Abstract
We describe herein herpesvirus-associated genital lesions in a Guiana dolphin (Sotalia guianensis) from the northern Brazilian coast. Papillary lesions on the vulva, with epithelial hyperplasia, swollen keratinocytes, and intranuclear inclusions, were positive for a herpesvirus (Gammaherpesvirinae subfamily).
Small dolphins from South American waters are affected by a variety of congenital, traumatic, infectious, and parasitic diseases.8,10 Potential contributing factors to some of these conditions may be the release of agricultural, industrial, and domestic waste. 10 Various pathogens cause a variety of infectious diseases in Guiana dolphins (syn. Guyana river dolphin, Sotalia guianensis).3,13 A novel cetacean morbillivirus was reported in 2014 in a Guiana dolphin from southeastern Brazil. 4
Several reports have described herpesviral infections in cetaceans from stranding events.1,5,18 Molecular analyses have allowed the identification of nucleotide sequences of Alphaherpesvirus and Gammaherpesvirus in skin lesions and mucous membranes of dolphins and whales.7,9,11,14 Although herpesvirus has been associated with the occurrence of plaque lesions in the mucosa of the penis and/or vulva in a variety of cetacean species,11,13,14 and herpesviral DNA has been detected in genital mucosa from odontocetes and mysticetes,5,9 herpesviral infection has not been reported previously in S. guianensis to our knowledge.
On September 13, 2014, an adult female Guiana dolphin, 104 cm long, was found recently dead, trapped in a fishing net, off the coast of Marajó Island, Pará State, Brazil. The carcass was kept on ice in a fishing boat and delivered to the Group of Studies of Aquatic Mammals in the Amazon (GEMAM; Paraense Museu Emílio Goeldi [MPEG]), maintained at −20°C until autopsy (Animal Pathology Laboratory of UFPA) on November 26, 2014. During the autopsy, tissue samples were collected in 10% neutral-buffered formalin, routinely processed, embedded in paraffin, sectioned at 5 µm, and stained with hematoxylin and eosin for histologic examination. Tissue samples were stored at −20°C for molecular studies. Genomic DNA was extracted from the frozen vulva lesion (Illustra tissue & cells genomicPrep Mini Spin Kit, GE Healthcare Life Sciences, Buckinghamshire, UK). The extracted DNA sample was forwarded to the Institute for Animal Health at the University of Las Palmas de Gran Canaria (Canary Islands, Spain), in a card (Whatman FTA Card, Sigma-Aldrich, Dorset, UK) to capture nucleic acids for molecular analysis. A conventional nested PCR using degenerate primers designed to amplify a region of the DNA polymerase gene, which contains highly conserved amino acid motifs within the Herpesviridae, was performed to detect herpesviral DNA. 17
The herpesviral PCR product was electrophoresed in 2% agarose gel, and an amplicon of ~250 bp was obtained (expected size). The sequenced product (209 nucleotides) was deposited in GenBank (accession KU666557) and compared with sequences available in GenBank using BLAST (https://goo.gl/PmPsjq). Sequence alignment was performed using Clustal W, 16 and phylogenetic analysis was performed using MEGA. 15 A maximum likelihood phylogenetic tree was constructed, and a bootstrap resampling (1,000 replicates) was used to assess the reliability of the tree. 15
The specimen was in poor body condition, and linear abrasions were observed on the skin of the face. Grossly, the most remarkable finding consisted of papillary lesions in the vulva, measuring ~0.5 cm diameter and covered with a thin layer of yellow material (Fig. 1A). The only other significant macroscopic finding was the presence of a few slender nematodes compatible with Halocercus spp. in the lumen of the trachea and bronchi. In addition, mesenteric lymph nodes and the spleen were slightly enlarged.
