Abstract
We tested samples from 1,015 chelonians in Europe for Mycoplasma spp., herpesviruses, ranaviruses, picornaviruses, and ferlaviruses by PCR. Mycoplasma spp. were detected in 42.1% and herpesviruses were detected in 8.0% of tested chelonians. Differentiation of the herpesviruses revealed that 46.9% of the detected chelonian viruses were testudinid herpesvirus 1 (TeHV-1) and 54.3% were TeHV-3, including co-detections of TeHV-1 and -3 in 3 tortoises. TeHV-4 was detected in a leopard tortoise (Stigmochelys pardalis), and a herpesvirus that could not be further characterized was found in a pond slider (Trachemys scripta). Picornaviruses (topiviruses) were detected in 2.2% of the tested animals; ferlaviruses were found in 0.6%; no ranaviruses were detected in any of the animals tested. Mycoplasma spp. were detected significantly more often in Horsfield’s tortoises (Testudo horsfieldii), leopard tortoises, and Indian star tortoises (Geochelone elegans) than in other species. Horsfield’s tortoises were also significantly more often positive for TeHV-1. Mycoplasma and TeHV-1 were co-detected in 3.0%, and mycoplasma and TeHV-3 in 2.3%. The TeHV-4–positive tortoise was also positive for mycoplasma. Mycoplasma and picornaviruses were co-detected in 1.2% of the tortoises. A spur-thighed tortoise (Testudo graeca) was positive for mycoplasma and a ferlavirus. In some cases, >2 pathogens were detected. A significant correlation between mycoplasma and herpesvirus detection was found. Of all tested animals, 47.6% were positive for at least one pathogen, demonstrating the importance of pathogen detection in captive chelonians.
Keywords
Introduction
Respiratory disease in chelonians is a well-known problem. Infection of the respiratory tract is often caused by mycoplasma and herpesviruses, which are well-investigated pathogens in chelonians and have been detected regularly in tortoises and turtles in Europe and the United States. The first isolation of mycoplasma from a tortoise was from the cloaca of a spur-thighed tortoise (Testudo graeca). The isolated species was called Mycoplasma testudinis and is not associated with clinical disease. 24 The best characterized Mycoplasma sp. in tortoises is Mycoplasma agassizii, causing upper respiratory tract disease (URTD) in various tortoise species. 7 URTD is associated with chronic rhinitis and conjunctivitis, nasal and ocular discharge, and palpebral edema.7,8,27,28,62 M. agassizii has been detected in numerous wild and captive tortoise species particularly in desert tortoises (Gopherus agassizii)7,9,27,28 and gopher tortoises (Gopherus polyphemus)8,9 in the United States, and in spur-thighed tortoises, Hermann’s tortoises (Testudo hermanni), Horsfield’s tortoises (Testudo horsfieldii), and marginated tortoises (Testudo marginata)61,65 in Europe, but also in other tortoise species. Mycoplasma testudineum has been found in desert tortoises6,29 and gopher tortoises (Wendland LD. Epidemiology of mycoplasmal upper respiratory tract disease in tortoises [PhD dissertation]. University of Florida, Gainesville, FL, 2007) and is also associated with URTD, 6 but is believed to be less pathogenic than M. agassizii. 29
Other Mycoplasma spp. have been described in free-ranging European tortoises (T. hermanni, T. graeca, T. marginata) 38 with and without URTD. Phylogenetic analyses have shown a close relationship to M. agassizii. 38 A novel Mycoplasma sp. was obtained from the phallus of a wild desert tortoise (Wellehan JFX Jr, et al. Identification of a novel herpesvirus and a novel Mycoplasma sp. in samples from translocated wild desert tortoises. Proc Ann Meeting Symp Desert Tortoise Council; Ontario, CA; 2014:35).
Although the majority of studies have concentrated on mycoplasmosis in tortoises, Mycoplasma spp. have also been described in emydid turtles including free-ranging eastern box turtles (Terrapene carolina carolina),18,54 western pond turtles (Actinemys [Emys] marmorata), red-eared sliders (Trachemys scripta elegans), 63 bog turtles (Glyptemys muhlenbergii), spotted turtles (Clemmys guttata), 54 and three-toed box turtles (Terrapene carolina triunguis) 57 in the United States, and a captive ornate box turtle (Terrapene ornata ornata) in Hungary. 16 All of these Mycoplasma spp. show high sequence identity. 54 Mycoplasma-positive turtles have been described with URTD-like syndromes (nasal and ocular discharge, swollen eyelids), 18 but also without any clinical signs.16,54,63
Herpesviruses described in chelonians belong to the subfamily Alphaherpesvirinae, family Herpesviridae.32,44,52,68 In tortoises, 4 genetically distinct herpesviruses have been described, with the proposed names testudinid herpesvirus 1–4 (TeHV-1–4). 40 Infections with herpesviruses in tortoises are often associated with ulcerative to diphtheroid-necrotizing stomatitis, glossitis, rhinitis, dyspnea, conjunctivitis, edema of the neck, anorexia, and lethargy; 48 meningoencephalitis 21 and necrotizing hepatitis 22 have been described. TeHV-1 has been described most frequently in Horsfield’s tortoises, 69 but also in other species.40,66 TeHV-2 was detected in a captive California desert tortoise with stomatitis 32 and in wild desert tortoises from the United States. 30 TeHV-3 has been detected in many different tortoise species, most commonly in European tortoises (T. graeca, T. marginata, T. hermanni, T. horsfieldii). Infections in Hermann’s tortoises and Horsfield’s tortoises are associated with higher morbidity and mortality than in spur-thighed tortoises,42,48 but have also been described in multiple other species. TeHV-4 was detected in a clinically healthy bowsprit tortoise (Chersina angulata) in the United States. 5
Herpesviruses have also been described in freshwater turtles. First reports were based only on electron microscopic findings of necrosis associated with intranuclear viral inclusion bodies mostly in the liver in Pacific pond turtles (Clemmys marmorata), 19 painted turtles (Chrysemys picta), 13 and map turtles (Graptemys spp.) 26 with fatal systemic diseases. In recent years, herpesviruses have been detected in several freshwater turtles by PCR, and these have been further characterized by partial genome sequencing and phylogenetic analysis. The novel herpesviruses detected include emydid herpesvirus 1 in eastern river cooters (Pseudemys concinna concinna), 35 northern map turtles (Graptemys geographica), and painted turtles 56 ; terrapene herpesvirus 1 64 and 2 72 in eastern box turtles; glyptemys herpesvirus 1 in bog turtles; glyptemys herpesvirus 2 in wood turtles (Glyptemys insculpta); and emydid herpesvirus 2 in bog turtles and spotted turtles. 55 Another herpesvirus, named pelomedusid herpesvirus 1, was detected in 2 West African mud turtles (Pelusios castaneus). 46 All of these herpesviruses show close genetic relationships to each other and to TeHVs, as well as to herpesviruses found in sea turtles, including species Chelonid alphaherpesvirus 5, the type species of genus Scutavirus (International Committee on Taxonomy of Viruses, http://ictvonline.org/virusTaxonomy.asp ). Moreover, a herpesvirus that was not further characterized by sequencing was found in an Australian Krefft’s river turtle (Emydura macquarii krefftii) by electron microscopy. 12
