Abstract
Swine dysentery is characterized by mucohemorrhagic diarrhea and can occur following infection by Brachyspira hyodysenteriae or “Brachyspira hampsonii ”. A definitive diagnosis is often based on the isolation of strongly beta-hemolytic spirochetes from selective culture or by the application of species-specific polymerase chain reaction (PCR) assays directly to feces. While culture is highly sensitive, it typically requires 6 or more days to complete, and PCR, although rapid, can be limited by fecal inhibition. Fluorescent in situ hybridization (FISH) has been described in formalin-fixed tissues; however, completion requires approximately 2 days. Because of the time constraints of available assays, a same-day FISH assay was developed to detect B. hyodysenteriae and “B. hampsonii ” in pig feces using previously described oligonucleotide probes Hyo1210 and Hamp1210 for B. hyodysenteriae and “B. hampsonii”, respectively. In situ hybridization was simultaneously compared with culture and PCR on feces spiked with progressive dilutions of spirochetes to determine the threshold of detection for each assay at 0 and 48 hr. The PCR assay on fresh feces and FISH on formalin-fixed feces had similar levels of detection. Culture was the most sensitive method, detecting the target spirochetes at least 2 log-dilutions less when compared to other assays 48 hr after sample preparation. Fluorescent in situ hybridization also effectively detected both target species in formalin-fixed feces from inoculated pigs as part of a previous experiment. Accordingly, FISH on formalin-fixed feces from clinically affected pigs can provide same-day identification and preliminary speciation of spirochetes associated with swine dysentery in North America.
Introduction
Since 2006, there has been an increase in swine dysentery (SD) diagnoses in growing and finishing pigs in the United States 6 with similar reports from Canada. 11 Swine dysentery is characterized by mucohemorrhagic diarrhea and typhlocolitis and can result in significant economic losses in affected production systems. 8 Classically, SD is associated with infection with Brachyspira hyodysenteriae. 8 However, a recently proposed novel Brachyspira species, “Brachyspira hampsonii”, 5 has been isolated from pigs with mucohemorrhagic diarrhea, and experimental infection with “B. hampsonii” strains has consistently resulted in clinical disease and gross lesions that are similar to, if not indistinguishable from, B. hyodysenteriae infection.3,12,14
A definitive diagnosis of SD is commonly based on the isolation of strongly beta-hemolytic, ring phenomenon–positive spirochetes from culture of mucohemorrhagic feces or colonic tissue and/or by species-specific polymerase chain reaction (PCR) assays run on extracts of such samples. 4 While Brachyspira culture using selective agars is a highly sensitive assay, it can be technically challenging, time consuming, typically requires speciation using PCR following isolation, and often requires 6 days or longer to complete, which can result in a delay in disease diagnosis. 4 Polymerase chain reaction assays, while rapid, can be limited by fecal inhibition, particularly in pigs. 13 Fluorescent in situ hybridization (FISH) assays have been described for the identification of B. hyodysenteriae and “B. hampsonii” in formalin-fixed tissues.2,4 However, these assays require approximately 2 days or more to complete following fresh tissue collection. 4 Because of the time constraints of currently available assays, the current study reports the development of a same-day FISH assay to detect B. hyodysenteriae and “B. hampsonii” in pig feces using previously described oligonucleotide probes Hyo12102 and Hamp12104 targeting the 23S ribosomal RNA (rRNA) of B. hyodysenteriae and “B. hampsonii”, respectively. In the present study, the newly developed FISH assay was simultaneously compared with both culture and real-time PCR (qPCR) on pig feces spiked with progressive dilutions of B. hyodysenteriae and “B. hampsonii” to determine the threshold of detection for each assay at 0 and 48 hr after sample preparation. As well, the assay was used on formalin-fixed feces collected as part of previous experiments. Additionally, the specificity of each probe was tested against Brachyspira murdochii and Brachyspira pilosicoli, which are the most commonly isolated weakly hemolytic Brachyspira spp. at the Iowa State University Veterinary Diagnostic Laboratory (ISU VDL; Ames, Iowa).