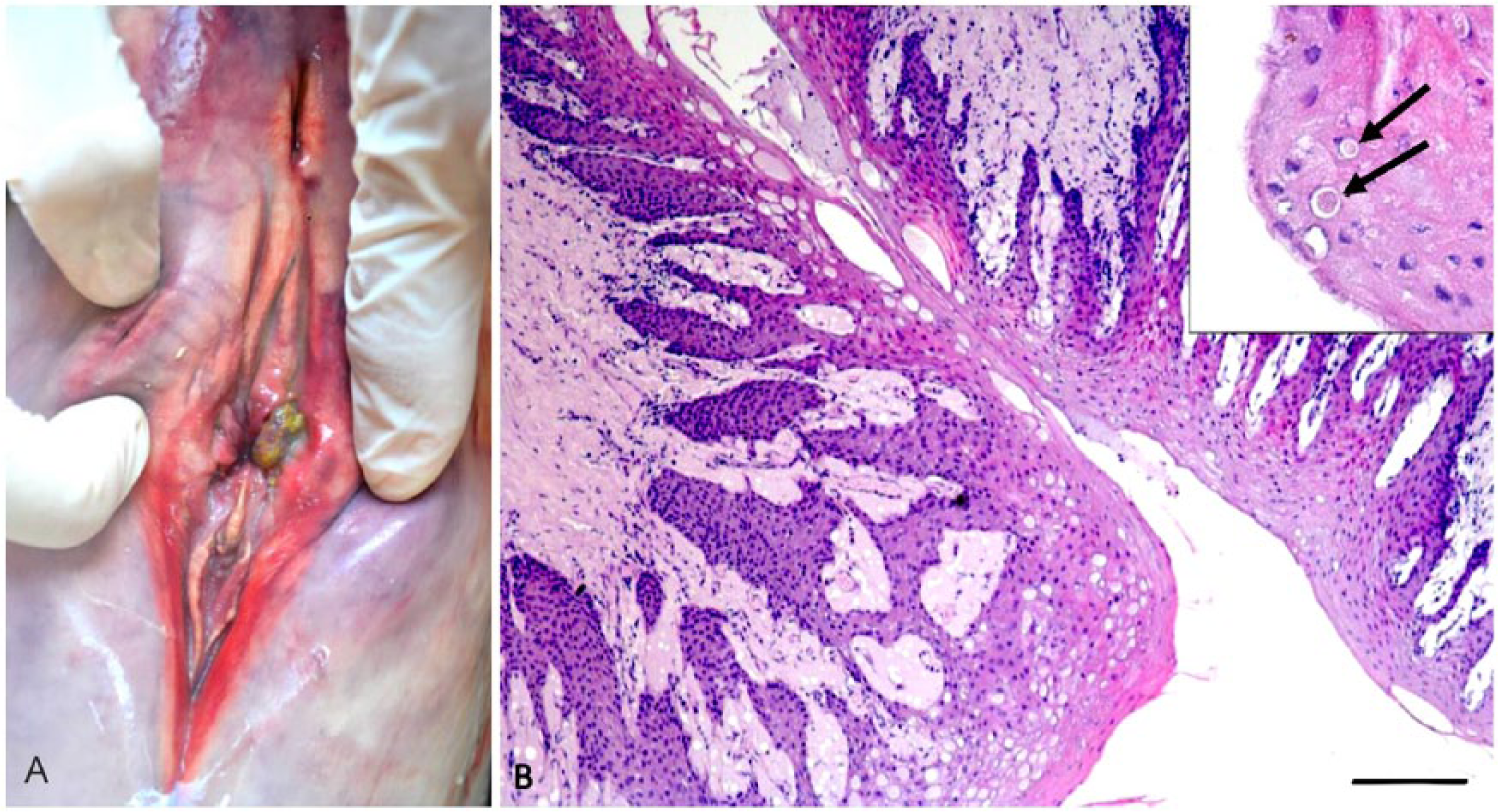
Histologically, there was moderate epithelial hyperplasia of the mucosa of the vulva (Fig. 1B), with moderate-to-severe nuclear swelling and cytoplasmic vacuolation of keratinocytes in the intermediate layers. In some areas, there was lysis of keratinocytes individually or in small groups, with discrete vesicle formation. Some keratinocytes had increased cytoplasmic eosinophilia and nuclear pyknosis. Intranuclear eosinophilic inclusions were occasionally observed in keratinocytes, particularly in the superficial layers. Herpesviral infection was suspected based on the gross appearance of the lesion and on the microscopic findings. 13
The meninges of the medulla oblongata had diffuse and moderate infiltrates of lymphocytes and macrophages. There were also rare foci of periarteritis with fibrinoid necrosis of arterioles. The cerebellar meninges had mild multifocal mononuclear cell infiltrates. A few areas of coagulative necrosis were observed in a mesenteric lymph node. The involvement of herpesvirus in the brain and lymph node lesions was not confirmed. Herpesvirus causing similar lesions has been reported previously in other cetacean species.2,6 However, the possibility of co-infection with a morbillivirus cannot be ruled out. 12