Other important pathogens of ectothermic animals are ranaviruses, which, in reptiles, have been mostly described in chelonians. 15 Ranaviruses have been found in numerous tortoise and turtle species, including Horsfield’s tortoises, 39 Hermann’s tortoises, 43 Chinese soft-shell turtles (Pelodiscus [Trionyx] sinensis), 10 eastern box turtles,2,3,14,34 leopard tortoises (Stigmochelys [Geochelone] pardalis), 4 Burmese star tortoises (Geochelone platynota), ornate box turtles, red-eared sliders, 33 gopher tortoises, Florida box turtles, 34 marginated tortoises, spur-thighed tortoises, and Egyptian tortoises (Testudo kleinmanni; Uhlenbrok C. [Detection of ranavirus infections in tortoises and characterization of virus isolates]. [Dissertation]. Justus-Liebig-Universität Gießen, Gießen, Germany, 2010). Infections with ranaviruses in these species have been associated with lethargy, anorexia, ulcerative stomatitis, ocular and nasal discharge, conjunctivitis, subcutaneous cervical edema, and “red-neck disease.” 10 A ranavirus isolated from soft-shelled turtles in China was completely sequenced 25 and shown to be very closely related to species Frog virus 3, the type species of genus Ranavirus. A previous study described 2 complete ranaviral genomes from isolates obtained from a Hermann’s tortoise (CH8/96) and an Egyptian tortoise (tortoise ranavirus 1, ToRV-1) as well as a partial genome analysis of another ranavirus from a Hermann’s tortoise (ToRV-2). 67 Phylogenetic analyses showed that reptilian ranaviruses were often more closely related to amphibian ranaviruses than to each other, and strains tend to cluster according to geographic origin of the specimen. 67 These and other studies suggest that some isolates of ranaviruses can be transmitted not only between different families, but between different classes of ectothermic animals.15,31,67
Additional viruses have been detected in tortoises and associated with clinical disease, including picornaviruses and paramyxoviruses. Picorna-like viruses were first isolated in cell culture and have been called virus “X” because it was difficult to characterize these pathogens (Marschang RE, Ruemenapf TH. Virus “X”: characterizing a new viral pathogen in tortoises. Proc Assoc Reptil Amphib Vet; Oct 2002; Reno, NV). These viruses have most frequently been isolated from spur-thighed tortoises, but also from marginated tortoises, Hermann’s tortoises, leopard tortoises, Egyptian tortoises, Horsfield’s tortoises, and Indian star tortoises (Geochelone elegans). Infections with picornaviruses have been associated with softening of the carapace in young tortoises, 23 and diphtheroid-necrotizing stomatitis, rhinitis, pneumonia, conjunctivitis, enteritis, and ascites in tortoises of various ages, but have also been detected in clinically healthy animals (Marschang RE. [Isolation and characterization of irido-, herpes- and reovirus from tortoises and description of an uncharacterized cytopathogenic agent]. [Dissertation]. Justus-Liebig-Universität Gießen, Gießen, Germany, 2001). Complete genome sequencing has shown that the picornaviruses found in tortoises in Europe are genetically distinct from previously described picornaviruses, and the name “topivirus” (tortoise picornavirus) has been suggested for these viruses. 17 A genetically distinct picornavirus has been detected in Sulawesi tortoises (Indotestudo forstenii) in the United States, and the name “rafivirus” has been suggested for a new genus for this virus. 50
Paramyxoviruses (PMVs) in the genus Ferlavirus have been described regularly in squamates, especially in snakes with respiratory disease, and the ferlaviruses can be divided into 4 genogroups: A, B, C, and tortoise. There are only a few reports of ferlavirus infections in chelonians. Electron microscopic findings of PMV-like particles have been described in Mediterranean tortoises with dermatitis (Zangger N, et al. [Viral dermatitis in the spur-thighed (Testudo graeca) and Hermann’s (Testudo hermanni) tortoises in Switzerland]. Proc Internat Colloq Pathol Med Reptil Amphib; Sept 1991; Bad Nauheim, Germany). A PMV isolated from a Hermann’s tortoise with pneumonia was found to be related to, but distinct from, described ferlaviruses from snakes and lizards. 45 Two distinct ferlaviruses from genogroups A and B were detected in a leopard tortoise with pneumonia. 58
Although there are numerous studies about mycoplasma, herpesvirus, and ranavirus in captive and wild chelonians, there are no detailed studies on the incidence of these pathogens in European veterinary practice, to our knowledge. Furthermore, there are only a very limited number of reports on topivirus and ferlavirus detection in chelonians in Europe. We tested samples from 1,015 chelonians from Europe for mycoplasma, herpesviruses, ranaviruses, picornaviruses, and ferlaviruses by PCR. These pathogens were selected because they can all be detected by PCR, and have all been associated with clinical signs of respiratory disease. We aimed to gain insight into the incidence of these pathogens in veterinary practice and to analyze the correlation of detection of specific pathogens and the chelonian species involved as well as the incidence of multiple pathogen detections in pet chelonians.
Material and methods
Samples from 1,015 chelonians were included in our study. Five chelonian families, Testudinidae (n = 832), Emydidae (n = 26), Geoemydidae (n = 16), Kinosternidae (n = 2), and Chelidae (n = 2), were involved, as well as chelonians (n = 133) and aquatic turtles (n = 4) of unknown species (Table 1). Samples were submitted from veterinary clinics or zoologic collections from Europe and Asia (1 case) between January 2014 and June 2015, mainly for testing for herpesvirus and mycoplasma. The case history was unknown in most cases and sometimes no species was provided. Samples included mainly oral swabs (n = 952) and nasal washes (n = 84), and in a few cases tissues (n = 26) and blood (n = 13). For 60 animals, 2 sample materials, in most cases a swab and nasal wash (n = 56), and in 4 cases a swab and blood, were submitted.
Animals tested and results for herpesviruses, mycoplasma, picornaviruses, and ferlaviruses by PCR.
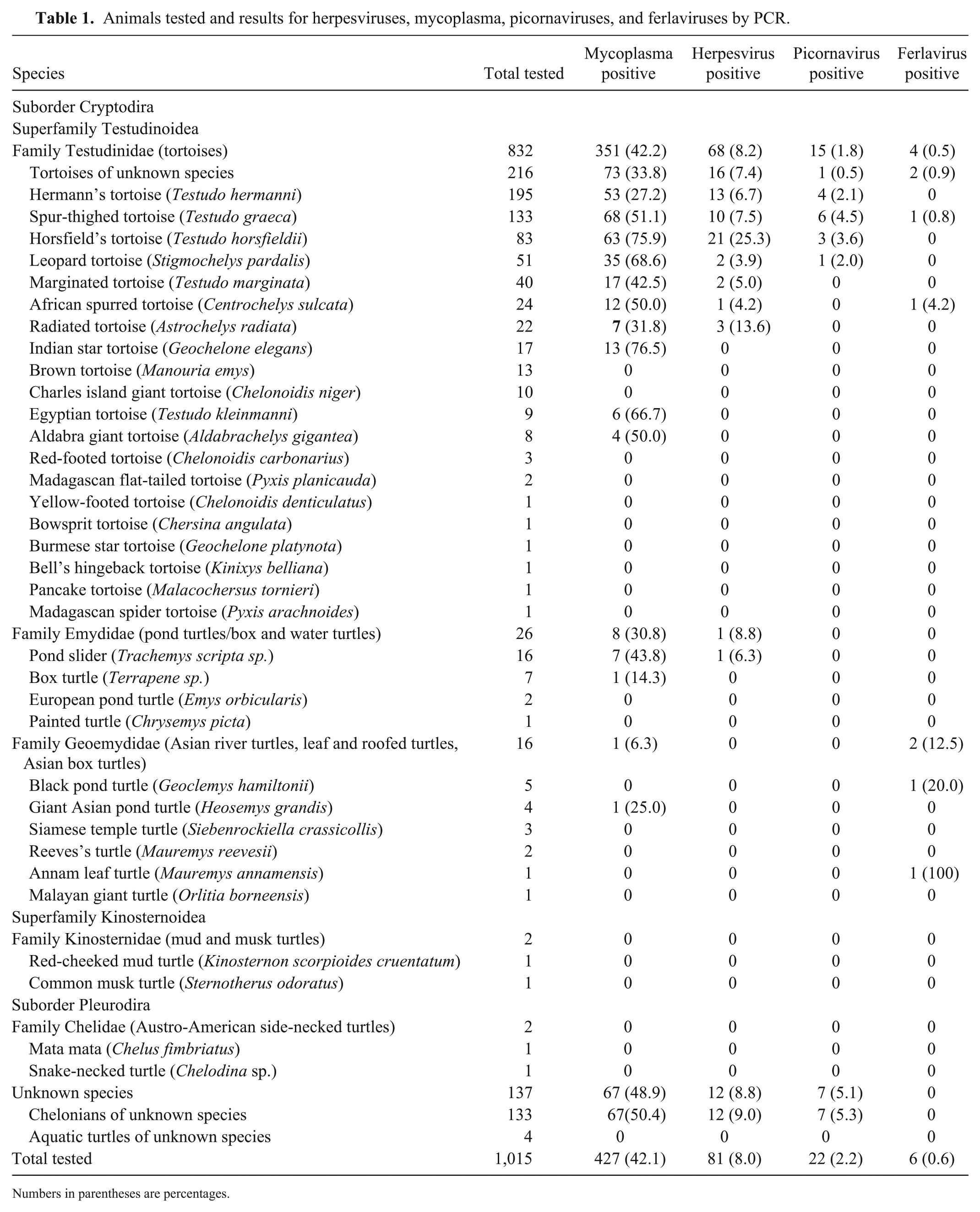
Numbers in parentheses are percentages.