Materials and methods
Bacterial strains and growth conditions
Strains used in the current study were obtained from the culture collection at the ISU VDL. The “B. hampsonii” clade II strain (EB107) was originally recovered from a clinical case of SD in 2011 and has been previously used in multiple pig inoculation experiments.3,14 The B. hyodysenteriae strain (B204) was originally recovered from a clinical case of SD in 1972. The B. murdochii (KC63) and B. pilosicoli (P43/6/78) strains were obtained from a clinical case in 2008 and the American Type Culture Collection, respectively.
Strains were plated onto selective agars containing either colistin, vancomycin, and spectinomycin, 9 or pig feces extract, spiramycin, rifampin, vancomycin, colistin, and spectinomycin. 10 Following 4 days of incubation in an anaerobic environment provided by a commercial system a at 41 ± 1°C, strains were transferred to trypticase soy agar with 10% bovine blood (10% TSA) and incubated as described previously. 3 Plate media used in the present study were prepared in-house and passed the quality assurance standards of the ISU VDL.
Preparation of fecal dilutions and standard plate counts
All strains were harvested from 10% TSA plates and rinsed into 0.85% physiological saline to a McFarland 0.5 standard. Tenfold serial dilutions were performed on “B. hampsonii” and B. hyodysenteriae to make 8 dilutions ranging from 10−1 to 10−8. One milliliter of the varying dilutions of each isolate was added to 7 ml of phosphate buffered 0.85% physiological saline, which contained 2 g of homogenized normal and antibiotic-free pig feces. One milliliter of a suspension of B. murdochii and of B. pilosicoli (each adjusted to a McFarland 0.5 standard) was added to 9 ml of phosphate buffered saline (PBS). At 0 and 48 hr after fecal dilution preparation, fecal dilutions of B. hyodysenteriae and “B. hampsonii” were plated onto trypticase soy agar with 5% bovine blood. Plates were incubated as stated above. Plates were observed and, while confluent growth was observed at the concentrated dilutions, discrete colonies were observed on the more dilute plates and were counted to obtain an estimation of the colony-forming units (CFU) per milliliter of each dilution on days 2, 4, and 6 of incubation. The final CFU/ml was calculated when the count was between 30 and 300 CFU/ml. Between the 2 time points, fecal dilutions were stored at 4°C. Isolation of any strongly beta-hemolytic, ring phenomenon–positive spirochetes was considered a positive culture.
Fluorescent in situ hybridization
Custom oligonucleotide probes (Hamp1210 and Hyo1210) were purchased from a commercial source b and were 5′-labeled with orange fluorescent c dye. One milliliter of the varying fecal dilutions of B. hyodysenteriae or “B. hampsonii” was added to 9 ml of PBS and passed through a commercially available filter d (pore size: 200–300 µm) from which 0.5 ml was transferred to 1.5 ml of 10% neutral buffered formalin (NBF) or 1.5 ml of PBS and incubated at 20–25°C for 20 min. Using a hydrophobic marking pen on positively charged slides, a 10–15 mm in diameter circle was drawn, and 20 µl of filtered fecal dilutions in 10% NBF or PBS was placed inside the circle. Half a milliliter of a 1:10 dilution of the McFarland 0.5 standard suspension of B. murdochii and B. pilosicoli was added to 1.5 ml of 10% NBF, incubated at 20–25°C for 20 min, and then a 20-µl aliquot was placed inside a marked circle. Slides were then dried in an incubator at 42°C for 20–30 min. The Hyo1210 and Hamp1210 probes were reconstituted using DNase- and RNase-free water and diluted to a working concentration of 10 ng/μl in hybridization buffer (20 mM of Tris, 0.9 M of NaCl, 20% sodium dodecyl sulfate [SDS] buffer, 40% formamide, 10% dextran sulfate [pH 7.2]). Each of the dried fecal samples of B. hyodysenteriae and “B. hampsonii” and dried PBS samples of B. murdochii and B. pilosicoli were covered with 50 µl of the hybridization solution containing the desired probe and placed in a moisture chamber. Hybridization was carried out for 6 hr at 42°C. Following hybridization, slides were washed in a wash buffer (hybridization buffer without SDS or formamide and prewarmed to the hybridization temperature) for 20 min. After washing, the slides were rinsed with sterile water, allowed to air dry, and mounted using a commercial anti-fade reagent. e A fluorescent microscope with a filter appropriate for orange fluorescent dye c was used for all analyses. Ten 60× high power fields of each circumscribed area were reviewed. The identification of a total of 3 or more spirochetes in the reviewed fields was designated as a positive sample. The most concentrated samples were reviewed first and, after 2 consecutive negative dilutions, additional dilutions were not reviewed. All fecal dilutions were stored at 4°C for 48 hr and processed again in the same manner. Two replicates with each strain were performed.
