Abstract
Avian intestinal spirochetosis (AIS), an important but neglected disease in laying hens, is caused by Brachyspira pilosicoli, B. intermedia, and B. alvinipulli. Poultry are also frequently colonized by putatively nonpathogenic species such as B. murdochii and B. innocens. We evaluated the differentiation of Brachyspira species by 3 methods: sequencing of the reduced nicotinamide adenine dinucleotide (NADH) oxidase gene (nox), matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS), and a new multiplex (m)PCR targeting genes such as the tryptophanase A gene (tnaA) and the p-aminobenzoyl-glutamate hydrolase subunit B gene (abgB). Sequencing of 414 bp of the nox PCR amplification products generated from 41 pure cultures of avian Brachyspira isolates allowed presumptive species identification in 33 isolates with at least 99% identity in basic local alignment search tool analysis, including B. pilosicoli, B. intermedia, B. murdochii, B. innocens, and “B. pulli”. MALDI-TOF MS analysis was found to be a reliable tool for differentiation after extension of the manufacturer’s database. In the mPCR, all isolates identified as B. pilosicoli and B. intermedia were positive for abgB and tnaA, respectively. The mPCR might be very useful in detecting Brachyspira species in mixed cultures including not only nonpathogenic species, such as B. innocens, but also one of the AIS pathogens. We found that MALDI-TOF MS analysis combined with the mPCR targeting tnaA and abgB was suitable for the identification of avian isolates of B. pilosicoli and B. intermedia, 2 important agents of AIS.
Keywords
Introduction
Avian intestinal spirochetosis (AIS) is an important disease in poultry, affecting mainly layer flocks but also breeders.11,12,34,35 The disease has been reported in various countries around the world.1,11,25,35 In the UK, high economic losses in the laying hen industry have been attributed to AIS. 18
Typical clinical signs of AIS are diarrhea and reduced egg production. Fecal staining of eggs might occur. 33 Brachyspira pilosicoli and B. intermedia are the main causative agents of AIS.7,17,34 Further species of the genus have been isolated from poultry, but their role in AIS is less clear. B. innocens, B. murdochii, and “B. pulli” are generally presumed to be nonpathogenic species, but B. alvinipulli is also considered an avian pathogen.30,36 B. hyodysenteriae, an important porcine pathogen, has been detected in poultry in a few cases. 7 B. hampsonii and B. suanatina are new species that are pathogenic for pigs but also associated with waterfowl.19,20,22
Pathologic changes have been induced experimentally in poultry by application of B. pilosicoli, B. intermedia, and, in one study, B. alvinipulli.6,12,34,36 Pathologic alterations recorded in experimentally infected chickens are generally mild or moderate and include lymphoplasmacytic typhlocolitis. 36
Numerous methods have been used to differentiate Brachyspira species, including biochemical tests as well as various PCR-based methods, sequencing of the amplification product of the reduced nicotinamide adenine dinucleotide (NADH) oxidase gene (nox), and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS).2,27,37 Hippurate hydrolysis is a typical reaction recorded in B. pilosicoli, and it has also been described for B. alvinipulli.2,9,15 Importantly, there is no generally accepted gold standard method for differentiation of Brachyspira species. Clear criteria for the identification of B. alvinipulli and “B. pulli” have not been published, to our knowledge. Furthermore, differentiation of B. murdochii and B. innocens is known to be difficult. 27 Thus, we sought to improve and evaluate identification tools for the differentiation of Brachyspira isolates from poultry. We compared differentiation by nox sequencing, MALDI-TOF MS, and a newly developed multiplex (m)PCR.
Materials and methods
Isolate collection
From October 2014 until March 2016, cloacal swabs (n = 162) of laying hens in 27 apparently healthy commercial flocks with different housing systems (floor housing, free-range housing, and organic farming) were collected. Furthermore, swabs from the cecum of breeding chickens (n = 29) in 5 flocks with floor housing systems were investigated. The poultry farms ranged from 225 to 70,000 (mean: 16,443) hens, 25–77 wk old. Samples were held on ice and transported in Amies medium within 6 h to the laboratory.
Culture for Brachyspira spp
For evaluation of cultural detection of Brachyspira, fecal samples from poultry were contaminated arbitrarily with different Brachyspira strains (B. murdochii C301, B. intermedia AN26/93, B. pilosicoli 102/06, B. hyodysenteriae 404/06). These samples were comparatively cultured on trypticase soy agar (TSA) containing 10% bovine blood and 3 different antibiotic supplementations: 1) 6.25 µg/mL of vancomycin, 6.25 µg/mL of colistin, 12.5 µg/mL of rifampicin, 25 µg/mL of spiramycin, and 200 µg/mL of spectinomycin; 2) 6.25 µg/mL of colistin, 12.5 µg/mL of rifampicin, and 200 µg/mL of spectinomycin; or 3) no antibiotic supplementation. The first combination of 5 antibiotics is frequently used as selective medium for isolation of Brachyspira from porcine feces. 5 Plates were incubated at 37°C and at 42°C under anaerobic conditions for 7 d. For evaluation, the extension of swarming of the Brachyspira colonies and the separation from colonies of other bacteria were determined.
