Abstract
An emerging novel spirochete in swine, provisionally designated “Brachyspira hampsonii,” has been detected worldwide. It has been associated with swine dysentery and cannot be differentiated from B. hyodysenteriae, the classical etiologic agent of this disease, using standard phenotypic methods. We evaluated cross-reactions of “B. hampsonii” isolates recovered from avian species in some of the currently available species-specific polymerase chain reaction (PCR) assays for the identification of swine Brachyspira species. Ten avian “B. hampsonii” isolates recovered from wild waterfowl were used. No false-positive results were recorded with a B. pilosicoli–specific PCR based on the amplification of a fragment of the 16S rRNA gene. However, the percentage of false-positive results varied, with a range of 10–80%, in the evaluated B. hyodysenteriae–specific assays based on the amplification of the 23S rRNA, nox, and tlyA genes. Similarly, results of the B. intermedia–specific PCR assays yielded poor specificity, with up to 80% of the “B. hampsonii” isolates tested giving false-positive results. Finally, 2 “B. hampsonii” avian isolates yielded a positive result in a B. innocens– and B. murdochii–specific PCR. This result should be interpreted very cautiously as these 2 isolates could represent a recombinant genotype.
The bacterial genus Brachyspira includes a number of officially named and unofficially proposed species of spirochetes that colonize the large intestine of different mammals and birds. 12 Seven Brachyspira species have been identified from swine. 7 Brachyspira hyodysenteriae is the etiologic agent of swine dysentery, a well-known disease of pigs characterized, in the classical form, by severe mucohemorrhagic colitis. 7 Brachyspira pilosicoli causes porcine intestinal spirochetosis, a less severe condition that resembles the early stages of swine dysentery. 7 Brachyspira innocens is considered a nonpathogenic species; B. intermedia and B. murdochii are variably associated with diarrheal disease. 7 In 2007, a new species within the genus, “B. suanatina,” was isolated from pigs and mallard ducks, and clinical signs and lesions compatible with swine dysentery were shown in experimentally infected pigs. 18 Similarly, in 2012, atypical Brachyspira isolates were isolated from outbreaks of mucohemorrhagic colitis indistinguishable from swine dysentery in the United States and Canada. Phylogenetic analysis of these isolates showed that they represent a novel species for which the name “B. hampsonii” was proposed. 5 In contrast to “B. suanatina”, this new proposed species has emerged as an important swine pathogen in the United States and Canada, even replacing B. hyodysenteriae as the predominant etiologic agent of swine dysentery. 21 It has also been described in European pigs,11,19 as well as in wild waterfowl in the Canadian Arctic 21 and in Spain. 13
Definitive detection of Brachyspira infection is commonly based on the isolation of the bacteria from colonic tissues or feces using selective media containing blood and antimicrobials incubated under an anaerobic atmosphere. 22 The degree of hemolysis is normally used for presumptive identification; B. hyodysenteriae, “B. suanatina”, and “B. hampsonii” are strongly β-hemolytic, whereas B. pilosicoli, B. innocens, B. intermedia, and B. murdochii are normally weakly β-hemolytic.5,7
Identification of pathogenic Brachyspira species is generally completed using specific molecular methods such as the polymerase chain reaction (PCR) assay, either directly from feces or colonic tissues or from primary cultures. 15 However, false-positive results with “B. hampsonii” isolates in supposedly specific PCR assays for the tlyA gene of B. hyodysenteriae and for the nox and 23S ribosomal RNA (rRNA) genes of B. intermedia have been described. 11 We evaluated cross-reactivity in some of the currently available species-specific PCR assays for identification of swine Brachyspira species using a collection of “B. hampsonii” isolates of avian origin.