Real-time polymerase chain reaction primers, probes, and protocol
One milliliter of each filtered fecal dilution in 10% NBF or PBS was submitted at 0 and 48 hr after sample preparation for qPCR. The DNA was extracted from the fecal samples according to manufacturer’s instructions using a commercial extraction kit f and an automated extraction procedure using a commercial instrument. g
The primers (forward: 5′-TTGCTACTGGTTCTTGGCCTG-3′; reverse: 5′-GAATGCTTCTATAAGTTCAACACCTAT-3′) 13 and probes (B. hyodysenteriae: FAM-5′-CGAAGGCTTAAAACAAGAAGGA-3′-IABlkFQ 13 ; “B. hampsonii”: cyanine 5-5′-CGCTAAATTATTCCAACA AGGACAGG-3′-IABkFQ) were obtained from a commercial source. h Real-time PCR was run using a commercially available master mix i and an exogenous internal positive control j (IPC) with a VIC c -labeled probe. Each Brachyspira probe was used separately, meaning a total of 2 reactions per DNA. Briefly, 12.5 µl of the master mix was combined with 1 µl of each primer diluted to 10 µM and 0.5 µl of probe diluted to 10 µM, 2.5 µl of 10× IPC mix, 0.5 µl of 50× IPC DNA, and 4.5 µl of PCR-grade water. The DNA template was added at 2.5 µl per reaction for a total reaction volume of 25 µl. The qPCR was run as follows: 95°C for 10 min, followed by 45 cycles of 95°C for 15 sec and 60°C for 45 sec. Resulting threshold cycle (Ct) values were reported through 45 cycles. A Ct value less than 45 was considered positive.
Preparation of samples from experimentally infected pigs
One milliliter of large intestinal luminal content was collected at necropsy during a previous inoculation trial 14 (B. hyodysenteriae (B204): n = 10; “B. hampsonii” (EB107): n = 10) and placed in 9 ml of brain heart infusion broth from which 0.5 ml was added to 1.5 ml of 10% NBF and stored at −80°C for 6 months. At time of collection, sample consistency varied from semisolid (clinically unaffected pigs) to liquid (clinically affected pigs). Samples were thawed and passed through a commercially available filter. d A 20-µl aliquot of the resulting filtrate was placed onto positively charged slides, and FISH was performed as described above.
Mucohemorrhagic feces were collected from the cement floor of pens containing experimentally infected pigs during a previous inoculation trial (B. hyodysenteriae (B204): n = 5; “B. hampsonii” (EB107): n = 3). 15 A 1-ml sample of dysenteric feces was placed in 9 ml of 10% NBF. On the day of collection, samples were passed through a commercially available filter. d The filtrate was divided and stored at either −80°C or 20–25°C. Fluorescent in situ hybridization was run as described above with a shortened hybridization time (3–4 hr). Samples stored at 20–25°C were run at 24–96 hr after fixation.
Results
Culture
The culture results of the varying dilutions at the 0- and 48-hr time points for all replicates are presented in Table 1. Briefly, culture was positive at dilutions of 10−1 to 10−7 immediately after sample preparation. After 48 hr of refrigerated storage, cultures from these same samples were positive at dilutions of 10−1 to 10−4, and there was a consistent loss of 2–3 log-dilutions between the 48-hr time point and time 0 for all replicates.