The samples collected from the different flocks in our study were all cultured on TSA containing 10% bovine blood, 6.25 µg/mL of colistin, 12.5 µg/mL of rifampicin, and 200 µg/mL of spectinomycin at 37°C under anaerobic conditions for 7 d. Primary cultures were screened for colonies typical for Brachyspira. In positive cases, colonies were subcultured onto Columbia blood agar (Oxoid, Wesel, Germany) under anaerobic conditions for the next 7 d. In negative cases, incubation of the primary cultures was continued for another 5 d. Gram stains of smears were conducted at every step. If macro- or microscopic inspection indicated mixed cultures, subculturing was repeated. If the inspection indicated pure cultures, DNA was extracted as described below and the isolates stored at −80°C in brain–heart infusion broth (Oxoid) containing 10% glycerol and 10% calf serum. Isolates identified as B. murdochii or B. innocens in MALDI-TOF MS analysis, but generating a tryptophanase A gene (tnaA) or p-aminobenzoyl-glutamate hydrolase subunit B gene (abgB) amplification product, were streaked out on fastidious anaerobe agar (FAA; LabM, Lancashire, United Kingdom) supplemented with 5% horse blood to induce single colony formation. Subsequently, 3 single colonies of each original isolate were subcultured and investigated again using MALDI-TOF MS and the new mPCR targeting tnaA, abgB, and nox.
Reference strains and sequencing
We included 10 reference strains representing 8 species in our study (Table 1). A pure culture on Columbia agar was suspended in 1 mL of phosphate-buffered saline (PBS; pH 7.2). DNA was isolated (DNeasy blood and tissue kit, Qiagen, Hilden, Germany) as recommended by the manufacturer. The nox PCR was conducted as described. 29 The PCR program included 30 cycles of amplification. DNA was initially denatured at 94°C for 3 min. Each cycle involved denaturation at 94°C for 30 s, annealing at 59°C for 40 s, and extension at 72°C for 54 s. The amplification was finished after a final extension step at 72°C for 10 min. A 10-μL aliquot of the PCR reaction was loaded on 1% agarose gel for electrophoretic separation of DNA that was then stained (Roti-GelStain, Carl Roth, Karlsruhe, Germany). Every nox amplicon was purified (innuPREP DOUBLEpure kit, Analytik Jena, Germany) as recommended by the manufacturer. Nucleotides 512–926 of the amplified nox gene were determined by Sanger sequencing using the oligonucleotide primers described previously. 29 Nucleotide one was defined as the A in the start codon of the nox gene of B. pilosicoli published under accession CP003490. The quality of the sequence was verified visually. Sequencing was repeated in the case of low quality using the same primers or 676-695nox_rev (TTAACTCTGTCATCACCTTC) and 805-826nox_for (GGTGCTATTGTAGTTGATACTA). All nox sequences obtained in our study were submitted to GenBank (KY216077–KY216109, KY196395–KY196402). After completing sequencing for nucleotides 512–926 of the nox gene, National Center for Biotechnology Information (NCBI) basic local alignment search tool analysis (BLAST) analysis (https://www.ncbi.nlm.nih.gov/BLAST/) was conducted. Identities ≥99% were considered a likely species identification.
Brachyspira reference strains included in the current study.
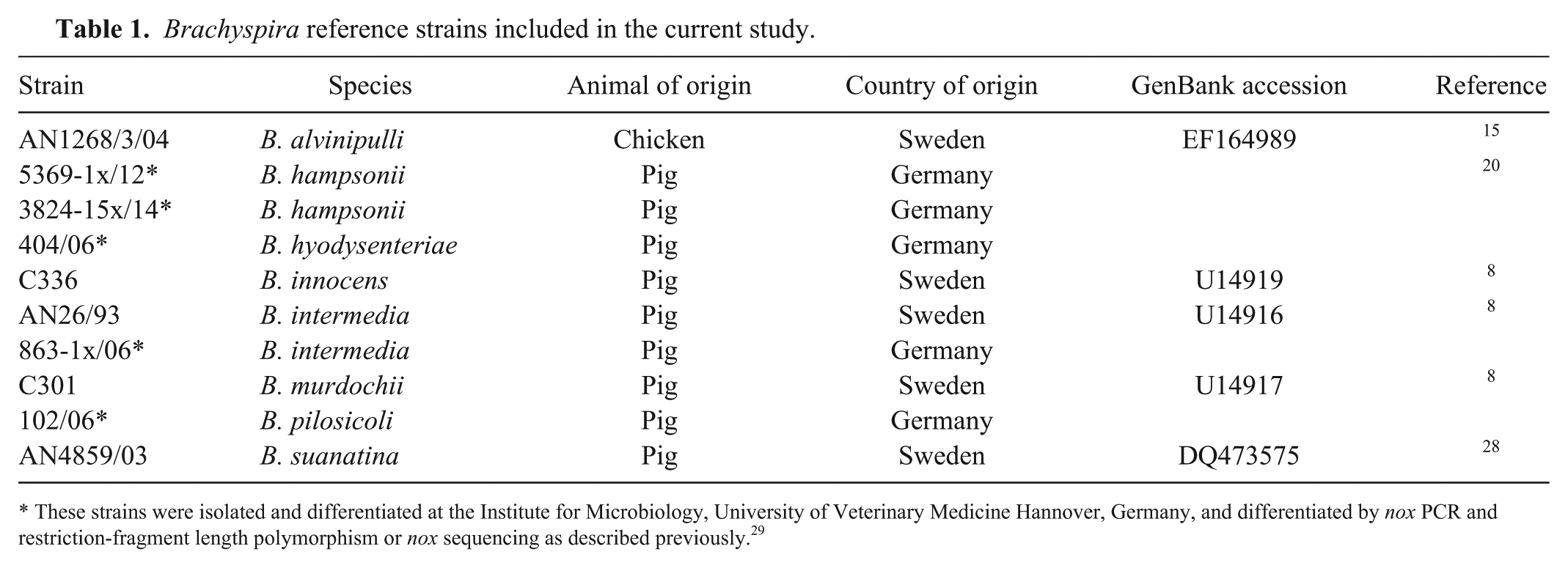
These strains were isolated and differentiated at the Institute for Microbiology, University of Veterinary Medicine Hannover, Germany, and differentiated by nox PCR and restriction-fragment length polymorphism or nox sequencing as described previously. 29
MALDI-TOF MS
A MALDI-TOF MS (Microflex LT, Bruker Daltonics, Bremen, Germany) was used with the following protocol for sample preparation. A total of 900 μL of ethanol was added to 400 µL of a Brachyspira suspension in PBS. The mixture was centrifuged at 14,560 × g for 2 min to remove the supernatant. The pellet was dried at room temperature and resuspended in 40 µL of 70% formic acid. A 40-µL aliquot of acetonitrile was carefully added and, after mixing, the sample was centrifuged at 14,560 × g for 2 min. A 1-μL aliquot of the supernatant of each strain or isolate was spotted 12 times onto a main spectrum profile (MSP) 96 polished steel target plate. Directly after drying, every spot was overlaid with 1 μL matrix. The matrix is composed of a saturated solution of α-cyano-4-hydroxycinnamic acid (Bruker Daltonics) dissolved in 47.5% water (Bioscience), 50% acetonitrile, and 2.5% trifluoroacetic acid.
