Abstract
It has been proposed, based on taxonomic and molecular studies, that all canine isolates belonging to Staphylococcus intermedius group (SIG) should be renamed Staphylococcus pseudintermedius. However, isolates of SIG and other coagulase-positive staphylococci share many phenotypic characteristics, which could lead to misidentification. The accuracy of matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) for identifying S. pseudintermedius isolates obtained from canine infections was evaluated, using a polymerase chain reaction (PCR)-based identification as the gold standard. In addition, MALDI-TOF MS was compared with conventional biochemical tests. A central problem was the incorrect identification of S. pseudintermedius isolates as S. intermedius by either MALDI-TOF MS or biochemical identification. From the 49 S. pseudintermedius isolates identified by the molecular method, only 21 could be assigned to this species by the biochemical approach and only 12 by MALDI-TOF MS. The 6 S. aureus isolates were correctly identified by all 3 techniques. However, using biochemical tests, 9 S. pseudintermedius were mistakenly classified as S. aureus, indicating a reduced specificity relative to the MALDI-TOF MS system. Analysis with the MALDI-TOF MS platform allowed rapid and accurate identification of the 49 isolates to the S. intermedius group but the approach was very limited in identifying S. pseudintermedius isolates, as only 12 of 49 isolates were correctly identified, a sensitivity of 0.24 (95% confidence interval: 0.13–0.39).
The Staphylococcus intermedius group (SIG), which includes the majority of coagulase-positive staphylococci isolated from animals, causes a variety of infections such as otitis externa, pyoderma, abscesses, mastitis, and wound infections.8,13,19 SIG includes 3 distinct species: Staphylococcus intermedius, Staphylococcus pseudintermedius, and Staphylococcus delphini.1,14 The newest described species, S. pseudintermedius, is the most common cause of canine staphylococcal infections.9,13,19 It has been proposed that all canine isolates identified as S. intermedius should be reclassified as S. pseudintermedius, unless proven otherwise by genetic typing methods. 6 The accurate classification of SIG isolates is quite problematic due to the fact that they share many phenotypic characteristics among each other and with other Staphylococcus species.3,14 Thus, reliable and simple identification methods are desirable. Molecular approaches have been suggested for accurate species identification of veterinary coagulase-positive staphylococci.6,14 A common approach is based on polymorphisms within the thermonuclease (nuc) gene.3,15 Matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) has been used for rapid and accurate identification of microorganisms. 10 The aim of the current study was to evaluate the reliability of a commercial MALDI-TOF MS system a for identifying S. pseudintermedius isolates obtained from canine infections, using a nuc gene polymerase chain reaction (PCR)-based test 15 as the reference (gold standard) method. In addition, the MALDI-TOF MS approach was compared with a biochemical identification system.
The 49 S. pseudintermedius isolates tested were collected from April 2011 to May 2013, from canine infection submissions to the Department of Small Animal Medicine of the Veterinary Hospital, State University from Norte Fluminense Darcy Ribeiro, located in Campos dos Goytacazes city, Rio de Janeiro, Brazil. Twenty-one (38.2%) isolates were recovered from pyoderma cases, 18 (32.7%) from otitis media, 14 (25.5%) from soft-tissue tumor, and 2 (3.6%) from surgical wound infections. The S. pseudintermedius strain BMBP02, known to be of sequence type (ST) 71, 12 was also used as a control in the tests. As MALDI-TOF MS has been reported as an accurate method for identifying S. aureus,16–18 6 S. aureus isolates were also included for quality control purposes. This study was approved by the Ethics Committee for Animal Care and Use from Universidade Estadual Norte Fluminense (no. 145/2011).