Summary of culture, fluorescent in situ hybridization (FISH), and real-time polymerase chain reaction (qPCR) results for each strain and replicate at 0 and 48 hr after sample preparation.*
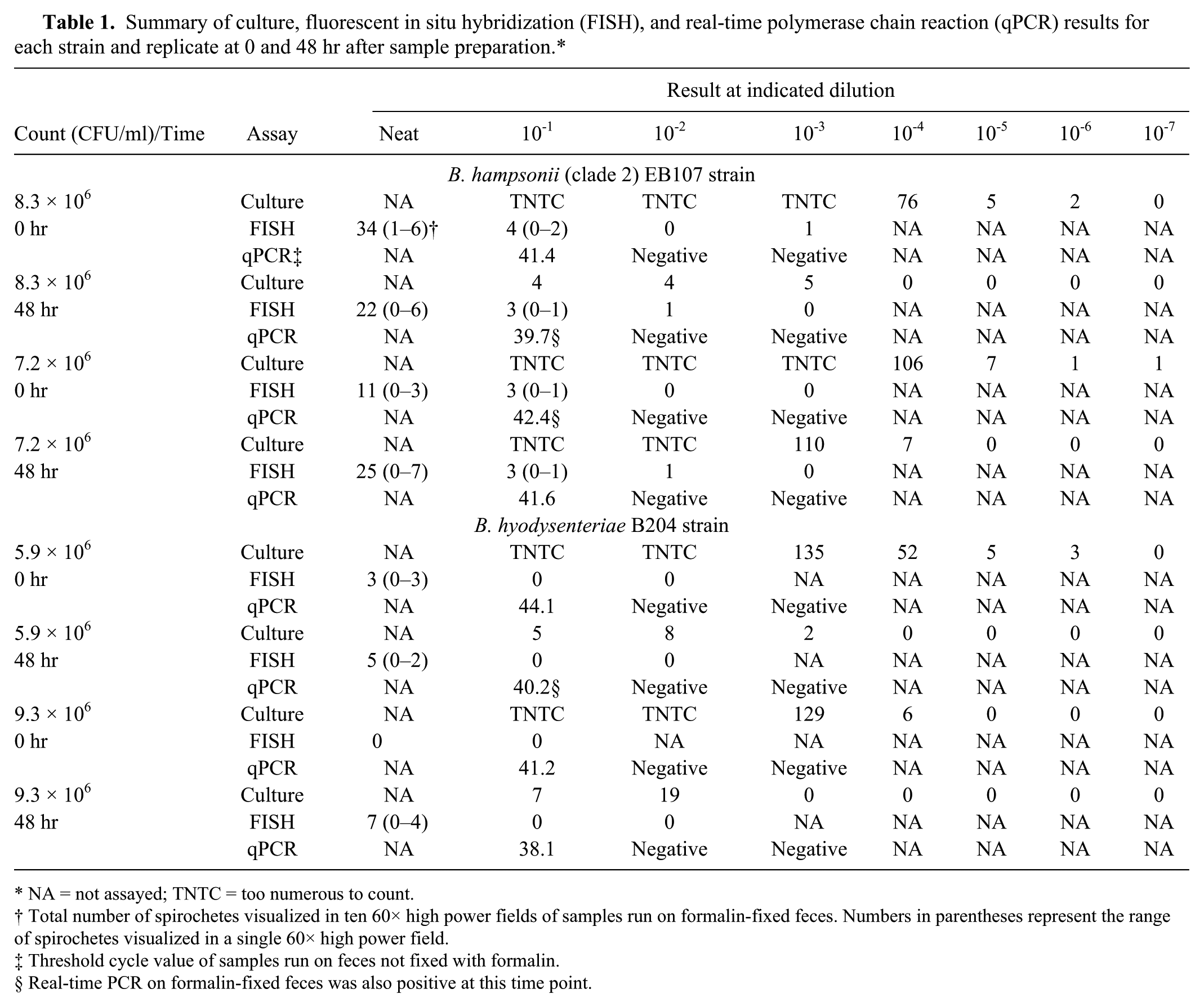
NA = not assayed; TNTC = too numerous to count.