The commercial database (Bruker Daltonics) included only 4 Brachyspira species with 11 entries in total. We extended the database by generating MSP using reference strains and strains with unambiguous results of species determination by nox sequencing and MALDI-TOF MS analysis.
Each time a database addition was performed, the MALDI-TOF MS was calibrated (Bruker Bacterial Test Standard, Bruker Daltonics) according to the manufacturer’s recommendation. Criteria for including the spectrum of a specific spot in the generation of a main spectrum in the database were numbers of peaks, peak height, and peak deviation as recommended by the manufacturer (quantification by the software). Furthermore, spectra were evaluated visually. A single spectrum was only used for extension of the database if the background noise was very low and the maximal intensities as well as the number of peaks were comparable to the other spectra included for this strain or isolate (very high quality spectrum as shown in Supplementary Fig. 1).
New mPCR for Brachyspira
We designed a new mPCR for differentiation of Brachyspira. Primers were constructed to detect genes encoding enzymes putatively involved in distinct biochemical phenotypes. A 325-bp amplicon of the tnaA gene was obtained with the oligonucleotide primer pair tnaAfor (GTAAAATGTCTTAATATAGGAGGCT) and tnaArev (GACATGCTTTATTTGTAGATGCTAA). A gene putatively involved in hippurate hydrolysis was identified by similarity searches blasting the gene encoding the hippurate hydrolase of Campylobacter jejuni 32 and designated accordingly as abgB gene. The oligonucleotide primer pair abgBfor (GGTAATATGCATGCTTGCGGAC) and abgBrev (TCCAGGCATTTTATGCTGATATTC) was designed based on the in silico–identified abgB genes of B. alvinipulli and B. pilosicoli (see WP_028330113.1 for B. alvinipulli and CP002873.1 for B. pilosicoli). The abgB amplification product is expected to have a size of 744 bp. To detect nox, tnaA, and abgB in a new mPCR, 2 µL of template (~50 ng DNA) was added to 48 µL of a PCR pre-mixture (leading to final concentrations of 2 µM for each forward and reverse primer), 1.5 mM MgCl2 (Invitrogen, Karlsruhe, Germany), Taq reaction buffer (20 mM Tris, pH 8.4, 50 mM KCl; Invitrogen), 0.2 mM of each deoxynucleoside triphosphate (Invitrogen), and 1 unit of Taq polymerase (Invitrogen). After initial denaturation at 94°C for 3 min, 30 cycles of denaturation at 94°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 75 s were run. The amplification was finished after a final extension step at 72°C for 5 min. Visualization of amplification products was conducted as described above.
Results
Optimization of cultural detection of Brachyspira spp. in fecal samples from poultry
Prior to our investigation of samples from the field, we evaluated culture-based detection of Brachyspira in fecal samples from poultry. A selective medium recommended for detection of Brachyspira in samples from pigs, namely TSA with 10% bovine blood and 5 supplemented antibiotics, did not allow isolation of Brachyspira in many samples from poultry artificially contaminated with different Brachyspira strains. Re-isolation of Brachyspira was most successful by culturing avian feces on TSA with 10% bovine blood supplemented with 6.25 µg/mL of colistin, 12.5 µg/mL of rifampicin, and 200 µg/mL of spectinomycin at 37°C.