The isolates were initially identified by routine tests (colony morphology, Gram stain reaction, and catalase and coagulase tests), and to the species levels by a PCR assay. The PCR assay used as the reference method was based on the amplification of the nuc gene using specific primers, an assay which is considered a reliable method for identifying coagulase-positive staphylococci of veterinary origin. 15 The PCR was performed using 200 µM of each deoxynucleotide triphosphate (dNTP), 0.01 µM of each primer, 0.125 U of Taq polymerase, b 1× enzyme buffer with MgCl2, and 100 ng of the DNA. Because S. pseudintermedius and S. aureus are the staphylococcal species most frequently recovered from canine pyoderma and otitis media, the PCR reactions were optimized to work as a single PCR rather than a multiplex PCR system, as previously suggested. 15 Amplification was carried out in a 96-well thermal cycler c using the following program: pre-denaturation at 95°C for 4 min; 35 cycles of 95°C for 30 sec, 51°C (reaction for S. aureus identification) or 56°C (reaction for S. pseudintermedius) for 30 sec, 72°C for 45 sec; and a final extension at 72°C for 2 min. The presence of a DNA fragment of 359 bp (S. aureus) or 926 bp (S. pseudintermedius) was confirmed by gel electrophoresis (1.5% agarose in 0.04 M Tris–acetate–ethylenediamine tetra-acetic acid, pH 8.3). The gel was then treated with 0.5 µg/mL of ethidium bromide for 15 min, and visualized in an imaging system. d
The MALDI-TOF MS analyses were performed using a commercial mass spectrometer a operated by the on-board software, e as recommended by the manufacturer. f At first, 2 protocols were tested: the extended direct transfer (EDT) and the formic acid extraction (FAE). Briefly, for the EDT, a single fresh colony was smeared (in duplicate) with the aid of a toothpick to form a thin film directly onto a polished steel target plate. f Then, 70% formic acid (1 µL) was overlaid on the film and dried at room temperature. Subsequently, the matrix solution (1 µL of α-cyano-4-hydroxycinnamic acid [HCCA]) was also overlaid, the material again dried, and analyzed by mass spectrometry with 280 shots per sample spot, using the recommended manufacturer settings for bacterial identification. For the FAE method, the bacterial suspension in distilled water was mixed with absolute ethanol (1:3), and centrifuged (15,000 × g, 2 min). The pellet was resuspended in a solution of 70% formic acid and ≥99.9% acetonitrile (20 µL each; 1:1 [v/v]), incubated at room temperature for 5 min, and centrifuged (15,000 × g, 2 min). An aliquot of supernatant (1 µL) was applied on the polished steel target plate f (in duplicate), dried at room temperature, overlaid with 1 μL of HCCA, dried again, and analyzed using the recommended manufacturer e settings. Species-level identification was assumed if the logarithmic identification score, expressing the degree of concordance with the best matching spectrum from the reference database, reached or exceeded the threshold of 2.0, as proposed by the manufacturer. f Escherichia coli strain DH5α was used for instrumental calibration. Data was obtained from at least 3 independent experiments with duplicates.
Species-level identification was also performed by conventional biochemical tests as previously described, 4 and included colony morphology, Gram stain reaction, pigmentation, hemolysin, and catalase, oxidase, and coagulase reactions. The expression of clumping factor, heat stable nuclease, alkaline phosphatase, arginine arylamidase, pyrrolidonyl arylamidase, ornithine decarboxylase, urease, β-glucosidase, β-galactosidase, and β-glucuronidase was also determined. As well, arginine utilization, acetoin production, nitrate reduction, esculin hydrolysis, and susceptibility to acriflavine and polymyxin were tested. Finally, a series of carbohydrate fermentation reactions was also included.
Student t-test (unpaired data) was used for comparing the MALDI-TOF MS scores using the EDT or FAE method. The estimated sensitivity, specificity, and accuracy, as well as the exact (Clopper–Pearson) 95% confidence interval (CI) compared with a perfect standard (nuc gene PCR-based method) were calculated following the recommendations of Food and Drug Administration statistical guidance (http://www.fda.gov/MedicalDevices/DeviceRegulationandGuidance/GuidanceDocuments/ucm071148.htm).