Total number of spirochetes visualized in ten 60× high power fields of samples run on formalin-fixed feces. Numbers in parentheses represent the range of spirochetes visualized in a single 60× high power field.
Threshold cycle value of samples run on feces not fixed with formalin.
Real-time PCR on formalin-fixed feces was also positive at this time point.
Fluorescent in situ hybridization
The FISH results on formalin-fixed feces of the varying dilutions at the 0- and 48-hr time points of all replicates are presented in Table 1. Figure 1 is a representative image of strong positive signal labeling spirochetes using a probe specific for “B. hampsonii”. Detection of 3 or more spirochetes was only observed in the formalin-fixed samples. At both 0 and 48 hr after sample preparation for replicate 1 and 2 of “B. hampsonii”, FISH was positive at the 2 most concentrated dilutions with a threshold of detection of 7.2 × 105 CFU/ml. The threshold of detection of FISH at 48 hr after sample preparation was 5.9 × 106 CFU/ml in replicate 1 of B. hyodysenteriae. Additionally, FISH failed to detect any positively labeled spirochetes at 0 hr after sample preparation for replicate 2 of B. hyodysenteriae, but was positive at 48 hr. A positive signal was not identified in the circumscribed area after hybridization of B. murdochii and B. pilosicoli with either the Hyo1210 or Hamp1210 probe.
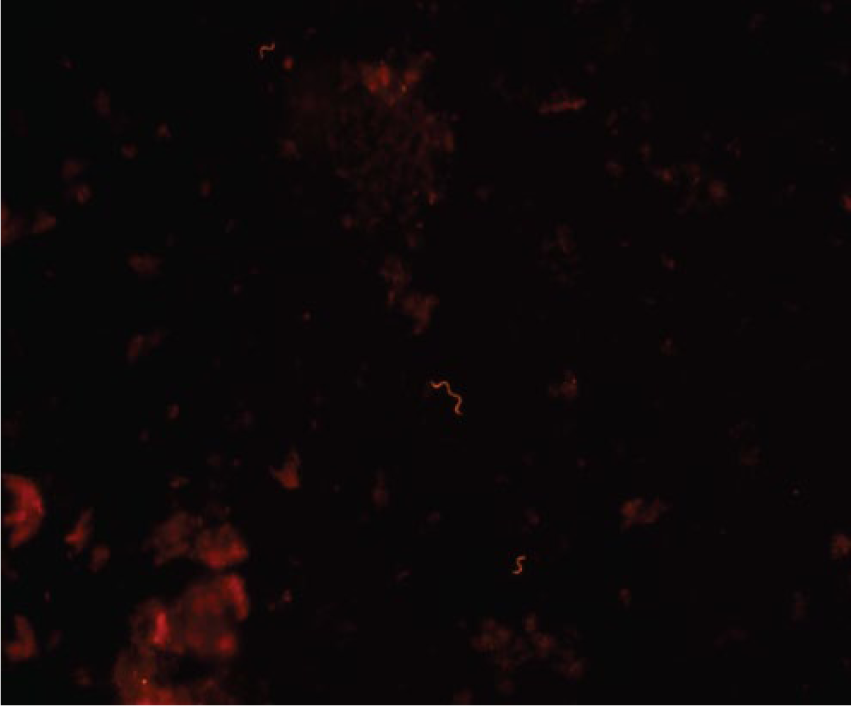
Photomicrograph of a formalin-fixed pig fecal sample spiked with “Brachyspira hampsonii” (strain EB107) 0 hr after sample preparation (replicate 2) and after fluorescent in situ hybridization using an orange fluorescent probe (Hamp1210) specific for “B. hampsonii”.
Real-time polymerase chain reaction
The Ct values of qPCR on feces not fixed in formalin of the varying dilutions at the 0- and 48-hr time points of all replicates are presented in Table 1. The IPC showed little fluctuation in Ct value indicating minimal qPCR inhibition (Ct: 28.7–30.7). A total of 11 positive qPCR results were obtained, 3 of which were from the formalin-fixed samples. Real-time PCR was consistently positive at the first fecal dilution for both spirochetes and at both time points, with the lowest level of detection for qPCR being 5.9 × 105 CFU/ml at 48 hr after sample preparation with B. hyodysenteriae.
