Abstract
We describe the histopathologic, immunohistochemical, and molecular features of a case of meningoencephalitis in a Thomson’s gazelle (Eudorcas thomsonii) naturally infected with zebra-borne equid herpesvirus 1 (EHV-1) and the implications for the molecular detection of zebra-borne EHV-1. A 4-y-old female Thomson’s gazelle was submitted for postmortem examination; no gross abnormalities were noted except for meningeal congestion. Microscopic evaluation demonstrated multifocal nonsuppurative meningoencephalitis with intranuclear eosinophilic and amphophilic inclusion bodies and EHV-9 antigen in neurons. PCR demonstrated the presence of a herpesvirus with a nucleotide sequence 99–100% identical to the corresponding sequences of zebra-borne EHV-1 and of EHV-9 strains. To determine whether EHV-1 or EHV-9 was involved, a PCR with a specific primer set for EHV-9 ORF59/60 was used. The sequence was identical to that of 3 recognized zebra-borne EHV-1 strains and 91% similar to that of EHV-9. This isolate was designated as strain LM2014. The partial glycoprotein G (gG) gene sequence of LM2014 was also identical to the sequence of 2 zebra-borne EHV-1 strains (T-529 isolated from an onager, 94-137 from a Thomson’s gazelle). The histologic lesions of encephalitis and antigen localization in this gazelle indicate prominent viral neurotropism, and lesions were very similar to those seen in EHV-1– and EHV-9–infected non-equid species. Histologic lesions caused by EHV-9 and zebra-borne EHV-1 are therefore indistinguishable.
Keywords
Infections of horses with species Equid herpesvirus 1 (EHV-1; family Herpesviridae, subfamily Alphaherpesvirinae, genus Varicellovirus) occur worldwide and can cause abortion, neonatal mortality, respiratory disease, and myeloencephalopathy (equine herpesvirus myeloencephalopathy [EHM]). The lesions associated with EHM are endothelial necrosis, thrombosis, perivascular lymphocytic cuffing, and hemorrhage, resulting in ischemic central nervous system injury. The virus in EHM cases replicates in endothelium and vascular myocytes but not in neurons. A subset of neuropathogenic EHV-1 strains associated with EHM have been shown to have a single mutation of the ORF30 gene at nucleotide position 2254 (G2254), which differentiate them from non-neuropathogenic genotypes (A2254) of EHV-1. 19 However, horses infected with the “non-neuropathogenic” strain of EHV-1 can also develop neurologic disease, indicating that other unidentified factors contribute to the onset of EHM.
Zebra-borne EHV-1 has been isolated from a Grevy’s zebra fetus, a Persian onager fetus, Thomson’s gazelles, polar bears, black bears, guinea pigs, and an Indian rhinoceros in association with neurologic disease and abortion.1,11,16,18,28,29 Three strains, 94-137 from a gazelle (Eudorcas thomsonii), T616 from a zebra (Equus grevyi), and T-529 from an onager (Equus hemionus onager), share 99% identities.12,16,18,29 These zebra-borne EHV-1 strains shared 98% and 95% identities with the horse-derived EHV-1 and EHV-9, respectively. 12 Zebra-borne EHV-1 also has a mutation at ORF30 similar to that seen in neuropathogenic EHV-1 in horses and therefore is considered a neuropathogenic herpesvirus. 12
Neurologic disorders caused by EHV-9 were first described in an outbreak of epizootic encephalitis of Thomson’s gazelles kept at a zoological garden in Japan.8,30 Initially the isolate was called gazelle herpesvirus 1, but was renamed EHV-9 based on its genetic homology to other members of the equid herpesvirus family. The histologic lesions in Thomson’s gazelles infected with EHV-9 are nonsuppurative meningoencephalitis with intranuclear inclusion bodies in neurons that are also observed in animals infected with neuropathogenic EHV-1.9,28,30
Herein we describe the histopathologic, immunohistochemical, and molecular features of a case of meningoencephalitis in a Thomson’s gazelle naturally infected with zebra-borne EHV-1 in North America, and include the molecular techniques recommended for the identification of zebra-borne EHV-1.