The 49 S. pseudintermedius isolates were confirmed as S. pseudintermedius by the nuc gene PCR (the reference method). Only a few MALDI-TOF scores were above 2.0 when the EDT methodology was used for the identification of the S. pseudintermedius isolates. The mean score was 2.25 ± 0.11 and 2.03 ± 0.17 for FAE and EDT methodologies, respectively, and this difference was statistically significant (P < 0.0001). Therefore, only the data from the FAE method was considered for these analyses. The manufacturer-recommended score of (at minimum) 2.0, for identification at species level, was achieved for 54 of the total 55 isolates (98.2%). The lowest score achieved (isolate NF851) was 1.974, even after 3 independent experiments (Table 1). Thus, considering the highest MALDI-TOF score obtained for each of the 49 S. pseudintermedius isolates, only 12 (24.5%) could be correctly identified as S. pseudintermedius, and the remaining 37 (75.5%) were classified as S. intermedius. As expected, all 6 S. aureus were correctly identified, with scores greater than 2.3 (Table 1).
Discriminating Staphylococcus pseudintermedius from Staphylococcus intermedius isolated from dog infections using polymerase chain reaction (PCR), conventional biochemical tests, and the matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) system. Staphylococcus aureus isolates were used as a control.*
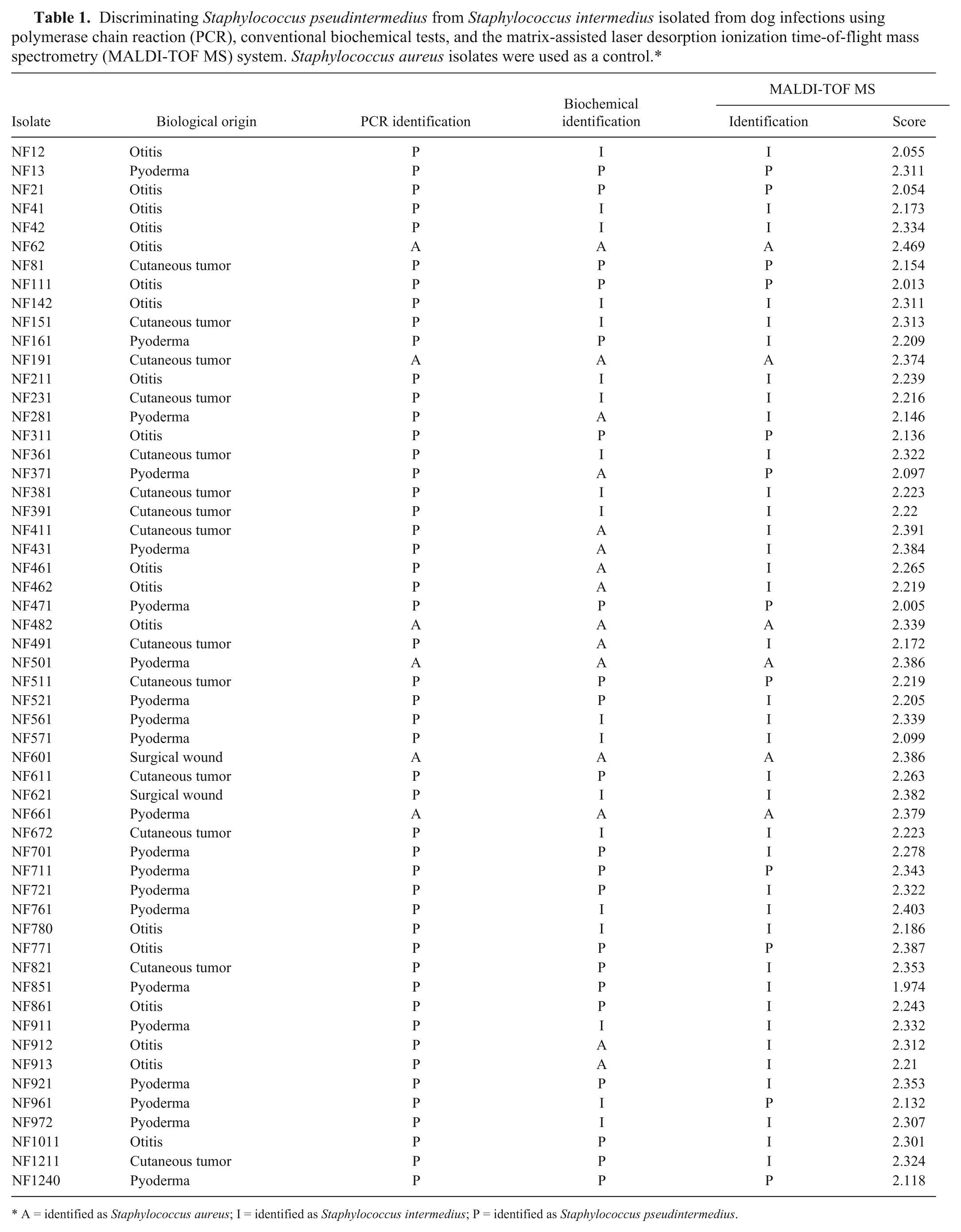
A = identified as Staphylococcus aureus; I = identified as Staphylococcus intermedius; P = identified as Staphylococcus pseudintermedius.