Differentiation of avian Brachyspira isolates by nox sequencing
Gram-negative, spiral-shaped bacteria with typical swarming colony morphology were isolated from 45 of the 191 samples (162 cloacal and 29 cecal swabs) from poultry investigated. Forty-four of the 45 isolates generated a nox PCR amplification product (singleplex PCR) and were thus included in nox sequencing. In 41 cases, the sequence from base pair (bp) 512–926 was determined in high quality. For the 3 remaining amplicons (isolates N5.6, N10.2, W1.4), only sequences with poor quality were obtained in numerous repetitions. The species were determined for the isolates with the 41 validated sequences by blasting nucleotides 512–826 against the respective NCBI database. The nox sequence (position 512–926) used for species determination was mainly defined by the region of high-quality sequences in all amplicons using the same primers as for amplification. BLAST analysis of nox sequences 512–926 identified 19 isolates as B. murdochii (all 100% identity), 4 isolates as B. innocens (3 with 100% and 1 with 93% identity), 5 isolates as “B. pulli” (all 99% or 100% identity), 3 isolates as B. intermedia (all 99% identity), and another 3 as B. pilosicoli (all 100% identity). In 7 cases, no clear species designation was possible (Table 2). Thus, BLAST analysis generated a species designation with an identity of 99% or 100% in 33 of 41 investigated avian isolates with a high-quality nox sequence (bp 512–926).
Comparative differentiation of 39 avian Brachyspira isolates by nox sequencing and mass spectrometry.
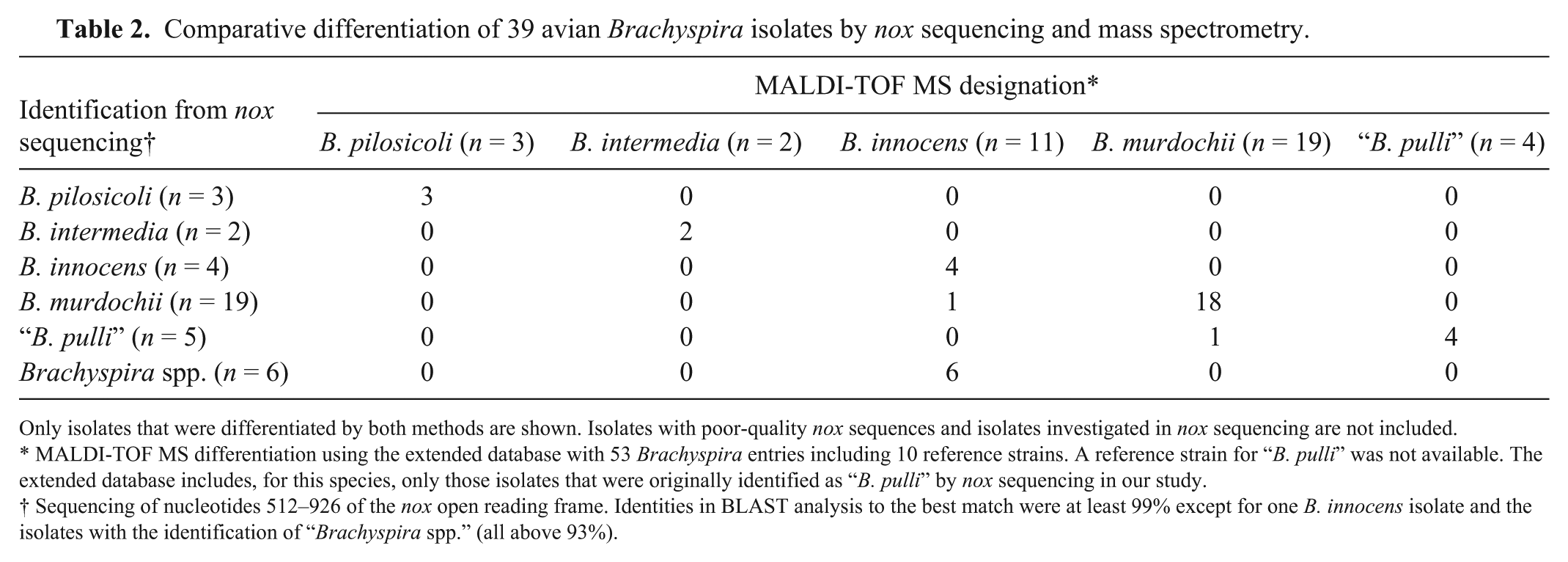

Only isolates that were differentiated by both methods are shown. Isolates with poor-quality nox sequences and isolates investigated in nox sequencing are not included.
MALDI-TOF MS differentiation using the extended database with 53 Brachyspira entries including 10 reference strains. A reference strain for “B. pulli” was not available. The extended database includes, for this species, only those isolates that were originally identified as “B. pulli” by nox sequencing in our study.
Sequencing of nucleotides 512–926 of the nox open reading frame. Identities in BLAST analysis to the best match were at least 99% except for one B. innocens isolate and the isolates with the identification of “Brachyspira spp.” (all above 93%).