After arrival at the laboratory, samples were processed within 24 h. Prepared DNA and RNA were stored at 4°C for 1 wk and then at −18°C. After DNA and RNA isolation, PCRs were performed within 6 mo in groups of 20–40 samples. DNA and RNA were extracted from samples using a commercial kit (MagNA Pure 96 DNA and viral NA small volume kit, Roche) according to the manufacturer’s instructions. PCRs were carried out for the detection of Mycoplasma spp., 8 ranaviruses,39,43 and ferlaviruses 1 as described previously. For detection of herpesviruses, 3 different PCRs were performed (Table 2). The consensus PCR detects a small region of the DNA polymerase gene of herpesviruses and is able to detect all known testudinid herpesviruses, 71 as well as many other members of the family Herpesviridae. The second PCR also amplifies a part of the DNA polymerase gene, but is only able to detect TeHV-1.44,49 The third PCR detects a part of the UL5 gene and is specific for TeHV-3.44,68 For the detection of topiviral RNA, PCRs were carried out using primers targeting a portion of the picornaviral polyprotein (Table 2). Reaction mixes for the topivirus PCRs were prepared (RealTime Ready RNA virus master kit, Roche) according to the manufacturer’s instructions. Thermocycler conditions were 50°C for 15 min, 95°C for 7 min, 9 “touch down” cycles at 95°C for 10 s, 62.5°C for 30 s (–1°C per cycle), and 72°C for 50 s, and 26 cycles at 95°C for 10 s, 54.5°C for 30 s, and 72°C for 50 s, and a final extension step at 72°C for 7 min. Positive samples yielded 400-bp products. Amplified PCR products from all of the PCRs listed were visualized with gel electrophoresis (E-Gel electrophoresis system, Invitrogen). Selected samples with positive PCR products were purified (MinElute purification kit, Qiagen) according to the manufacturer’s instructions. Sequencing was performed (Big-Dye Terminator v3.1 cycle sequencing kit, Life Technologies) and analyzed (ABI 3130 sequencer, Applied Biosystems). Sequences were manually edited, and primer sequences were removed for further analysis. The sequences were compared to other sequences in GenBank using BLASTN (https://goo.gl/jrtHqS).
Primers used and references for detection of different pathogens by PCR.
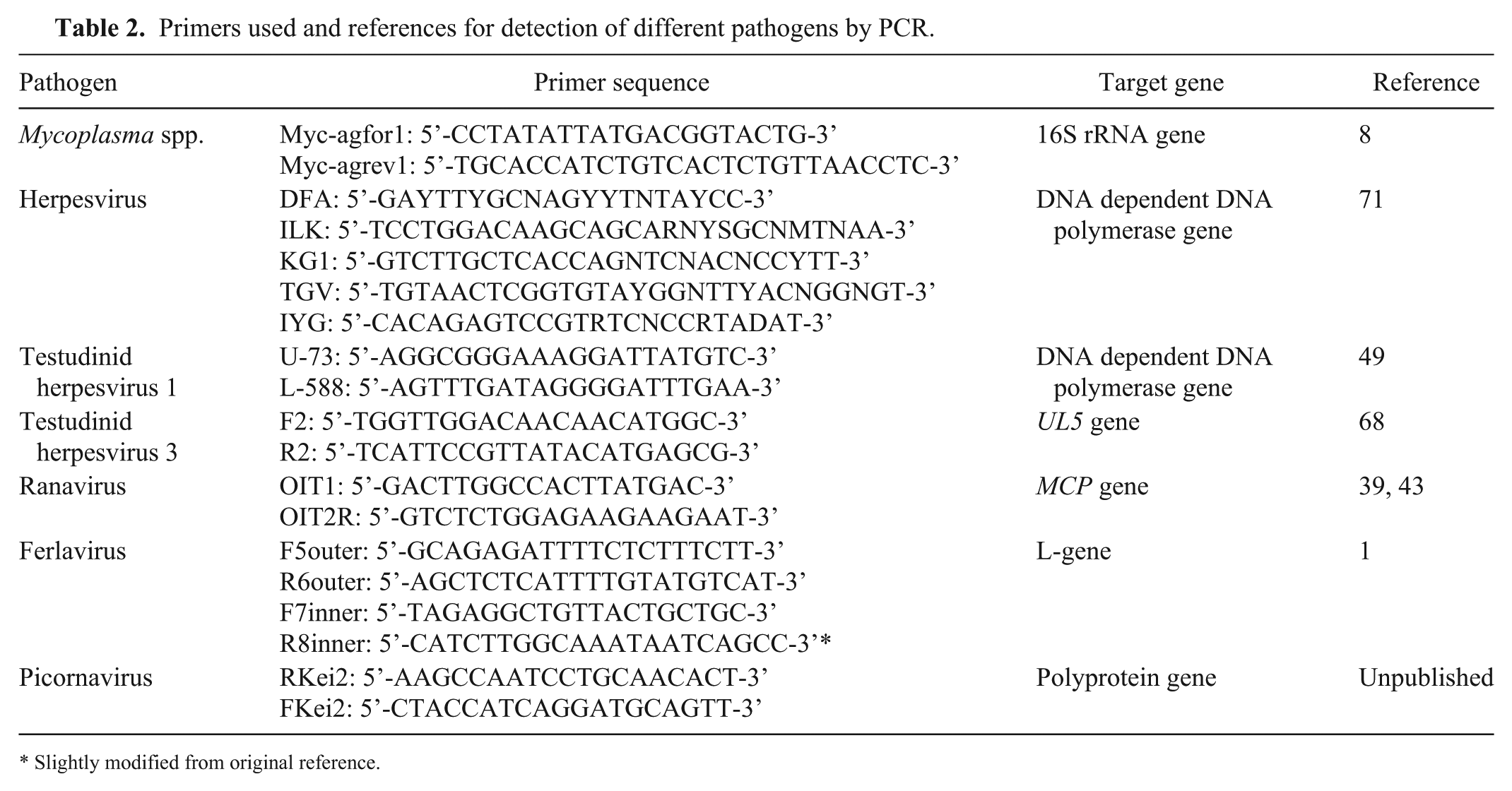

Slightly modified from original reference.
Confidence intervals (95% CI) for prevalences in chelonian species were determined by the Clopper and Pearson method using the Graph Pad Software Prism (Graph Pad Software, San Diego, CA). The Pearson chi-squared test was used with a type I error α of 0.05 to test the independence of compared prevalences. Fisher exact test was used for small sample sizes tested (n < 30). Bonferroni correction was used to control Pearson chi-squared tests computed with multiple values. The odds ratio (OR) was calculated testing the association of double-detections with Mycoplasma spp. and herpesviruses. A p value < 0.05 was considered significant.