Fluorescent in situ hybridization: samples from experimentally infected pigs
Fluorescent in situ hybridization run on large intestinal content samples collected as part of a previous experimental inoculation trial was positive for all culture-positive, qPCR-positive, and clinically affected pigs at necropsy inoculated with either B. hyodysenteriae (n = 9) or “B. hampsonii” (n = 5). However, FISH was negative on samples from 2 inoculated but clinically unaffected pigs, one of which was culture positive but qPCR negative and the other of which was culture negative but qPCR positive. Fluorescent in situ hybridization was also negative on samples from all other culture-negative pigs at necropsy.
A representative image of a pen-level fecal sample with strong positive signal labeling spirochetes using a probe specific for “B. hampsonii” is displayed in Figure 2. Of the 5 samples collected from the pen of a B. hyodysenteriae–inoculated group, 3 were positive when fixed in formalin for 48 hr or less. The 2 samples that were fixed for greater than 48 hr were negative. One sample that was positive at 48-hr fixation was rerun at 72-hr fixation and was negative, while a formalin-fixed frozen aliquot of the same sample remained positive. Similarly, of the 3 samples collected from the pen of a “B. hampsonii”–inoculated group, 2 were positive when fixed in formalin for 48 hr or less. The single sample that was fixed for greater than 48 hr was negative.
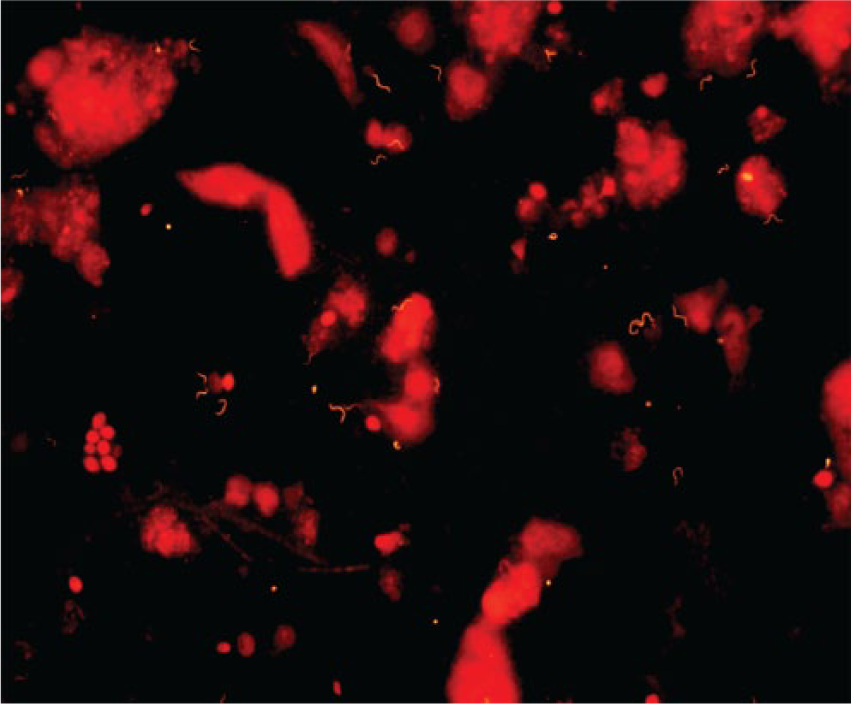
Photomicrograph of a formalin-fixed pig fecal sample collected from the pen of pigs inoculated with “Brachyspira hampsonii” (strain EB107) 24 hr after formalin fixation and following 4 hr of in situ hybridization using an orange fluorescent probe (Hamp1210) specific for “B. hampsonii”.
Discussion
There has been an increase in SD diagnoses associated with infection with B. hyodysenteriae or “B. hampsonii” in growing and finishing pigs in the United States since 2006 6 with similar reports from Canada since 2009. 11 Because of the time constraints and limitations of currently available assays, the current study has developed a same-day FISH assay to detect B. hyodysenteriae and “B. hampsonii” in pig feces using previously described oligonucleotide probes Hyo1210 and Hamp1210 targeting 23S rRNA of B. hyodysenteriae and “B. hampsonii”, respectively.