Differentiation of avian Brachyspira isolates by MALDI-TOF MS
All morphologically typical isolates and the reference strains were also subjected to MALDI-TOF MS analysis. Two of the 45 isolates could not be recovered from glycerol stocks (including one putative B. intermedia isolate) and were thus not investigated in MALDI-TOF MS. Mass spectrometry using the original Bruker database and a score of 2 as cutoff did not lead to a reliable species designation in 15 of 43 investigated isolates from poultry (Table 3). Thus, the Bruker database was extended by master spectra using reference strains (Table 1) as well as isolates from our study after species determination by nox sequencing and MALDI-TOF MS analysis. In 93% of the cases, a new MSP was based on at least 8 single-spectra of high quality (see Supplementary Fig. 1 for examples of high-quality spectra). Four MSPs were generated on 6 or 7 spectra, but with very high quality. As no reference strain was available for “B. pulli” and the MALDI-TOF database did not include this organism, detection of this species relied on the results of the BLAST analysis of the nox sequence only. The final extended database included 53 entries for 9 different Brachyspira species. Re-evaluation of the isolates using the extended database resulted in mean log scores >2 for the 43 avian Brachyspira isolates from Germany (Table 3). The distribution was as follows: B. murdochii (n = 20), B. innocens (n = 14), “B. pulli” (n = 4), B. intermedia (n = 2), and B. pilosicoli (n = 3). Noteworthy, B. innocens isolate N5.1 did not generate a nox amplification product. A distance dendrogram based on the mass spectrometric data was generated to identify similarities and clusters of the different Brachyspira strains and isolates (Fig. 1). All B. pilosicoli strains and isolates clustered together with a high degree of similarity (distance level below 100) and were clearly distinct from all of the other Brachyspira species (distance level 1,000). A further cluster with a maximal distance level of 400 was formed by the different B. intermedia strains and isolates and the 2 reference strains for B. suanatina and B. hyodysenteriae added in our study. Isolates of other species were not included in this cluster. The B. alvinipulli reference strain AN1268/3/04 was quite dissimilar to all of the other investigated Brachyspira strains and isolates (distance level above 500). A further large cluster contained all B. innocens and B. murdochii strains and isolates investigated in our study. Isolates designated as “B. pulli” and the B. hampsonii reference strains were also found in this cluster. A clear separation of the different species within this cluster was not observed (Fig. 1).
Comparison of the log scores obtained for differentiation of Brachyspira species using the original database of the manufacturer and the extended database in MALDI-TOF MS analysis.
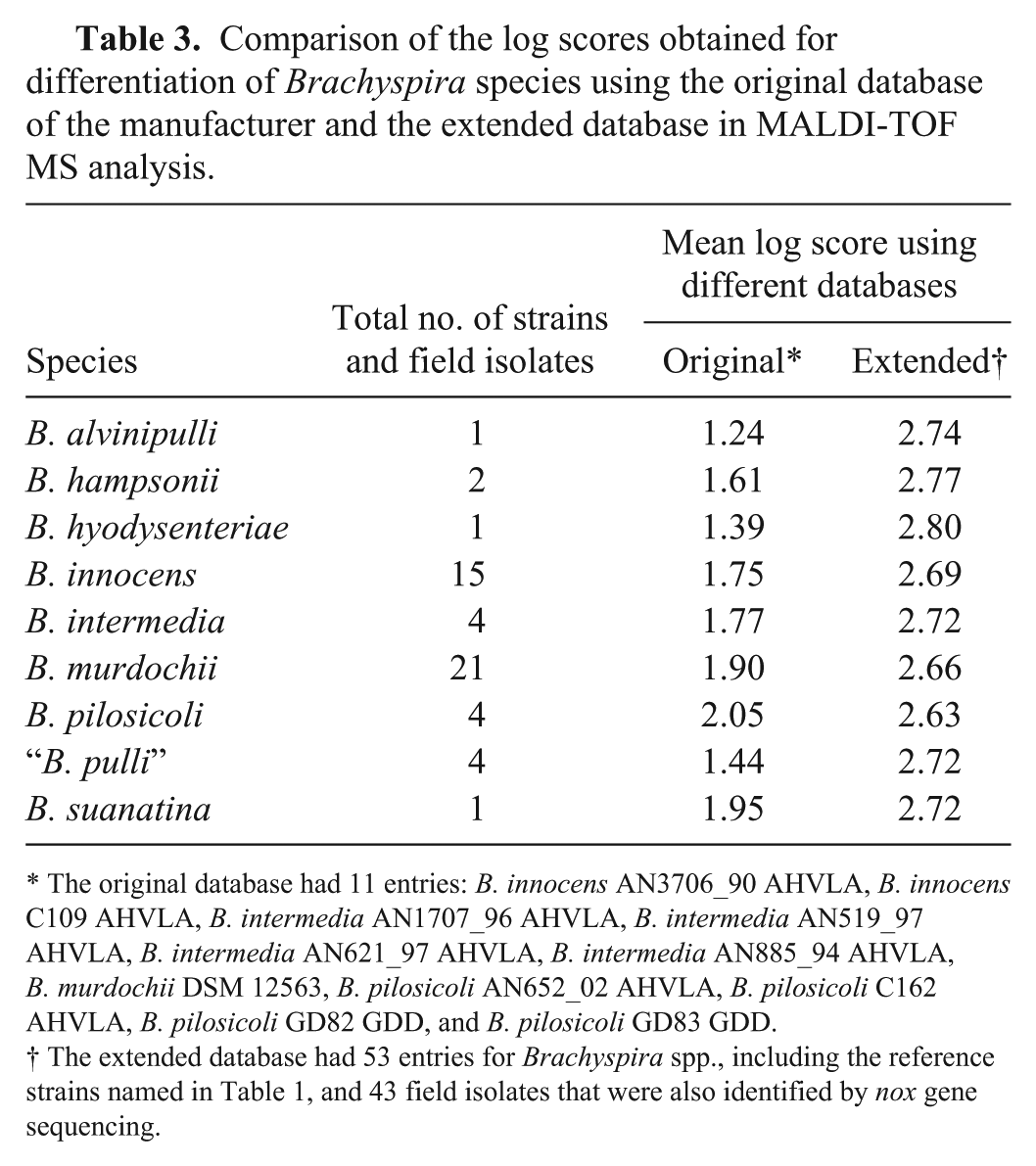
The original database had 11 entries: B. innocens AN3706_90 AHVLA, B. innocens C109 AHVLA, B. intermedia AN1707_96 AHVLA, B. intermedia AN519_97 AHVLA, B. intermedia AN621_97 AHVLA, B. intermedia AN885_94 AHVLA, B. murdochii DSM 12563, B. pilosicoli AN652_02 AHVLA, B. pilosicoli C162 AHVLA, B. pilosicoli GD82 GDD, and B. pilosicoli GD83 GDD.