Results
Mycoplasma spp. were detected in 427 (42.1%, 95% CI: 39.1–45.1%) of the 1,015 chelonians tested (Table 1), of which the majority were tortoises, with 351 (42.2%, 95% CI: 38.9–45.6%) of 832 tortoises, including positive tests in 68 (51.1%) of 133 spur-thighed tortoises, 63 (75.9%) of 83 Horsfield’s tortoises, 53 (27.2%) of 195 Hermann’s tortoises, 35 (68.6%) of 51 leopard tortoises, 17 (42.5%) of 40 marginated tortoises, 13 (76.5%) of 17 Indian star tortoises, 12 (50.0%) of 24 African spurred tortoises (Centrochelys sulcata), 7 (31.8%) of 22 radiated tortoises (Astrochelys radiata), 6 (66.7%) of 9 Egyptian tortoises, 4 (50.0%) of 8 Aldabra giant tortoises (Aldabrachelys gigantea), and 73 (33.8%) of 216 tortoises of unknown species. Mycoplasma spp. were also detected in 7 (43.8%) of 16 pond sliders (Trachemys scripta sp.), 1 (14.3%) of 7 box turtles, 1 (25.0%) of 4 giant Asian pond turtles (Heosemys grandis), and 67 (50.4%) of 133 chelonians of unknown species.
A total of 64 (15%) PCR products were sequenced. Positive samples for sequencing were selected randomly from each species that tested positive. Selected products were obtained during different years and months throughout the project. Comparison of the obtained nucleotide sequences with those available in GenBank using BLASTN showed 95–100% sequence identity with previously described Mycoplasma spp. in tortoises (GenBank accessions HQ326265.1–HQ326277.1). 38 Sequence identities to M. agassizii (accession NR114450.1) and M. testudineum (accession JF666138.1) were 99–95% and 94%, respectively. All positive samples from the pond sliders and the positive samples from the box turtle and the giant Asian pond turtle were also sequenced and showed 99–100% sequence identity with previously described Mycoplasma spp. of emydid turtles (accessions KJ623622.1–KJ623625.1 and FJ59564.1)16,54 and 98% sequence identity with Mycoplasma spp. described in tortoises (accessions HQ326265.1–HQ326277.1). 38 Sequence identities to M. agassizii (accession NR114450.1) and M. testudineum (accession JF666138.1) were 98% and 95%, respectively. Unique sequences have been submitted to GenBank and given the accessions KY031330–KY031336. There were no significant differences in the frequency of mycoplasma detections among Horsfield’s tortoises, leopard tortoises, and Indian star tortoises. These 3 species were positive for Mycoplasma spp. significantly more often than all other chelonians (p < 0.0001). Comparing species regarding their origin, European tortoises (T. hermanni, T. graeca, T. marginata) were less frequently positive for Mycoplasma spp. than African tortoises (A. radiata, C. sulcata, S. pardalis, A. gigantea, T. kleinmanni) and Asian tortoises (T. horsfieldii, G. elegans, Manouria spp., H. grandis, Geoclemys hamiltonii; p = 0.0001).
Herpesviruses were detected in 81 (8.0%, 95% CI: 6.5–9.8%) of 1,015 animals (Table 1), including 68 (8.2%, 95% CI: 6.5–10.2%) of 832 tortoises, 1 (6.3%) of 16 pond sliders, and 12 (9.0%) of 133 chelonians of unknown species. Tortoise species positive for herpesviruses included 21 (25.3%) of 83 Horsfield’s tortoises, 13 (6.7%) of 195 Hermann’s tortoises, 10 (7.5%) of 133 spur-thighed tortoises, 3 (13.6%) of 22 radiated tortoises, 2 (5.0%) of 40 marginated tortoises, 2 (3.9%) of 51 leopard tortoises, 1 (4.2%) of 24 African spurred tortoises, and 16 (7.4%) of 216 tortoises of unknown species.
Differentiation of the herpesviruses by genotype-specific PCRs (Table 3) showed that 38 (46.9%) animals were positive for TeHV-1, including 19 (90.5%) Horsfield’s tortoises, 3 (23.1%) Hermann’s tortoises, 2 (20.0%) spur-thighed tortoises, 1 (50.0%) leopard tortoise, 8 (50.0%) tortoises of unknown species, and 5 (41.7%) chelonians of unknown species. TeHV-3 was detected in 44 (54.3%) animals, including 3 (14.3%) Horsfield’s tortoises, 10 (76.9%) Hermann’s tortoises, 9 (90.0%) spur-thighed tortoises, 3 (100%) radiated tortoises, 2 (100%) marginated tortoises, 1 (100%) African spurred tortoise, 8 (50.0%) tortoises of unknown species, and 8 (66.7%) chelonians of unknown species. TeHV-1 and TeHV-3 were co-detected in a spur-thighed tortoise, a Horsfield’s tortoise, and a chelonian of unknown species. In 1 (50.0%) leopard tortoise, TeHV-4 (1.2%) was detected. 36
Differentiation of herpesviruses in positive tortoise species.
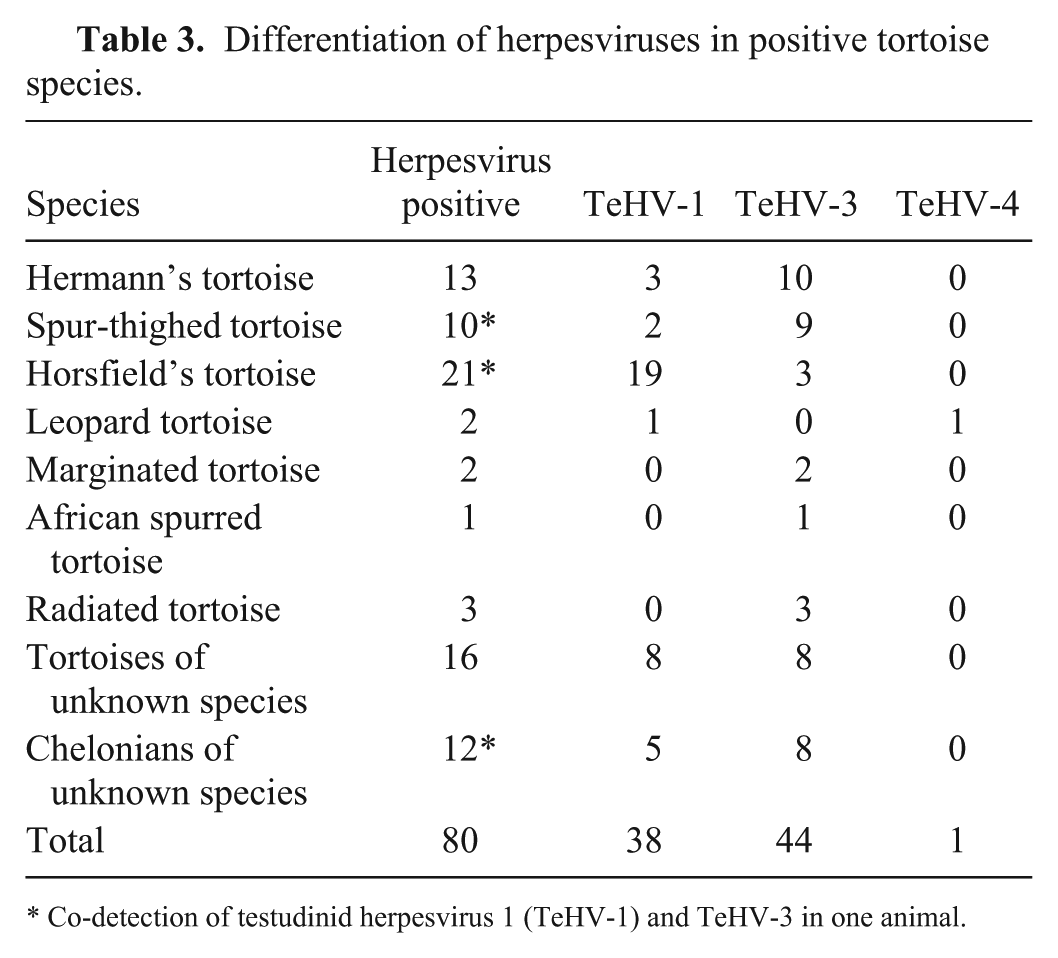
Co-detection of testudinid herpesvirus 1 (TeHV-1) and TeHV-3 in one animal.