Biochemical identification was performed for the 55 isolates (including the 6 S. aureus isolates), and 21 (38.2%) were identified as S. pseudintermedius, 19 (34.5%) as S. intermedius, and 15 (27.3%) as S. aureus (Table 1). Therefore, from the 49 S. pseudintermedius isolates detected by the molecular method, only 21 could be allocated to this species by the biochemical approach, and just 12 by MALDI-TOF MS, corresponding to 42.9% and 24.5% of the isolates tested, respectively. All 6 S. aureus isolates tested could be identified by the 3 techniques. However, by biochemical testing, 9 PCR-identified S. pseudintermedius were mistakenly classified as S. aureus, indicating a reduced specificity in comparison with the MALDI-TOF MS system to allocate microorganisms within S. aureus species (Table 1).
Staphylococcus pseudintermedius is a major pathogen associated with canine pyoderma.8,13,20 Reports on the international spread of the highly multiresistant lineage ST71 of S. pseudintermedius over different countries,2,7,19 including Brazil, 12 raises important concerns on the correct identification of these staphylococcal species. The MALDI-TOF MS approach represents an opportunity for accurate, rapid, and reasonably priced identification of bacteria and other microorganisms in clinical microbiology laboratories. 10 Despite that, only a few studies have documented the efficiency of MALDI-TOF MS to identify S. pseudintermedius.5,11,16
In fact, the commercial MALDI-TOF system used in the current study a allowed an accurate identification of the 6 S. aureus isolates, used as control, with sensitivity and specificity of 1.00. All S. pseudintermedius isolates were classified into SIG (specificity and sensitivity of 1.00) when the FAE method was used. The specificity of the MALDI-TOF MS for discriminating the SIG isolates within S. pseudintermedius species was also 1.00. However, a lower sensitivity of 0.24 (95% CI: 0.13–0.39) was achieved, corresponding to an accuracy of just 0.33% (95% CI: 0.21–0.47; Table 2).
Specificity and sensitivity of the matrix-assisted laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) system and biochemical tests for the identification of the 55 staphylococcal isolates previously identified by a polymerase chain reaction assay.*
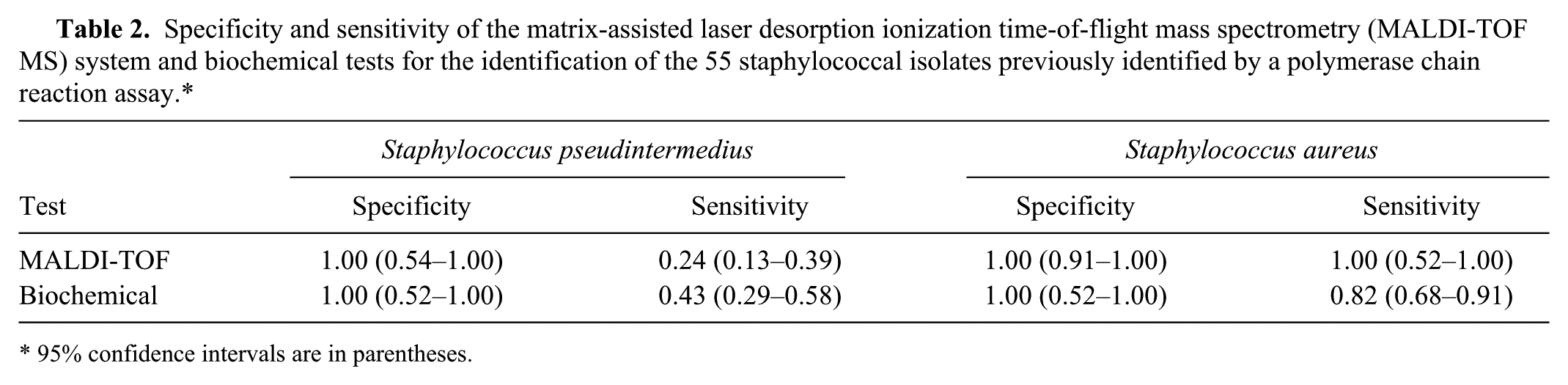

95% confidence intervals are in parentheses.