The extended database had 53 entries for Brachyspira spp., including the reference strains named in Table 1, and 43 field isolates that were also identified by nox gene sequencing.
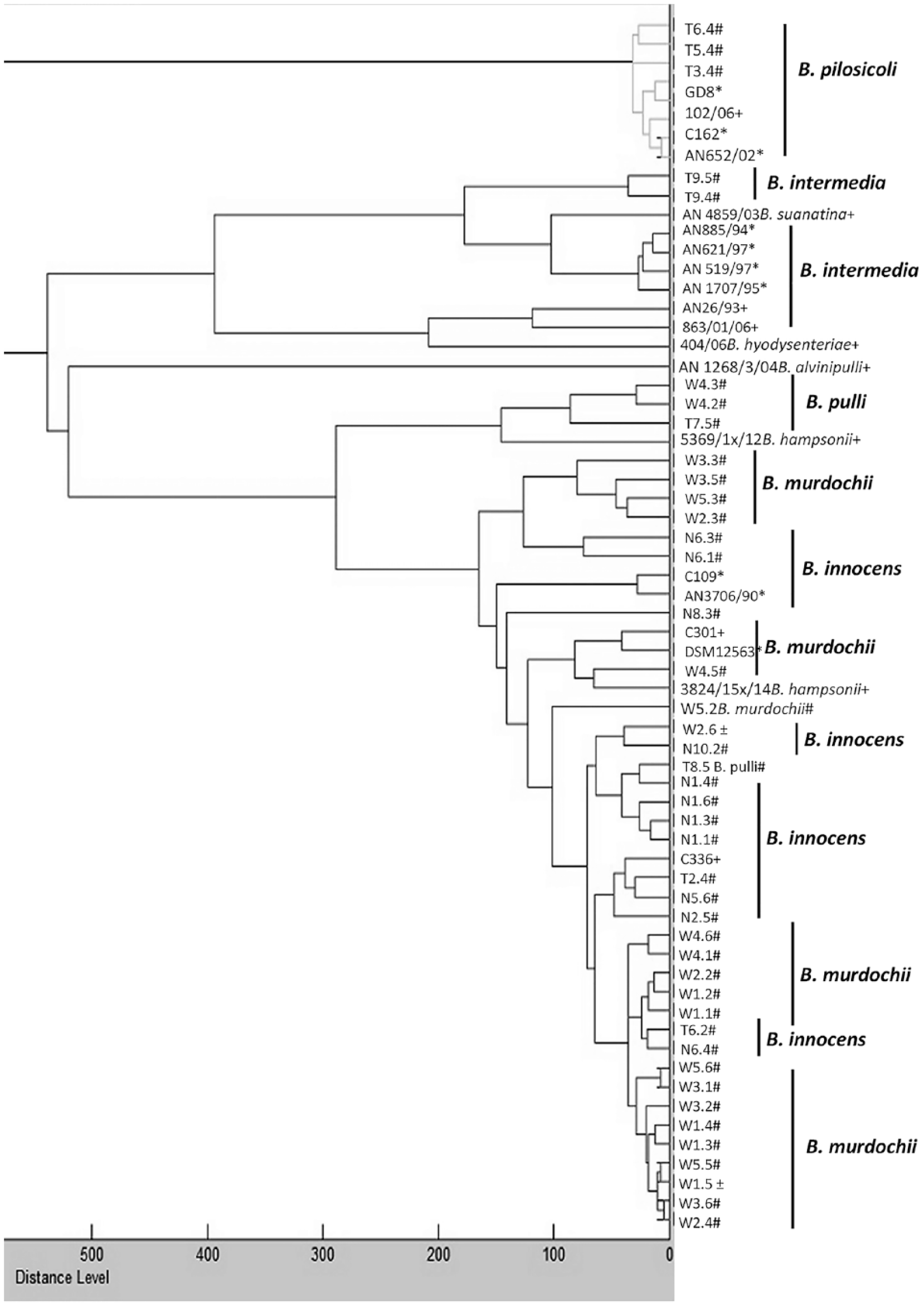
Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI–TOF MS) dendrogram based on master spectra profiles of 64 Brachyspira strains either isolated in our study or included as reference strains by our laboratory (n = 10) or by the manufacturer (n = 11, see also Table 1). Reference strains of the Bruker database are marked with *. Strains marked with + and # are reference strains and isolates from poultry included in our study, respectively. Sequencing results of the nox gene were not in accordance with species identifications obtained by MALDI-TOF MS in cases marked with ±. The distance level of the B. pilosicoli cluster to the other Brachyspira strains was 1,000.