All products from samples in which only 1 of the 3 PCRs used for herpesvirus detection was positive were sequenced. In addition, products from randomly chosen samples for each species, distributed throughout the study period, were also sequenced. Sixteen PCR products of TeHV-1–positive samples and 24 PCR products of TeHV-3–positive samples were sequenced. Sequences of PCR products obtained from TeHV-1–type viruses using the nested (n)PCR 71 were 100% identical to herpesviruses described previously (GenBank accession DQ343883.1). 44 Sequences from the PCR specific for TeHV-1 49 were also 100% identical to TeHV-1 sequences described previously (accession AB047545.1). 49 Sequences from TeHV-3 PCR products obtained using the nPCR 71 were 97–100% identical to herpesviruses described previously (accession KR363629.1). 20 Sequences from the PCR specific for TeHV-3 68 were 98–100% identical to TeHV-3 sequences described previously (accession KM924292.1). 20 Sequence alignment of the TeHV-4 PCR product showed 99% sequence identity to the TeHV-4 described previously (accession GQ222415.1).5,36 Remarkably, the PCR product from the pond slider was obtained with the PCR specific for TeHV-1, 49 and sequencing showed 90% sequence identity to TeHV-1 described previously (accession AB047545.1) 49 and 75% sequence identity to TeHV-3 (accession KR363629.1). 20 No product was obtained from this sample using the nPCR protocol. 71 Although no significant differences were observed for TeHV-3, the prevalence for TeHV-1 was significantly higher in Horsfield’s tortoises compared to all other testudinid species (p = 0.0001).
Comparisons of the 3 PCRs for herpesvirus detection sometimes revealed inconsistent results. In 5 animals, the specific TeHV-3 PCR was positive, whereas no virus was detected using the pan-herpesviral nPCR. In 3 animals, the TeHV-1–specific PCR was positive, whereas the pan-herpesviral nPCR was negative. In all cases in which discrepancies were noted, the specific PCR products were sequenced. In the 8 cases in which 1 of the specific PCRs was positive, but the pan-herpesviral nPCR was not, the identity of the detected viruses was confirmed by sequencing. In several cases, only the pan-herpesviral nPCR was positive, whereas the TeHV-1– and TeHV-3–specific PCRs were negative. Sequencing of the products showed that the PCR products were identical to corresponding TeHV-1 sequences in 2 cases. Nonspecific products were repeatedly obtained using this nPCR. Sequencing of PCR products in these cases showed that these were specific either for other, non-chelonian herpesviruses or were nonspecific. Five of the products were specific for equid alphaherpesvirus 1 (EHV-1), 10 for equid gammaherpesvirus 2 (EHV-2), 2 for equid gammaherpesvirus 5 (EHV-5), 5 for asinine herpesvirus 5, 5 for felid alphaherpesvirus 1, and 1 for human alphaherpesvirus 1. For 40 of the amplified PCR products with the expected size of 200 bp, sequencing revealed no significant similarity to known herpesvirus sequences in GenBank. The majority of these sequences are somewhat similar to various yeast or fungal DNA sequences.
No ranaviruses were detected in any of the samples tested using the PCR for the detection of part of the ranaviral major capsid protein gene.39,43
Picornaviruses (topiviruses) were detected in 22 (2.2%, 95% CI: 1.4–3.3%) of 1,015 animals by PCR (Table 1), including 6 (4.5%) of 133 spur-thighed tortoises, 4 (2.1%) of 195 Hermann’s tortoises, 3 (3.6%) of 83 Horsfield’s tortoises, 1 (2.0%) of 51 leopard tortoises, 1 (0.5%) of 216 tortoises of unknown species, and 7 (5.3%) of 133 chelonians of unknown species. Sequencing of 16 of these PCR products showed that the products were 86–98% similar to previously described topiviruses (GenBank accessions KM873611.1–KM873615.1). 17 Given the small number of picornavirus-positive samples, meaningful statistical analyses were not possible.
Ferlaviruses were detected in 6 (0.6%, CI: 0.2–1.3%) of 1,015 chelonians by PCR (Table 1), including 1 (0.8%) of 133 spur-thighed tortoises, 1 (4.2%) of 24 African spurred tortoises, 1 (20.0%) of 5 black pond turtles, 1 (100%) of 1 Annam leaf turtle, and 2 (0.9%) of 216 tortoises of unknown species. Sequencing of all 6 PCR products showed that these were 100% similar to each other and 99% similar to previously described snake ferlaviruses (GenBank accession AF286043.1). Because of the small number of ferlavirus-positive animals, no statistical analysis was carried out.
Mycoplasma and herpesviruses were co-detected in 51 (5.0%; 95% CI: 3.8–6.6%) tortoises. Of these, mycoplasma and TeHV-1 were detected in 30 (3.0%) animals, including 17 (20.5%) Horsfield’s tortoises, 3 (1.5%) Hermann’s tortoises, 1 (0.8%) spur-thighed tortoise, 1 (2.0%) leopard tortoise, 4 (1.9%) tortoises of unknown species, and 4 (3.0%) chelonians of unknown species. Mycoplasma and TeHV-3 were detected in 23 (2.3%) animals, including 3 (3.6%) Horsfield’s tortoises, 8 (4.1%) Hermann’s tortoises, 5 (3.8%) spur-thighed tortoises, 1 (2.5%) marginated tortoise, 2 (0.9%) tortoises of unknown species, and 4 (3.0%) chelonians of unknown species. The chance of being co-infected with herpesviruses and mycoplasma was twice as high (OR = 2.2; p = 0.013) as being infected with only a single pathogen in chelonians. In Testudo spp., the chance of co-detections of herpesviruses and mycoplasma was 5 times higher than for single pathogen detections (OR = 5.1; p = 0.0001). The leopard tortoise with TeHV-4 was also positive for mycoplasma. 36 Mycoplasma and picornaviruses were co-detected in 12 (1.2%) animals, including 3 (3.6%) Horsfield’s tortoises, 3 (2.3%) spur-thighed tortoises, 1 (0.5%) Hermann’s tortoise, 1 (2.0%) leopard tortoise, and 4 (3.0%) chelonians of unknown species. Mycoplasma and PMV were found in 1 spur-thighed tortoise. Detection of >2 pathogens was also observed in individual cases. Mycoplasma, TeHV-1, and TeHV-3 were detected in 1 Horsfield’s tortoise, 1 spur-thighed tortoise, and 1 chelonian of unknown species. Mycoplasma, TeHV-3, and picornavirus were detected in 1 spur-thighed tortoise.
Of the positive mycoplasma samples, 403 (42.3% of all tested swabs) were swabs (mainly oral swabs), 23 (27.4%) were nasal washes, and 12 (46.2%) were tissues. For 23 mycoplasma-positive animals, 2 samples, swab and nasal washes, were submitted. From these, both swab and nasal wash were positive in 11 chelonians; 8 chelonians were only positive in the swab sample, and 4 chelonians were only positive in nasal washes. Herpesviruses were also detected mainly in swabs, with 72 (7.6%) positives, but also in 7 (26.9%) tissue samples, and 2 (2.4%) nasal washes. In 2 cases, a herpesvirus-positive swab was submitted with a negative nasal wash, whereas in another case the nasal wash was positive and the swab was negative. Picornaviruses were detected in 19 (2%) swabs, 2 of which were submitted with nasal washes, but only 1 (1.2%) nasal wash was also positive. Furthermore, picornaviruses were detected in 3 (11.5%) tissue samples. The 6 ferlavirus-positive samples were swabs (0.6%). Statistical analysis to determine the best sample material was not possible because swabs were the main sample material, whereas nasal washes and tissue represent just a small proportion of the samples tested.