In the present study, the sample type greatly impacted the level of detection. Real-time PCR on fresh feces and FISH on formalin-fixed feces had similar levels of detection at both time points for both target spirochetes. Fluorescent in situ hybridization failed when used on fresh feces. Formalin fixation is commonly used in FISH to preserve the integrity of cells, prevent cell loss, and preclude nucleic acid degradation. 1 Yet, qPCR run on formalin-fixed samples was rarely positive. Although beneficial for FISH, formalin fixation is likely to inhibit primer access during PCR through the formation of cross-links. 7 Accordingly, formalin fixation of feces when done for 48 hr or less, while the most appropriate sample for FISH, is not suitable for the qPCR assay used in the current study.
Culture was the most sensitive method overall in the present study, commonly detecting both B. hyodysenteriae and “B. hampsonii” at less than 10 CFU/ml immediately after sample preparation. This is consistent with previous studies in which qPCR of rectal swabs 11 or mucosal scrapings obtained at necropsy 3 detected fewer positive samples than selective culture. However, a significant reduction in the number of culturable spirochetes was observed following 48 hr of storage of fecal dilutions at 4°C. Testing samples following 48 hr of storage was selected to replicate the average transport time (time between sample collection and laboratory receipt) for clinical diagnostic samples shipped in the United States and received by the ISU VDL. Although the current study was not a direct representation of laboratory submissions, the data suggests that culture is the best assay currently available for surveillance testing and identification of carrier animals; yet, diagnostic samples should be promptly processed and delivered to the laboratory to decrease the likelihood of false-negative cultures due to prolonged transit time.
Fluorescent in situ hybridization effectively detected the target Brachyspira spp. in formalin-fixed feces from infected pigs with clinical dysentery. Labeled spirochetes were also readily detectable in pen-level samples from infected groups of pigs with clinical disease. While culture remains the diagnostic assay of choice for surveillance situations where low numbers of spirochetes are anticipated, clinically affected pigs may be identified in a timelier manner using either qPCR or FISH. Although a positive signal was not identified after hybridization of B. murdochii and B. pilosicoli with either probe, it bears noting that cross-reactivity with the Hamp1210 probe and Brachyspira intermedia in FISH on formalin-fixed tissues has been observed. 4 However, given the relatively high threshold of detection for FISH on formalin-fixed feces in the current experiment, it seems unlikely that a clinically insignificant infection would be detected by this method. The diagnostic specificity of FISH on formalin-fixed feces under field conditions will become more apparent once it has been adapted into routine use. Accordingly, FISH performed on feces from clinically affected pigs and fixed in formalin for 48 hr or less will allow for same-day identification and preliminary speciation of spirochetes commonly associated with SD in North America. Concurrent culture for confirmation of hemolytic phenotype is recommended where available, and speciation from culture by molecular methods remains a more definitive assay.
Footnotes
a.
BD GasPak EZ anaerobe container system, BD Biosciences, Sparks, MD.
b.
Invitrogen custom oligos, Life Technologies, Carlsbad, CA.
c.
Alexa Fluor 555, VIC; Life Technologies, Carlsbad, CA.
d.
Fisherbrand disposable filter columns, Thermo Fisher Scientific, Waltham, MA.
e.
Invitrogen ProLong Gold, Life Technologies, Carlsbad, CA.
f.
Ambion MagMAX total nucleic acid isolation kit, Life Technologies, Grand Island, NY.
g.
KingFisher, Thermo Fisher Scientific, Waltham, MA.
h.
TaqMan, IDT Inc., Coralville, IA.
i.
Applied Biosystems VetMAX-Plus qPCR master mix, Life Technologies, Grand Island, NY.
j.
Applied Biosystems TaqMan, Life Technologies, Grand Island, NY.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Boehringer Ingelheim Vetmedica Professorship in Food Animal Infectious Disease.