Differentiation of avian Brachyspira isolates by a new mPCR
The mPCR assay gave the 939-bp amplicon of the nox gene for the 10 reference strains (Fig. 2). This amplification product was also generated for all field isolates except for N5.1, which also lacked the nox amplicon in the singleplex PCR. The B. hyodysenteriae reference strain (404/06), the 2 B. intermedia reference strains (AN26/93, 863x01/06), and the B. suanatina strain (AN4859/03) displayed, as expected, a second band of ~325 bp in accordance with a tnaA-positive genotype (Fig. 2). Furthermore, the field isolates identified as B. intermedia by nox sequencing were also tnaA positive (T1.6, T9.4, and T9.5). However, 2 B. murdochii isolates (W3.3 and W3.5), as determined by nox sequencing and MALDI-TOF MS analysis, also displayed the 325-bp tnaA band. The remaining isolates were negative for tnaA in the mPCR. All of the B. pilosicoli reference strains and the field isolates were positive for the nox gene and for abgB, the latter being a 744-bp product (Fig. 2). All other strains or isolates except for one putative B. innocens isolate (N6.3) were negative for the 744-bp abgB band. The abgB band was not detectable in the reference strain for B. alvinipulli AN1268/3/04.
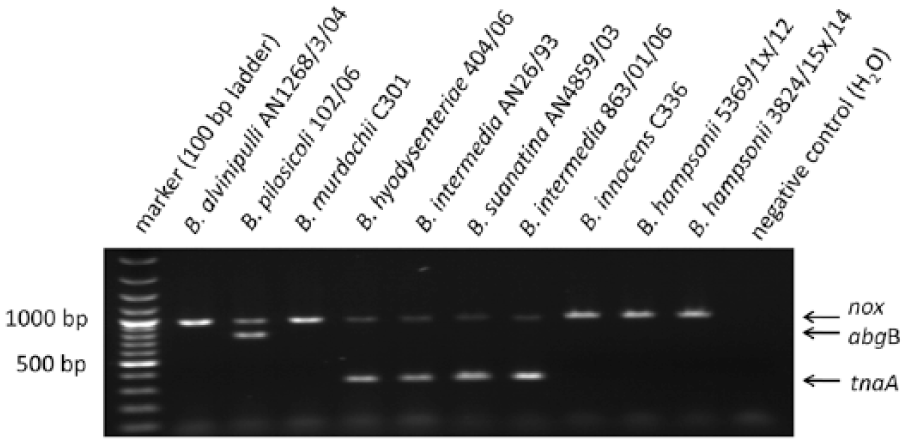
Multiplex (m)PCR targeting the reduced nicotinamide adenine dinucleotide oxidase gene (nox), the tryptophanase A gene (tnaA) and the p-aminobenzoyl-glutamate hydrolase subunit B gene (abgB) of the indicated Brachyspira species. The mPCR might be used to support identification of the avian intestinal spirochetosis causative agents B. pilosicoli and B. intermedia by detecting abgB and tnaA, respectively. The gel photo shows the results of the mPCR for the indicated reference strains included in our study.
Isolates with conflicting MALDI-TOF MS and mPCR results
The field isolates that were originally positive for tnaA (W3.3 and W3.5), but which were identified as B. murdochii in MALDI-TOF MS and nox sequencing, and the putative abgB-positive B. innocens isolate (N6.3), were streaked on FAA for single colony formation. Examination of 3 subcultured colonies of each sample revealed that these subcultures were all positive for the nox gene but negative for tnaA as well as abgB in the mPCR. Hence, the identification of B. murdochii was confirmed for all subcultures of W3.3 and W3.5. All subcultures of N6.3 received a score >2 for B. innocens.
Discussion
The NCBI database included 388 and 2,676 entries for nox and 16S ribosomal (r)DNA gene sequences of Brachyspira isolates in July 2016, respectively (mainly partial sequences). These 2 collections of sequences are by far the largest for any Brachyspira gene and have both been used in previous studies for differentiation by BLAST analysis.24,37 Because 16S rDNA sequencing turned out to have very limited differentiation power for some Brachyspira species, 31 nox sequencing has been used by different groups.19,37 Based on these findings, we decided to use nox sequencing in our study. We used a nox primer pair that has been used successfully over many years at the University of Veterinary Medicine in Hannover to identify hundreds of porcine Brachyspira isolates. 29 For most of the isolates generating a nox amplification product, we received a high identity index (33 of 41 with 99% or 100% identity) in BLAST analysis, indicating that the isolates belonged to the respective species. However, there is a pitfall in nox sequencing that should be further addressed in the future: the frequency of genetic transfer and recombination of nox sequences between different Brachyspira species is not known. Interestingly, a bacteriophage carrying nox sequences has been identified in B. hyodysenteriae. 14 If transduction of nox sequences between different Brachyspira species occurred frequently in the field, species determination based on nox sequences would not be reliable, even though it has been conducted over many years in various laboratories.23,29 Based on this uncertainty, we decided to establish and evaluate further methods for differentiation of Brachyspira. This need for additional identification tools was demonstrated by our finding that sequence analysis of the nox gene for species determination did not lead to a specific identification for various reasons in 12 of 45 isolates confirmed as Brachyspira by MALDI-TOF MS.
MALDI-TOF MS analysis has been used in 3 studies for differentiation of Brachyspira isolates obtained mainly from pigs.3,27,37 In our study, high MALDI-TOF MS scores were obtained for all isolates obtained from poultry after extending the manufacturer’s database. Thus, MALDI-TOF MS analysis is a powerful method for differentiating Brachyspira species, a conclusion reached in previous MALDI-TOF MS studies.3,27,37 The existing MALDI-TOF MS database was extended using strains or isolates that had been identified based on nox sequencing data. However, the database was extended very carefully using reference strains first and then isolates with high MALDI-TOF MS and BLAST scores. For further evaluation of MALDI-TOF MS–based differentiation, it would be expedient to analyze a further collection of avian field isolates differentiated by a method independent of nox sequencing and not used for extension of the MALDI-TOF MS database. However, such a collection of field isolates is currently not available in our laboratory.