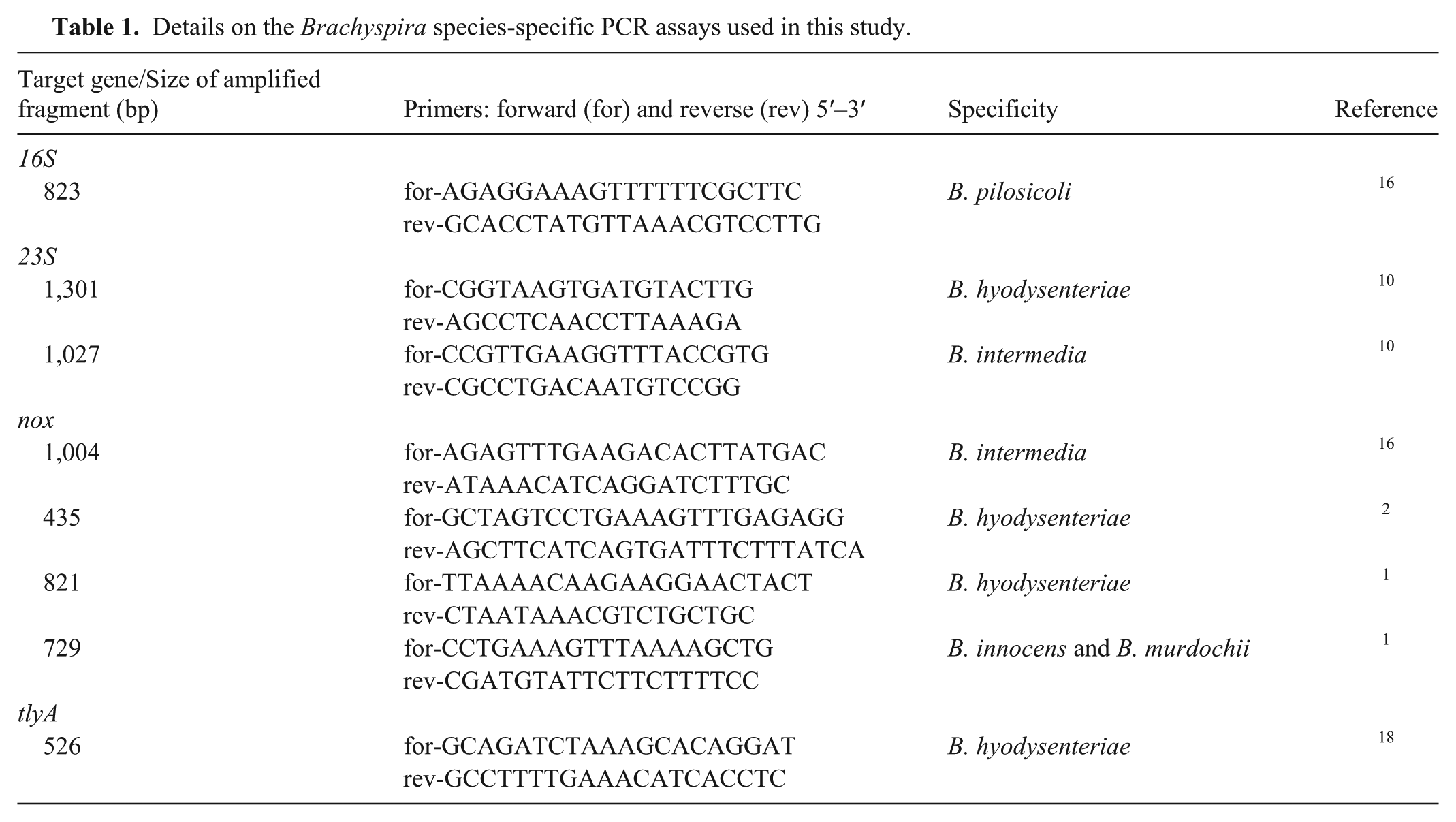
Bacterial isolates used in our study included 10 “B. hampsonii” isolates, the reference strain B204 of B. hyodysenteriae (ATCC 32121), the type strain P43/6/78 of B. pilosicoli (ATCC 51139), and 3 isolates from the bacterial collection held at the University of León identified as B. murdochii, B. intermedia, and B. innocens by PCR amplification and sequencing of the nox gene as described previously. 1 The “B. hampsonii” isolates were recovered from migratory wild waterfowl in Spain, 13 and they are the only isolates reported of this species in Spain to date. Using multilocus sequence typing, 7 of the 10 isolates were classified in 3 of the 4 proposed genetic groups of “B. hampsonii” and showed a close relationship with the swine origin genotypes reported in Europe. 14 Isolates were grown on sheep blood agar plates a at 42°C in an anaerobic atmosphere. b Genomic DNA was obtained from each isolate by boiling as described previously. 13 Eight previously described species-specific PCR assays were evaluated (Table 1). Target genes included nox, 16S rRNA, 23S rRNA, and tylA. Of these targets, the 16S rRNA and 23S rRNA genes are usually recognized as more conserved among different Brachyspira species, whereas the nox gene seems to be less conserved and is the most frequently used target for the differentiation of Brachyspira species.1,22 The PCR assays were performed using a commercial kit c in 50-µL reaction mixtures and always included a positive and a negative control consisting of the specific Brachyspira species and ultrapure water, respectively. Each reaction was performed using primer and PCR conditions described previously.1,2,10,16,18 The PCR products were subjected to agarose gel electrophoresis and visualized by ultraviolet illumination in the presence of a nucleic acid stain. d The PCR reaction was classified, for each sample, as positive (strong band similar to positive control), weakly positive (weak band), or negative.