The vulvar lesion tested positive for herpesvirus based on sequences of the DNA polymerase gene. Phylogenetic analysis showed that the sequence obtained from the vulvar mucosa of the Guiana dolphin belonged to the Gammaherpesvirinae subfamily. Sequences from striped (Stenella coeruleoalba; KM248274, GQ888672), bottlenose (Tursiops truncatus; DQ288667, AY952776), and Risso’s dolphins (Grampus griseus; KJ406184, DQ288666) were most closely related to this novel sequence (amino acid p-distance: 0.078; Fig. 2). Research with herpesvirus in marine mammals is relatively recent, with studies conducted mainly in cetaceans from the North Atlantic Ocean and the Mediterranean Sea.2,6,13 There are reports of skin disorders in Guiana dolphins of unknown etiology. 19
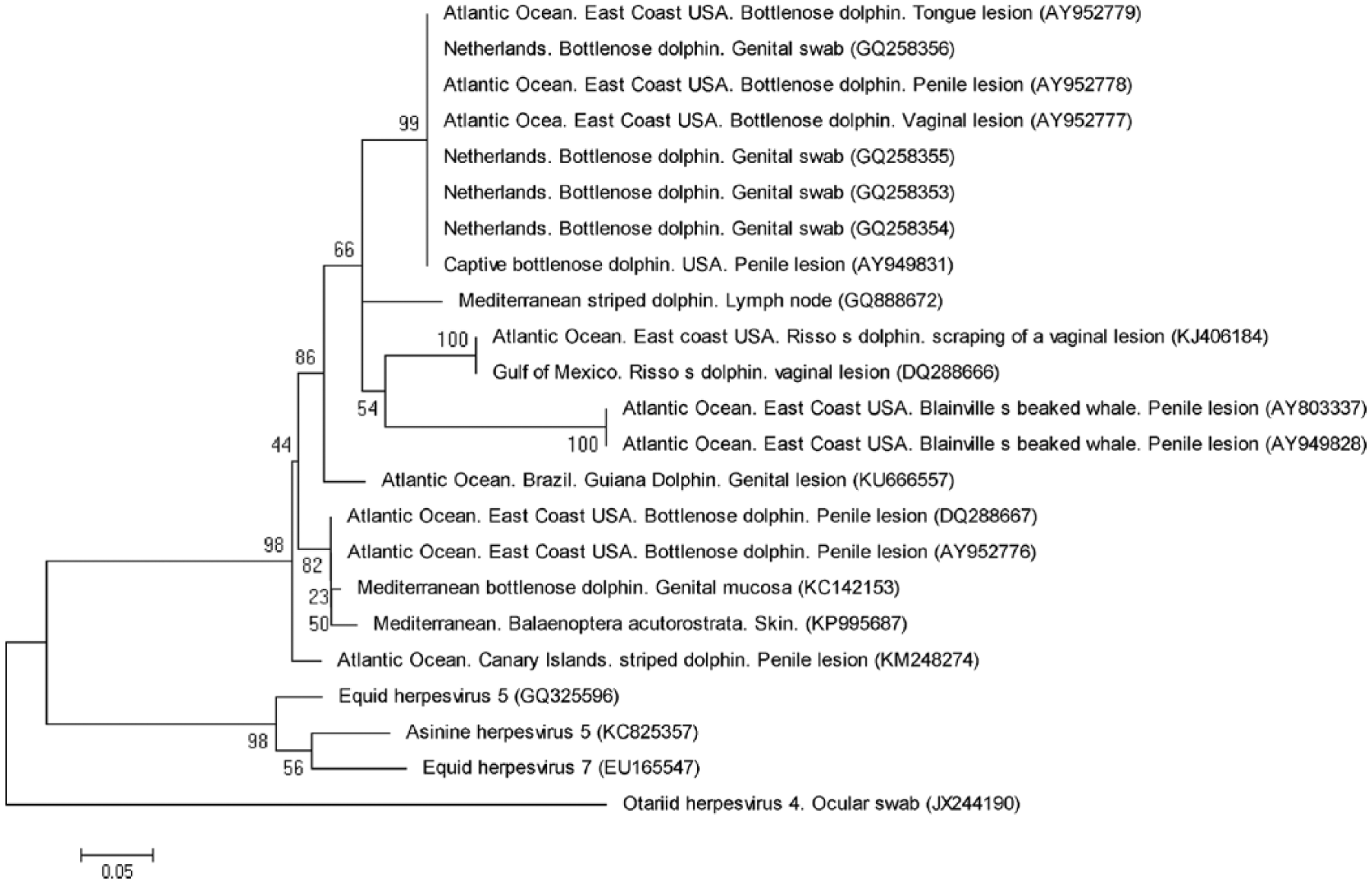
Neighbor-joining phylogenetic tree comparing a new isolate of herpesvirus from a Guiana dolphin (Sotalia guianensis) from the northern Brazilian coast with the related herpesviral polymerase gene sequences.
Footnotes
Acknowledgements
We thank PROPESP/UFPA for supporting the publication of our study.
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
R. Emin-Lima is supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq; Process 313356/2015-7). This work was authorized under number 38130-1 ICMBio-SISBIO.