The recommendation that all S. intermedius isolates from canine infections be renamed S. pseudintermedius 6 might explain, at least in part, why the MALDI-TOF MS databank is still limited in terms of a capacity to correctly differentiate between these species. The manufacturer’s instructions f recommend that identification of an isolate should only be considered a highly probable species identification when the score is greater than 2.3. However, in the current study, the score average was 2.25 (±0.11), and only 20 (40.8%) SIG isolates reached 2.3. For most of the isolates tested using FAE, even when 3 or more independent experiments were carried out, the score was lower than 2.3.
The MALDI-TOF MS results were also compared with those of the biochemical tests. For the S. aureus isolates, the biochemical identification showed a sensitivity of 1.00 but a specificity of 0.82 (95% CI: 0.68–0.91). Actually, in the case of S. aureus, 2 studies17,18 have previously validated the superiority of MALDI-TOF MS identification over traditional biochemical systems. For the S. pseudintermedius isolates, the biochemical method showed a specificity of 1.00. Nevertheless, the sensitivity was very low, 0.43 (95% CI: 0.29–0.58), with an accuracy of only 0.49 (95% CI: 0.35–0.63; Table 2).
Using a different MALDI-TOF MS system, a higher estimated sensitivity, specificity, and efficiency for S. pseudintermedius of 0.78 (95% CI: 0.60–0.90), 0.97 (95% CI: 0.86–0.99), and 0.88 (95% CI: 0.78–0.95), respectively, has been reported. 5 In this earlier work, the overall efficiency was quite high and ranged between 0.88 and 0.99 for S. pseudintermedius–S. delphini and S. intermedius, respectively. However, these results were only achieved by use of new reference spectra (SuperSpectra) based on the most discriminating peaks for a given species. 5 In fact, the spectra previously present in the MALDI-TOF MS database led to erroneous identification of strains. 5 Similarly, an in-house reference spectra has been created for the same commercial instrument used in the current study, and the use of that in-house reference spectra resulted in 17 isolates initially identified as S. intermedius being correctly recognized as S. pseudintermedius. 11
In summary, the MALDI-TOF MS commercial system a used in the current study allowed a rapid and accurate identification of the S. aureus and SIG within 1 day, but it showed a limited ability to identify, to the species level, the SIG isolates belonging to S. pseudintermedius. In view of this limitation, the MALDI-TOF MS database a used in the current study needs to be expanded and algorithms developed to solve the existing inadequate identification of SIG isolates.
Footnotes
a.
Biotyper 3.1, Bruker, Atibaia, São Paulo, Brazil.
b.
Promega Corp., Madison, WI.
c.
Veriti 96-well thermal cycler, Applied Biosystems, Foster City, CA.
d.
LumiBis gel imager, DNRBio-Imaging Systems, Jerusalem, Israel.
e.
MALDI flexControl 3.4 and the MALDI-Biotyper 3.1 software, Bruker, Atibaia, São Paulo, Brazil.
f.
Bruker Daltonik GmbH, Bremen, Germany.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Fundação de Amparo à Pesquisa do Rio de Janeiro (FAPERJ), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), and by European Commission’s Seventh Framework Programme (FP7), through the Marie Curie International Research Staff Exchange Scheme NANO_GUARD (PIRSES-GA-2010-269138).