Details on the Brachyspira species-specific PCR assays used in this study.

The B. hyodysenteriae, B. pilosicoli, B. innocens, B. murdochii, and B. intermedia isolates only yielded positive results in each relevant species-specific PCR (Table 2). However, some false-positive results were obtained when “B. hampsonii” isolates were tested in these species-specific PCR assays. The percentage of “B. hampsonii” isolates that give a false-positive result was 40–80% for the B. intermedia–specific PCRs, 10–80% for the B. hyodysenteriae–specific assays, and 20% for the B. innocens– and B. murdochii–specific PCR. On the other hand, the B. pilosicoli–specific PCR based on the amplification of the 16S rRNA gene 16 did not show any false-positive result when tested using “B. hampsonii” isolates, a finding that has been described previously with other Brachyspira species. 4
Results of Brachyspira species-specific PCR assays, by species and target gene.*
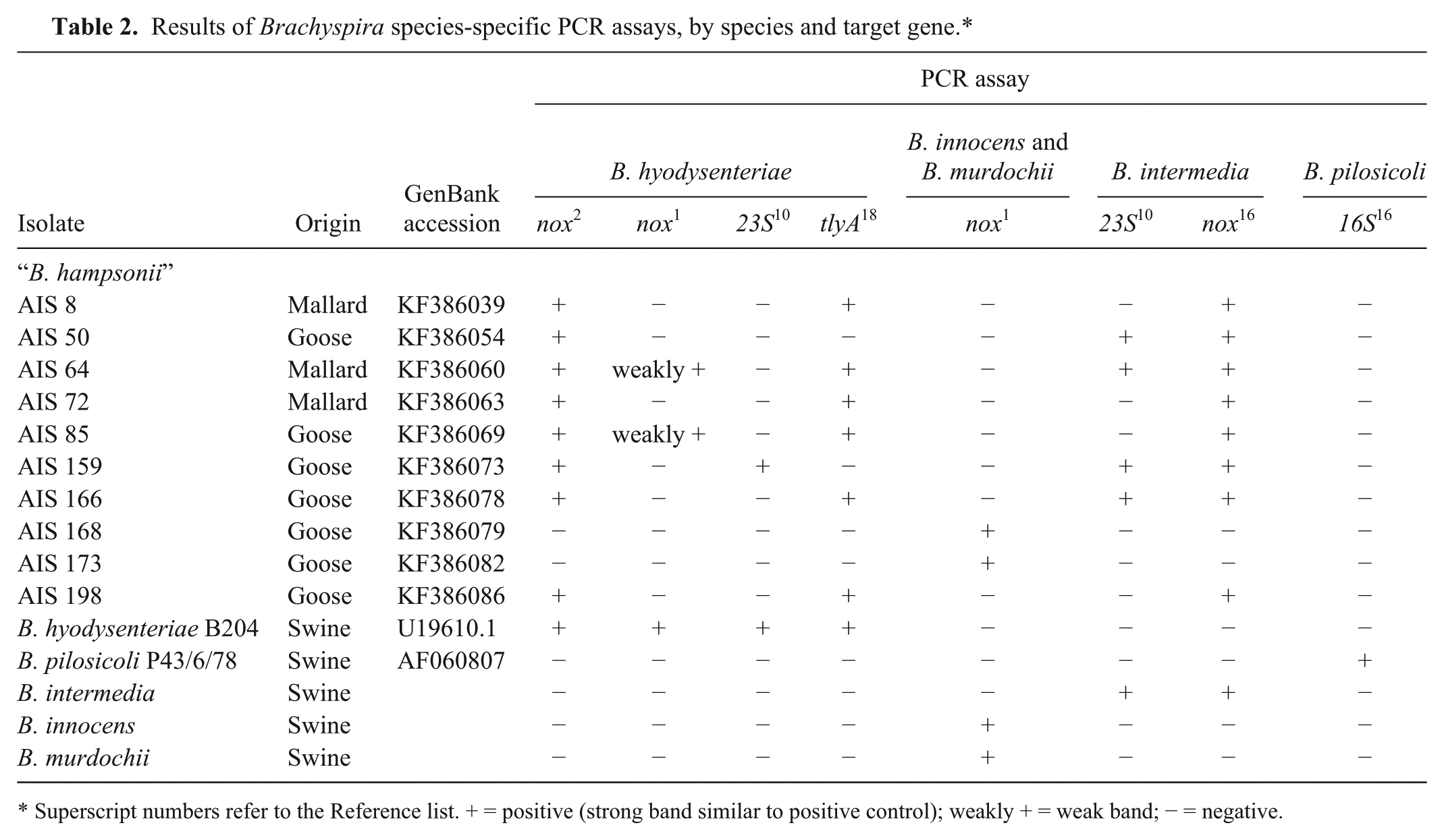
Superscript numbers refer to the Reference list. + = positive (strong band similar to positive control); weakly + = weak band; − = negative.
Since the first description in 2012, “B. hampsonii” has emerged as a relevant swine pathogen in North America and is associated with a disease indistinguishable from classical swine dysentery caused by B. hyodysenteriae.5,7,20 Although “B. hampsonii” has been isolated from Belgian pigs imported to Germany 19 and from 2 gilts brought from the Czech Republic to Belgium, 11 the actual distribution of the species in Europe is unknown. Several methods have been described for the detection or identification of “B. hampsonii”22,23; however, in most laboratories, the diagnosis of swine dysentery is still supported by microbial culture and/or species-specific PCR for the identification of B. hyodysenteriae.22,23 According to our results, some of the most commonly used B. hyodysenteriae–specific PCR assays are most likely to fail when differentiating between B. hyodysenteriae and “B. hampsonii” isolates, which could underestimate the real prevalence of this novel species.
In terms of the B. hyodysenteriae–specific PCR assays used on the “B. hampsonii” isolates, the most specific assay was based on the amplification of a 1,301-bp fragment located on the 23S rRNA gene, 10 and gave only 1 false positive. The next best performance was given by an assay based on the amplification of a 821-bp fragment of the nox gene, 1 with 2 weak false positives. These 2 weak false-positive results remained even when more stringent conditions were used, such as increasing the annealing temperature by 2°C. In contrast, 8 of 10 “B. hampsonii” isolates gave a positive result in the B. hyodysenteriae–specific PCR based on the amplification of a 