Statistical analysis of the 4 countries with the most samples (Germany, United Kingdom, Spain, and Switzerland) showed that Mycoplasma spp. (p = 0.0005) and herpesviruses (p = 0.0256) were detected significantly more often in animals from Spain than from the other 3 countries (Table 4). Statistical analysis revealed that mycoplasma could be detected significantly more often in the fall (p = 0.0082), whereas herpesviruses were detected significantly less frequently in the fall (p = 0.023; Table 5). Therefore, co-detections of mycoplasma and herpesviruses are significantly less frequent (p < 0.0001) in the fall.
Chelonian pathogen detection in different European countries.
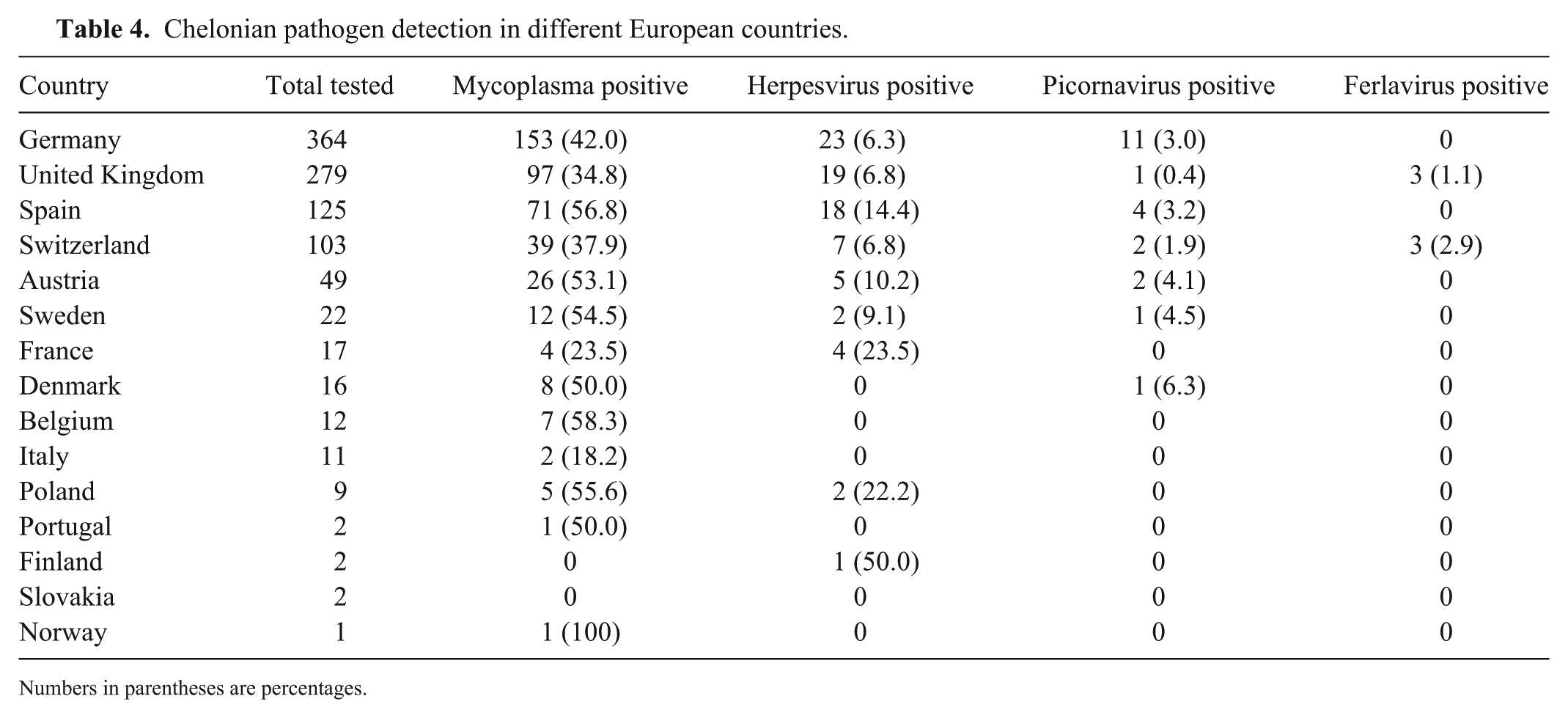
Numbers in parentheses are percentages.
Chelonian pathogen detection from the period of January 2014–June 2015 according to month.
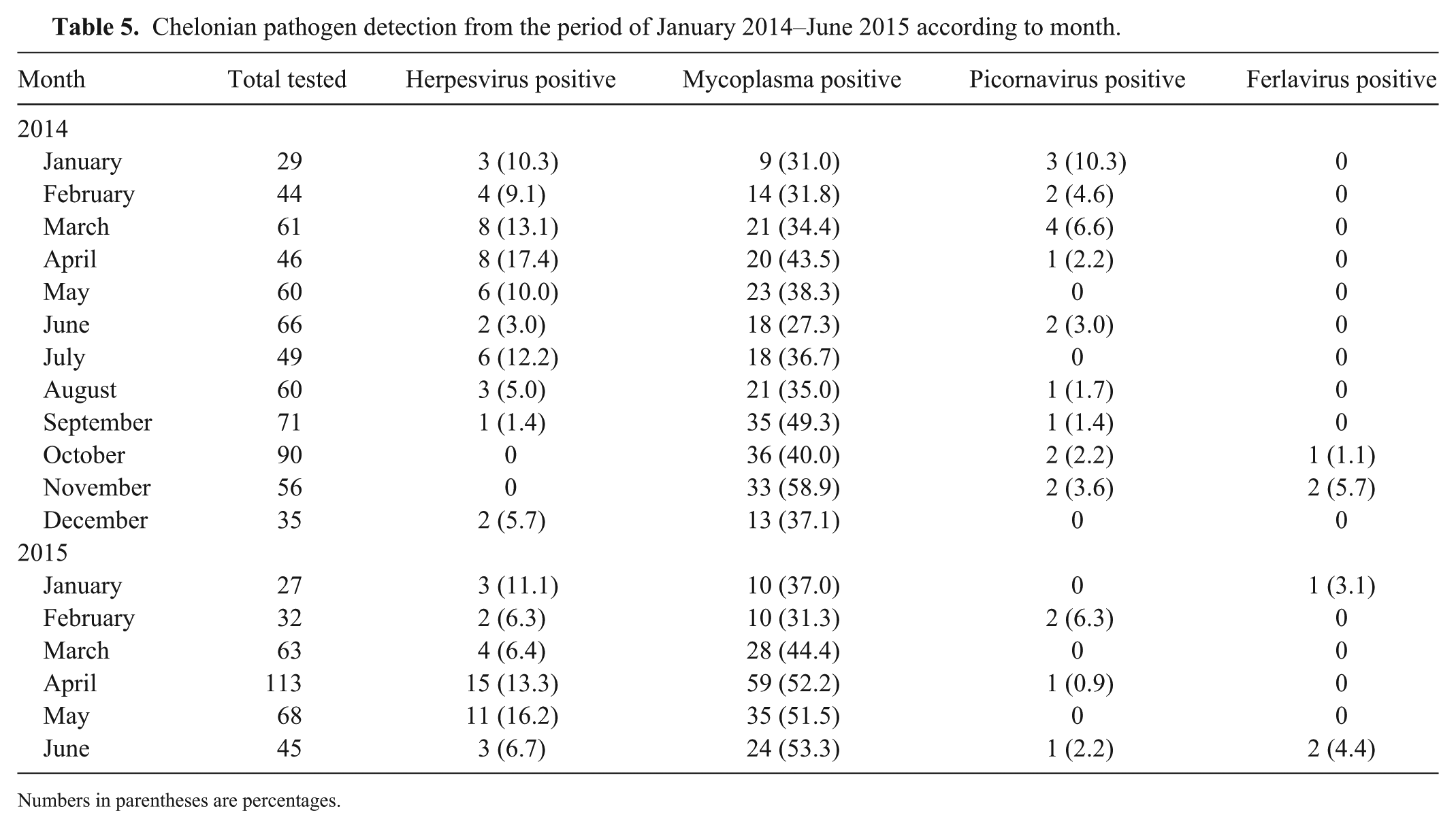
Numbers in parentheses are percentages.
