen secondary to ulceration. However, the high number of trophozoites may also indicate that E. polecki, under certain circumstances, is able to multiply extensively in tissues and cause severe local damage. Immune suppression may be a contributing factor; however, no significant lesions were observed in lymphoid organs, and 2 well-known immunomodulating viruses—PCV-2 and PRRSV—were not found by immunohistochemistry in the affected animal.
The proliferation of E. polecki may be explained by intestinal dysbiosis caused by B. hyodysenteriae infection. Changes in the intestinal nutrient content caused by inflammatory exudates, mucus, and blood can induce alterations in proportions of microorganisms in the intestinal lumen,1,4,7,8,17,19 including amoebae. 12 Loss of epithelial barrier integrity secondary to B. hyodysenteriae infection may have allowed invasion of E. polecki into the lamina propria, submucosa, and even invasion of lymphatic vessels.
To date, the pathogenicity of E. polecki in domestic pigs has not been fully studied. There are few reports of amoebiasis in the large intestine of pigs. E. polecki subtype 3 and E. suis have been detected in pigs from Japan with colonic ulcerative and hemorrhagic lesions,13,14 respectively. In the case of E. suis, the presence of Brachyspira spp. was ruled out. Interestingly, E. polecki has been identified in the small intestine of a pig with proliferative ileitis caused by L. intracellularis, and it was suggested that coinfection exacerbated the lesions. 12 Amoebae were found in the injured ileum, but not the colon in that case.
Balantidium spp., Blastocystis sp., and Trichomonas spp. were also detected in colonic feces. These protozoa are considered normal intestinal commensals in porcine intestine, and they do not typically cause disease or intestinal lesions. However, host immunosuppression, intestinal dysbiosis, or disruption of the epithelial layer can predispose to overgrowth of these intestinal commensals.1,17,19 The 2 latter possibilities may account for the proliferation of these protozoa, given that the farm used antimicrobials to control the problem (potential dysbiosis associated), and SD is a well-known cause of disruption of the mucosa epithelium. In any case, only B. coli was observed microscopically, and was limited to the colon lumen.
Footnotes
Acknowledgements
We thank Blanca Pérez and Ghizlane El Korchi (Servei de Diagnòstic de Patologia Veterinaria, from the Veterinary School of Barcelona, Spain) for their technical support. We thank Dr. Enric Vidal for his assistance in preparing the images.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