A 4-y-old female Thomson’s gazelle (Eudorcas thomsonii) housed in the Baton Rouge Zoo (Baton Rouge, Louisiana) suddenly developed lethargy, headshaking, and seizures that rapidly became more severe and frequent. Magnetic resonance imaging (MRI), complete blood count, chemistry panel, and cerebrospinal fluid analysis were performed. MRI revealed multifocal T2W hyperintense, T1W isointense, DWI hyperintense, FLAIR hyperintense, and non–contrast-enhancing lesions that were diffuse within the left olfactory bulb, a focal region of hyperintensity in the rostroventral aspect of the right and left frontal lobes, and a focal hyperintense lesion within the left thalamus (Supplemental Fig. 1). The T2W hyperintense region in the left thalamus was sharply marginated and wedge-shaped on a transverse DWI image, which also showed a focally extensive and ill-defined region of hyperintensity in the left parietal lobe. Multifocal hyperintensity regions were also noted in the temporal lobes, bilaterally, on a transverse FLAIR image. Cytologic evaluation of cerebrospinal fluid revealed nonsuppurative pleocytosis. No significant findings were revealed by the complete blood count and chemistry panel. A week after the onset of clinical signs, the animal was found dead in its habitat despite treatment and was submitted for postmortem examination. No rabies virus, bovine viral diarrhea virus, arboviruses, pseudorabies virus, bovine herpesvirus 1, or canine distemper virus was detected in several brain samples.
At autopsy, the Thomson’s gazelle was in thin body condition, and weighed 20 kg. The abomasum contained numerous Haemonchus nematodes. There was severe diffuse meningeal congestion. Selected organs and tissues were fixed in 10% neutral-buffered formalin and examined microscopically for processing for histologic examination using routine methods. Sections of the brain were obtained from various areas including cerebral cortex, hippocampus, thalamus, pons, medulla oblongata, cerebellum, and cervical spinal cord.
There was moderate multifocal nonsuppurative meningoencephalitis within the white and gray matter of the cerebral cortex, hippocampus, thalamus, midbrain, pons, medulla oblongata, and cervical spinal cord. The leptomeninges and Virchow–Robin spaces were expanded by moderate numbers of inflammatory cells, more frequently at the level of cerebral cortex, thalamus, and brainstem than in the hippocampus and cervical spinal cord, but not in the cerebellum. The inflammatory cells were predominantly small lymphocytes and a few macrophages. The gray matter of the cerebral cortex had multifocal-to-coalescing extensive neuronal necrosis with mild-to-moderate astrogliosis, microgliosis, and glial nodules. Necrotic neurons with intranuclear inclusion bodies were noted, particularly within the gray matter of the affected cerebral cortex. The inclusion bodies were either eosinophilic or amphophilic, indistinct and smudgy, filled the neuronal nuclei (diffuse-type inclusion body; Fig. 1), and were accompanied by chromatin margination. No neuronophagia was evident in any of the examined sections. Glial nodules and mild astrogliosis were more commonly observed within the white matter, which also had sparse dilated axons (spheroids). The brainstem often had random multifocal-to-coalescing, poorly demarcated malacic foci characterized by gitter cell infiltration, measuring 2 × 2 mm to 5 × 2 mm (Fig. 2). Within these areas there was also neuronal necrosis with rare intranuclear inclusion bodies, astrocytic gliosis, perivascular lymphocytic and histiocytic cuffing, and hyperemia. Here, the endothelial cells were swollen and hypertrophied; vasculitis, endothelial damage, or hemorrhage was not observed. Neuronal degeneration and intranuclear inclusion bodies within the degenerate neurons and glial cells were more commonly observed adjacent to the malacic areas that had prominent infiltration of gitter cells throughout the brainstem. Neuronal necrosis was most severe in the cerebral cortex of the frontal, piriform, temporal, and parietal lobes, thalamus, and brainstem, and less in the hippocampus and cervical spinal cord.