Discussion
We found a very high incidence of Mycoplasma spp. in samples from chelonians in Europe, including 42.2% of the tortoises tested and 30.8% of the emydid turtles tested. Previous studies in tortoises (Testudo sp.), also based on PCR, have shown Mycoplasma spp. prevalences of 2.4% in tortoises (n = 329) from France and Morocco (Mathes KA. [Investigations on the occurrence of mycoplasma and herpesviruses in wild and captive Mediterranean tortoises (Testudo hermanni, Testudo graeca graeca and Testudo graeca ibera) in France and Morocco]. [Dissertation]. Justus-Liebig-Universität Gießen, Gießen, Germany, 2003), 6% in tortoises (n = 63) from Spain, 61 15.8% in tortoises (n = 146) from the United Kingdom, 65 and 37% in tortoises (n = 30) from Italy. 38 Studies in emydid turtles from the United States have shown Mycoplasma spp. prevalences of 30% in eastern box turtles (n = 23) from Virgina, 18 7.3–22% (depending on the region) in western pond turtles and red-eared sliders (n = 178) from California, 63 and 58.3% in different emydid turtles from the northeastern United States. 54 The high percentage of Mycoplasma spp. in our study could be because the submitted samples were derived from animals that were under veterinary care.
We found significant differences in the frequency of mycoplasma detection in some species. Horsfield’s tortoises, leopard tortoises, and Indian star tortoises were mycoplasma-positive significantly more often than other species; detection of Mycoplasma spp. in European tortoises was less common. In a previous study, most mycoplasma were also detected in Horsfield’s tortoises. 65 It has been suggested that Mycoplasma spp. belong to the commensal flora in some emydid species because of their high prevalence. 54 Therefore, some species could harbor mycoplasma and be a source of infection for other chelonians. 65 Mycoplasma are, however, generally very species specific, although there are numerous reports of mycoplasma infections outside of their natural hosts leading to more severe diseases.11,59 In one case in our study, Mycoplasma spp. was detected in a giant Asian pond turtle, belonging to the family Geoemydidae, which has not been reported previously, to our knowledge. The Mycoplasma spp. sequence found in the giant Asian pond turtle was 99% identical to previously published sequences from emydid turtles 54 and the emydid mycoplasma detected in our study and clearly clustered with these in phylogenetic analysis (data not shown). Further studies are needed to investigate the specificity and the impact on clinical signs of Mycoplasma spp. in different chelonian species.
Background information on the health status of mycoplasma-positive chelonians was only available for a few animals. Many of these had typical signs of URTD, although Mycoplasma spp. was not always detected in animals in which these clinical signs were reported. There are, however, contradictory reports on the prevalence of Mycoplasma spp. and the correlation between infection and clinical disease available in the literature. Although M. agassizii is an important cause of URTD in tortoises, 7 other studies have shown high prevalences (58.3%) of Mycoplasma spp. in apparently clinically healthy aquatic species, 54 and low prevalences (6%) of Mycoplasma spp. in tortoises with rhinitis–stomatitis complex have been described. 61 The differences in reports about infections with Mycoplasma spp. and their clinical impact could be the result of the chronic character of these infections. In tortoises with M. agassizii infections, clinical signs occur intermittently, and the severity of the disease varies.8,62 Subclinical disease and silent carriers are common. 28 Other factors (e.g., stress or other pathogens) may also increase clinical signs. 7 Several different strains of M. agassizii with different pathogenic potential have been described.8,28 It is also believed that clinical signs of URTD may be more severe after re-infection of previously infected tortoises. 9
False-negative results are possible in our study because the majority of submitted samples were swabs. Mycoplasma colonize the ventrolateral mucosal surface of the upper respiratory tract in tortoises, 8 thus oral swabs can result in false negatives. Samples that have been recommended for mycoplasma detection in previous studies include nasal lavages that were incubated in SP4 medium for 24–48 h in order to detect even small amounts of mycoplasma. 8 This is, however, much more time-consuming and costly and is therefore difficult to implement in laboratory practice. It is interesting to note that, in those cases in which both oral swabs and nasal washes were tested, the oral swabs appeared to be somewhat more sensitive, although testing of both oral swabs and nasal washes increased detection rates.
The percentage of herpesvirus-positive chelonians found in our study (8.0%) lies between previously reported incidences, with 0.3% in tortoises from France and Morocco (Mathes KA. [Investigations on the occurrence of mycoplasma and herpesviruses in wild and captive Mediterranean tortoises]), 8.2% in tortoises from the United Kingdom, 65 17% in tortoises from Belgium, 47 and 25% in tortoises from Spain. 61 Few studies have further categorized the herpesviruses detected in large numbers of tortoises in Europe. In 1 study, 11 herpesvirus isolates from tortoises were compared. Two were determined to be TeHV-1 (called tortoise herpesvirus type 2), whereas 9 were TeHV-3 (called tortoise herpesvirus type 1). 44 In another study in Spain, TeHV-1 (called ChHV type II) was detected in 3% of the animals tested, whereas TeHV-3 (called ChHV type I) was detected in 24% of the animals. 61 A seroprevalence study of TeHV-1 and TeHV-3 in wild-caught spur-thighed tortoises in Turkey showed prevalences of 5.5% and 37%, respectively. 41 Although according to the previous studies it seems that TeHV-3 occurs more often than TeHV-1, in our study the incidence of TeHV-1 (46.9%) and TeHV-3 (54.3%) was almost equal. TeHV-1 and -3 also appear to differ in species affected. Previous studies have detected TeHV-1 most commonly in Horsfield’s tortoises.40,44,69 This was also true in our study. The apparent increase in TeHV-1 detections could therefore hypothetically be influenced by the pet trade and changes in popularity of specific species (in this case, Horsfield’s tortoises) as pets. There is a lack of information about the clinical impact of the different TeHVs in different species. It is believed that TeHV-3 is associated with higher morbidity and mortality in Hermann’s and Horsfield’s tortoises than in spur-thighed tortoises,40,42 and that TeHV-1 has a relatively low morbidity and mortality in Horsfield’s tortoises showing less severe disease, which could also explain the high detection rate of TeHV-1 in this species,66,70 and might contribute to the distribution of this strain in pet tortoises.
TeHV-4 has been detected previously only once in a bowsprit tortoise in the United States. In our study, TeHV-4 was detected for the first time in a leopard tortoise and for the first time in Europe. The leopard tortoise belonged to a mixed species collection of tortoises and showed respiratory signs. This animal was described previously in a case report. 36
The herpesvirus detected in the pond slider seems to be different from testudinid herpesviruses, although it was detected using the TeHV-1–specific PCR. The pan-herpesvirus nPCR was negative for this sample. Unfortunately, there was not enough DNA from this sample to repeat the PCRs and sequencing for confirmation. It is possible that this PCR product is a novel herpesvirus, which is not surprising considering the numerous descriptions of novel herpesviruses in freshwater turtles in recent years.12,35,46,55,56,64,72
TeHV-2 is believed to be endemic in Californian desert tortoises in the United States. 51 In our study, TeHV-2 could not be detected, likely because only chelonians from Europe were tested and Californian desert tortoises were not included.
Despite the apparent preference of TeHV genotypes for different species, it is important to note that different TeHV genotypes can infect various species. Resistant species can serve as carriers and can transmit TeHV to more susceptible species, which can result in fatal disease. It is therefore recommended to avoid mixing species within collections. 48 As in mammals, herpesviruses in tortoises can cause latent infections. Virus may not be detectable in live animals in these cases in the absence of viral shedding. Given that the majority of samples tested in this study were from live animals, it is possible that the real prevalence of herpesviruses in chelonians could be higher. However, the sampling of animals that were under veterinary care likely led to a sampling bias more in favor of infected animals.