One striking feature of the MALDI-TOF MS dendrogram (Fig. 1) is that B. innocens and B. murdochii isolates cluster together with no detectable distinction of the 2 species. Mixing of B. innocens and B. murdochii isolates was also observed previously when generating a phylogenetic tree based on partial 16S rDNA sequences of avian Brachyspira isolates. 7 The reason for this mixing is not known. The phylogenetic tree in the prior study 7 showed clustering of the B. pilosicoli strains clearly distinct from the other Brachyspira species, which was also found in the MALDI-TOF MS dendrogram in our study. Furthermore, all of the B. intermedia, B. hyodysenteriae, and B. suanatina strains form one cluster based on 16S rDNA analysis in the prior study7 and MALDI-TOF MS analysis in our study. The high concordance of these 2 studies regarding cluster formation of avian Brachyspira isolates underscores the power of MALDI-TOF MS analysis for the identification of isolates of Brachyspira.
The tnaA gene most likely encodes the enzyme responsible for the indole reaction, 16 which is a typical phenotypic feature of B. intermedia, B. suanatina, and B. hyodysenteriae, although isolates of B. hyodysenteriae strains negative for the indole reaction are well known.8,26,27 We observed discrepancies in tnaA and abgB genotyping with results of species determination by MALDI-TOF MS and nox sequence analysis in 3 cases. One isolate (N6.3) determined as B. innocens by MALDI-TOF MS and nox sequencing was positive for abgB (encoding putatively the hippurate hydrolase). Two B. murdochii isolates (W3.3 and W3.5) were at first positive for tnaA; however, these 3 abgB- and tnaA-positive genotypes, respectively, were not reproducible when subcultures of single colonies of these isolates generated on FAA were examined. Thus, we hypothesize that the original isolate N6.3 was a mixed culture with dominance of B. innocens over B. pilosicoli. Similarly, W3.3 and W3.5 are considered putative mixed cultures of B. murdochii and B. intermedia. Based on the high frequency of cultural detection of different Brachyspira species in poultry, mixed cultures have been discussed in previous work on AIS.7,13 The combination of MALDI-TOF MS analysis and the new mPCR is, in our opinion, a good approach to identify such putative mixed cultures. Generation and differentiation of single colonies on FAA is a promising method to dissect such mixed cultures. However, we did not succeed in detecting B. pilosicoli or B. intermedia in the 3 cases mentioned.
Hippurate hydrolysis has been used as an important criterion for identifying B. pilosicoli for more than 2 decades, although hippurate-negative biovariants of B. pilosicoli have been identified.4,10,21 We identified a gene by BLAST analysis that is putatively involved in hippurate hydrolysis. However, further studies including expression and characterization of recombinant AbgB and loss-of-function analysis are required to prove that abgB is necessary for hippurate hydrolysis in B. pilosicoli. Importantly, B. alvinipulli has also been described to hydrolyze hippurate, 30 but the reference strain included in our study was negative in the abgB PCR. In conclusion, the detection of abgB in all confirmed B. pilosicoli strains and isolates indicates that the mPCR targeting abgB is a useful tool in the identification of this avian pathogen.
We examined healthy flocks in our study because flocks with clear clinical signs of AIS were not available. We found that the main etiologic agents of AIS, B. pilosicoli and B. intermedia, might be detected in apparently healthy flocks, although with a very low prevalence. In the flocks we studied, B. murdochii and B. innocens appear to be more prevalent. The epidemiologic relevance of the high prevalence of these apparently nonpathogenic Brachyspira for AIS is not known. Further studies are needed to determine the prevalence of the different Brachyspira species in various poultry flocks in Germany including flocks with typical clinical signs. Based on the results of our study, we regard the combination of MALDI-TOF MS and the newly developed mPCR targeting nox, tnaA, and abgB as an improved approach to the investigation of samples from poultry for the etiologic agents of AIS.
Supplemental Material
DS1_JVDI_10.1177_1040638718772319 - Supplemental material for Differentiation of Brachyspira spp. isolated from laying hens using PCR-based methods and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry
Supplemental material, DS1_JVDI_10.1177_1040638718772319 for Differentiation of Brachyspira spp. isolated from laying hens using PCR-based methods and matrix-assisted laser desorption/ionization time-of-flight mass spectrometry by Monika Harms, Volker Schmidt, Tilo Heydel, Jutta Hauptmann, Christine Ahlers, Rene Bergmann and Christoph G. Baums in Journal of Veterinary Diagnostic Investigation
Footnotes
Acknowledgements
We thank Viveca Båverud (National Veterinary Institute, Uppsala, Sweden) for providing reference strains B. suanatina AN4859/03, B. intermedia AN26/3, B. innocens C336, B. murdochii C301, and B. alvinipulli strain AN1268/3/04. Judith Rohde (Institute for Microbiology, University of Veterinary Medicine Hannover, Germany) kindly provided B. hyodysenteriae 404/06, B. intermedia 863-1x/06, B. pilosicoli 102/06, and “B. hampsonii” strains 5369-1x/12 and 3824-15x/14. Furthermore, she is acknowledged for critically reading the manuscript. Finally, we thank Louise Eidam and Lena Dißmann for their excellent work in their traineeships.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