435-bp fragment of the nox gene. 2 The proportion of false-positive results was also high using a PCR based on the amplification of a 521-bp fragment of the tylA gene. 18 According to our results, some of the evaluated species-specific PCR assays are of little use for the definitive identification of B. hyodysenteriae isolates when used at the conditions originally proposed. A similar cross-reaction was previously described for “B. hampsonii” isolates of porcine origin 11 using the PCR assay based on the amplification of the tlyA gene. False-positive results in B. hyodysenteriae–specific PCR assays based on the amplification of the tlyA and nox genes have also been described for some avian isolates identified as B. alvinipulli. 9
In some laboratories, Brachyspira spp. isolates are further characterized using phenotypic methods. The isolation of Brachyspira spp. from clinical samples is a time-consuming and technically challenging procedure. 7 Furthermore, B. hyodysenteriae and “B. hampsonii” isolates have similar phenotypic profiles. The only phenotypic characteristic that can be used to differentiate the species is the lack of indole production in the case of “B. hampsonii” isolates. 5 However, indole-negative B. hyodysenteriae isolates have been reported.5,8
Cross-reactions with “B. hampsonii” were also very common in the 2 evaluated B. intermedia–specific PCR assays, a problem that has been described previously.3,11,19 These assays were based on the amplification of a 1,027-bp fragment of the 23S rRNA gene 10 (4 of 10 “B. hampsonii” isolates were positive) or a 1,004-bp fragment of the nox gene 16 (8 of 10 of the “B. hampsonii” isolates were positive). Brachyspira intermedia is an extremely diverse indole-positive and weakly hemolytic intestinal spirochete that includes a great variety of sequence types or clusters. 17 It has been reported that B. intermedia–specific PCR based on the amplification of the nox gene fails to detect all strains within the species, and this lack of sensitivity has been attributed to the genetic heterogeneity of this species. 17 The results of our study indicate that available B. intermedia–specific PCR assays should also be optimized in terms of specificity.
Only 2 of the evaluated isolates, identified as “B. hampsonii” AIS 168 and AIS 173, yielded a positive result in the specific B. innocens– and B. murdochii–specific PCR evaluated. However, these results must be considered very carefully. These 2 isolates were classified as “B. hampsonii” based on phenotypic characteristics and on the analysis of partial sequence of the nox gene, but they clustered with B. innocens and B. murdochii when 16S rRNA gene sequence was taken into account. 13 In the first recognition of “B. hampsonii” in wild European waterfowl, genomic rearrangements and recombination events were suggested as the explanation for the lack of correlation between the results obtained using different target genes. 13 These events have been described when different species of spirochetes infect the same individual concurrently,6,18 particularly in avian hosts,9,13 making the identification of Brachyspira species even more difficult.