Three different PCRs were used in our study for the detection of herpesviruses: 2 PCRs specific for TeHV-1 49 and TeHV-3, 68 and a pan-herpesviral nPCR 71 that is able to detect all known TeHVs as well as others. The pan-herpesviral nPCR can be a useful tool in the detection of herpesviruses in exotic animals, because it is capable of detecting a wide range of herpesviruses, including previously undescribed viruses. However, this can also lead to a lack of sensitivity and/or specificity. In our study, the pan-herpesviral nPCR did not detect virus in 8 cases in which one of the other PCRs used did. It also resulted in products that were not specific in 68 cases. These nonspecific results were either non-chelonian herpesviruses or were nonspecific. We do not know where the non-chelonian herpesviruses originated; most likely, these are contaminants from the tortoises’ surroundings or the veterinary surgeries in which sampling was carried out. Contamination in the laboratory during processing is also possible. To reduce the risk of contamination in the laboratory, only 20–40 samples were run per day and each run included negative controls after every tenth sample. At no time was there a run with a conspicuously high number of positive samples, and negative controls were never positive. We recommend sequencing of all products obtained only with the pan-herpesviral nPCR. 71 Previous studies have also reported that the pan-herpesviral nPCR was not very specific when used with chelonian samples. 51 It is nevertheless useful when testing chelonian samples to use several different PCRs in order to increase the chances of detecting a variety of herpesviruses.
No positive results were obtained from any of the samples tested using a PCR for the detection of a portion of the major capsid protein gene of ranaviruses.39,43 A positive control was included in every PCR run in order to verify the functionality of the PCR. The positive control was obtained from a cell culture isolate, and ranavirus specificity of the product was repeatedly confirmed by sequencing. Furthermore, an internal control to detect and exclude inhibited samples was run for every sample. There are different possible explanations for the lack of positive ranavirus samples in our study. The sensitivity of the conventional PCR used could have been insufficient for detection of small amounts of viral DNA. However, of the published PCRs for the detection of ranaviral DNA, the one used has been shown to detect the widest range of genetically distinct ranaviruses. This was considered an important criterion based on genetic diversity in ranaviruses in chelonians in Europe reported previously. 67 In a previous study, tissues from different organs were more suitable for the detection of ranavirus by PCR than oral swabs, probably because of the higher virus load in organs than in exfoliated epithelial cells obtained with a swab (Uhlenbrok C. [Detection of ranavirus infections in tortoises and characterization of virus isolates]. [Dissertation]. Justus-Liebig-Universität Gießen, Gießen, Germany, 2010). In our study, tissue samples were only tested from 26 animals; the majority of samples were oral swabs or nasal washes. In a study of free-ranging eastern box turtles, it was hypothesized that low detection rates could be caused by the association of ranaviral infections with high mortality, leading to underdiagnosis in wild animals. 3 In captive chelonians, low detection rates could be the result of the reluctance of owners to spend money on testing of dead animals. It is also possible that ranaviral infections in chelonians occur rhythmically, and that a longer screening period would be necessary to determine a pattern of infection. As well, chelonians may shed the virus for only a short time.
Topiviruses were detected in several species in which similar viruses have been described using the PCR described in our study. Infections with topiviruses can be associated with the same clinical signs as infections with herpesviruses or mycoplasma, so although the incidence of topiviruses seems to be low, topiviruses should be considered as a differential etiology for chelonians with stomatitis, rhinitis and/or conjunctivitis. Furthermore, animals with a soft carapace, enteritis, ascites, or nephritis should be tested for topiviruses.
Very little is known about ferlaviral infections in chelonians, and infections have only been described in Hermann’s tortoises, 45 spur-thighed tortoises (Zangger N, et al. [Viral dermatitis in the spur-thighed (Testudo graeca) and Hermann’s (Testudo hermanni) tortoises in Switzerland]), and a leopard tortoise. 58 In another study, antibodies against the 4 known genogroups of ferlaviruses (A, B, C, and tortoise) have been detected in different tortoise species with prevalences of 1–5%. 60 Our study expands the list of chelonian species in which ferlaviruses have been detected to include African spurred tortoises, black pond turtles, and Annam leaf turtles. The first species belongs to the family Testudinidae; the last 2 species belong to the family Geoemydidae. The sequences of the PCR products from all of the ferlavirus-positive tortoises were identical and 100% identical to a genotype B squamate ferlavirus,45,58 increasing the evidence available that these viruses are not species specific and can be transmitted between different orders of reptiles. Ferlaviruses were only detected in samples submitted from the United Kingdom and Switzerland. This is most likely an incidental finding, especially given that the prevalence of these viruses in chelonians appears to be very low. Nevertheless, it would be useful if more attention were given to this pathogen in order to prevent spread within chelonian collections and between chelonians and squamate reptiles.
For both of the RNA viruses detected in our study (topiviruses and ferlaviruses), it is possible that the instability of isolated RNA could have led to a reduced rate of detection. RNA degradation over time can certainly be a problem, and false-negative results are possible. However, in our experience, degradation of RNA within a storage period of 6 mo is negligible.
Co-detections of several different pathogens have been described previously in chelonians. In previous studies, detections of herpesviruses and Mycoplasma spp. in tortoises have been described with incidences of 1.37 65 –3%. 61 Serologic evidence of co-infections with TeHV-1 and TeHV-3 has been found in spur-thighed tortoises in Turkey. 41 Mycoplasma spp. were also detected in all 3 chelonians with double herpesvirus detection (TeHV-1 and TeHV-3). In our study, 5% of all animals were positive for mycoplasma and herpesviruses, and it could be shown that co-detections are more likely than detection of a single pathogen. Reasons for the high probability of co-detections could be the synergistic effect of Mycoplasma spp. and herpesviruses causing more severe disease, or that chronic diseases such as mycoplasmosis can reduce the immune response and lead to an increased susceptibility to further infections. 65 In a previous study on the detection of testudinid intranuclear coccidia (TINC) by real-time PCR in the same group of 1,011 animals with 27 (2.7%) TINC-positive tortoises, 37 37% of the TINC-positive animals were positive for Mycoplasma spp. and 11% were positive for TeHV-3. However, no significant correlation was found between TINC and mycoplasma detection or TeHV-3 detection.
We tested oral swabs and nasal washes most commonly, which is appropriate for minimally invasive samples for the detection of pathogens from the respiratory tract. In those cases in which tissue samples were tested, these were generally pooled samples from different organs, mostly including tongue, lung, liver, intestine, and kidney to represent the most important organ systems. The probability of false-negative results because of inappropriate tissue samples is therefore low. The high percentage of positive samples in tissues could be because autopsied animals were suspected to be infected with a certain pathogen and only obvious lesions were tested. In 9 of the animals included in our study, only blood was available, and these blood samples were all negative for all of the pathogens. Blood samples are unlikely to be appropriate samples for the detection of respiratory pathogens, given the likely limited duration or lack of viremia or bacteremia. False negatives are therefore to be expected if only blood samples are tested.
Mycoplasma and herpesviruses were detected significantly more frequently in animals from Spain than from other countries. However, this may be at least partially the result of sampling bias, and only 4 countries were compared given the small sample size in most other countries.
We detected mycoplasma most frequently in the fall, whereas herpesvirus detections in fall were the lowest. However, statistical analysis was only performed for an interval of 1 y. Comparison of at least 2 y would be valuable in order to draw reliable conclusions on the seasonality of mycoplasma and herpesvirus detections. Tortoises with herpesviral infections typically show clinical signs especially in the spring or late fall. 53
Footnotes
Declaration of conflicting interests
E Kolesnik and RE Marschang are employed by a commercial laboratory offering testing services for veterinarians. This employment did not influence study design, interpretation, or publication preparation.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