Misidentification of Brachyspira infections could occur very frequently.11,19 In the United States, it has been reported that >50% of the clinical Brachyspira isolates recovered from pig feces or intestines could not be identified at the species level using current PCR assays.5,6 Despite the low number of “B. hampsonii” isolates evaluated and their common avian origin, our results confirm that some of the currently available PCR assays for the identification of Brachyspira species could yield false-positive results, when used as initially described, given lack of detection of this novel proposed species. Our results suggest that PCR conditions should be carefully revised to increase specificity, and PCR results should be carefully interpreted if the amplified DNA products are not sequenced. In particular, B. hyodysenteriae– and B. intermedia–specific assays should be carefully evaluated for specificity and the inclusion of a “B. hampsonii”–specific PCR together with B. hyodysenteriae– and B. pilosicoli–specific assays is strongly recommended in routine panels for porcine colitis in post-weaning and fattening pigs.
Footnotes
Acknowledgements
We thank Gloria Fernández-Bayón for providing excellent technical assistance.
Authors’ note
Preliminary results were presented as a poster at the 6th European Symposium of Pig Health Management, Sorrento, Italy, 7–9 May, 2014.
Authors’ contributions
LM Aller-Morán contributed to conception and design of the study; contributed to acquisition, analysis, or interpretation of data; and drafted the manuscript. FJ Martínez-Lobo and A Carvajal contributed to conception and design of the study; contributed to acquisition, analysis, or interpretation of data; and critically revised the manuscript. P Rubio contributed to conception and design of the study and critically revised the manuscript. All authors gave final approval and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Oxoid Ltd., Basingstoke, Hampshire, United Kingdom.
b.
Bugbox anaerobic workstation, Ruskinn Technology Ltd., Bridgend, South Wales, United Kingdom.
c.
Invitrogen Platinum hot start PCR master mix, Thermo Fisher Scientific Inc., Madrid, Spain.
d.
GelRed, Biotium Inc., Hayward, CA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was funded by the Spanish Ministry of Economy and Competitiveness (project AGL 2010-18804).