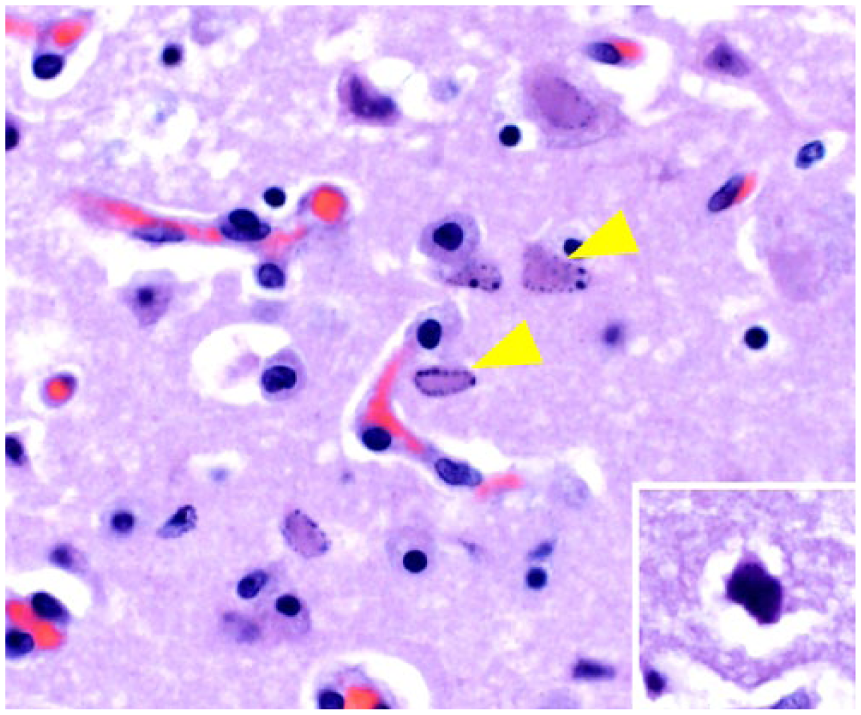
Intralesional necrotic neurons with intranuclear inclusion bodies (arrowheads) in the cerebral cortex of a Thomson’s gazelle (Eudorcas thomsonii). The inclusion bodies are generally eosinophilic, indistinct, smudgy, and fill the nuclei (diffuse-type inclusion body). Inset. Amphophilic diffuse-type inclusion body. H&E. 400×.
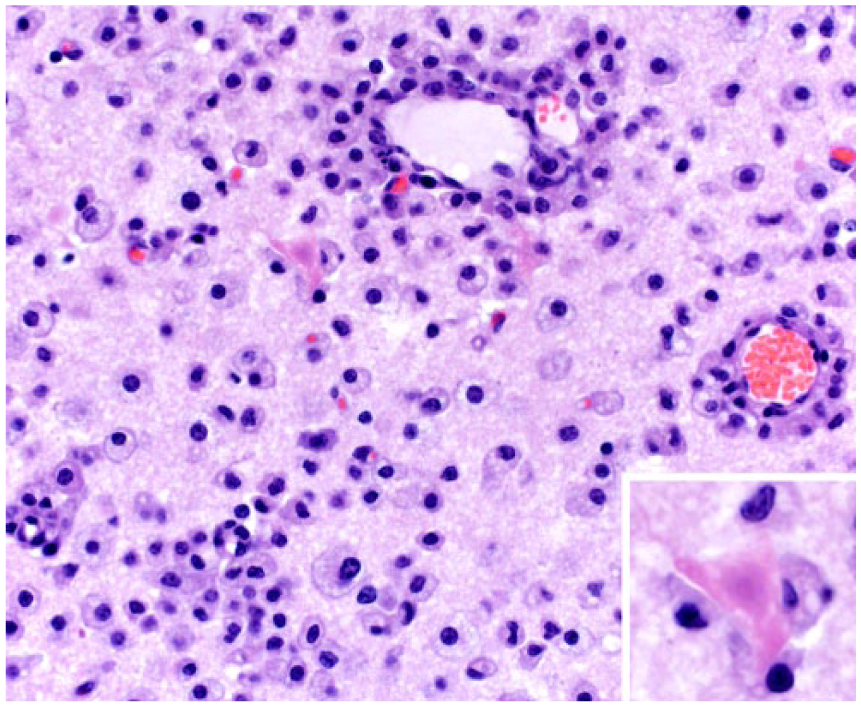
A focus of malacia in the brainstem of a Thomson’s gazelle is characterized by infiltration of the neuroparenchyma by large numbers of gitter cells with abundant foamy cytoplasm. These cells and lymphocytes also surround vessels. H&E. 200×. Inset. Necrotic neuron surrounded by gitter cells. 400×.
Brain sections including cerebral cortex, hippocampus, thalamus, pons, and medulla oblongata were sent to Gifu University in Japan for immunohistochemistry with a rabbit polyclonal serum specific for EHV-9 (1:800) by using the two-step polymer method (EnVision+ system HRP labelled polymer anti-rabbit, Dako, Tokyo, Japan). Positive control tissues for the anti–EHV-9 antibody were from a mouse experimentally infected with EHV-9. Additional sections were stained with murine monoclonal antibody 26A5 IgG2a isotype against the EHV-1 envelope glycoprotein C. 5 Equine fetal tissues containing lesions and EHV-1 were used as positive controls. This antibody also detected EHV-1 antigen within endothelium and myocytes of the blood vessels of equine central nervous system. Immunohistochemistry for EHV-9 revealed antigens in the nucleus, cytoplasm, and dendrites in degenerating and necrotic neurons, and occasionally in morphologically unremarkable neurons and glial cells in the gray matter and white matter of the cerebral cortex, thalamus, pons, and medulla oblongata (Fig. 3). Scattered EHV-9–labeled neurons were also confirmed within the malacic areas and adjacent areas. No antigen was observed in Gitter cells, in inflammatory cells surrounding vessels, and in endothelial cells. No EHV-1 antigen was observed.
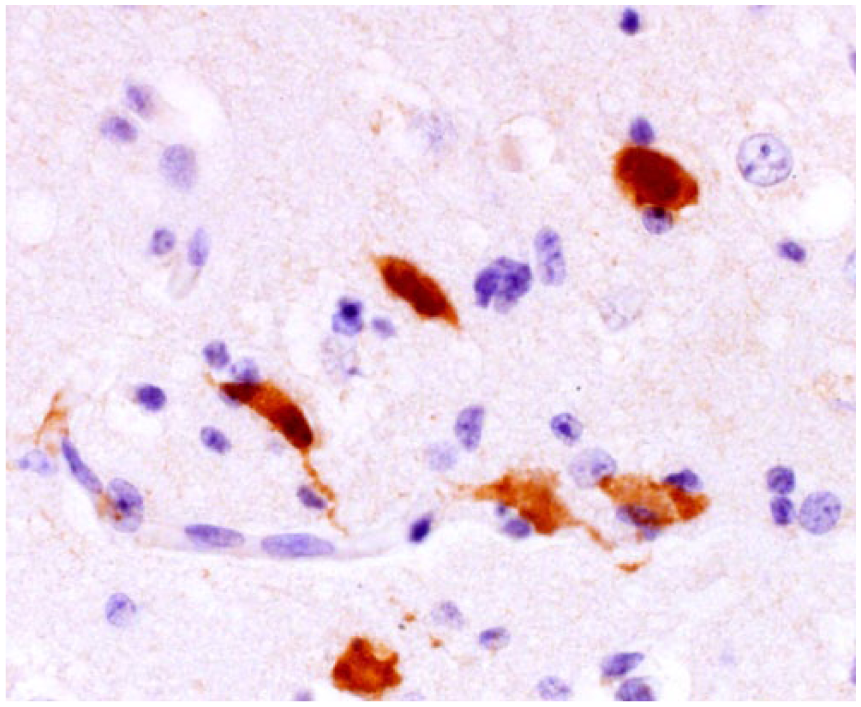
Anti–equid herpesvirus 9 immunoreactivity is noted within the cytoplasm, dendrites, and nuclei in the neurons in the cerebral cortex of a Thomson’s gazelle. Immunohistochemistry, counterstained with hematoxylin. 400×.
For PCR, DNA was extracted (QIAamp DNA FFPE tissue kit, Qiagen, Valencia, CA) from formalin-fixed, paraffin-embedded brain tissue. A PCR for the detection of the polymerase gene of herpesvirus was performed using the nested set of pan-herpesvirus primers according to a method described previously. 27 The amplicon derived from the generic herpesvirus PCR testing was sequenced. A unique 178-bp sequence was analyzed by BLAST against the GenBank database. 2 The nucleotide sequence was 99–100% identical to the corresponding partial DNA polymerase gene sequence of multiple EHV-1 and EHV-9 strains.
To determine whether EHV-1 or EHV-9 had infected the Thomson’s gazelle, a PCR assay that was reported to be specific for EHV-9 ORF59/60 detection was performed. 17 The expected size product of 679 bp was amplified from the sample DNA (Fig. 4). However, further analysis of both forward and reverse primer sequences revealed only minor mismatches with corresponding EHV-1 sequences that did not prevent the amplification of a neurotropic strain of EHV-1 in an assay specificity test conducted in our laboratory (data not shown). This unexpected cross-reactivity of the EHV-9 PCR assay to EHV-1 DNA necessitated follow-up sequencing of the product amplified from the case to verify the nature of the amplicon. A 571-bp sequence was derived and analyzed by BLAST against GenBank sequences and was found to be 100% similar to that of 3 recognized “zebra-borne” EHV-1 strains: T-529, 94-137, and T-616.16,18,29 The zebra and onager EHV-1 strains (T-529 and T-616) were isolated from aborted fetuses; strain 94-137 was isolated from a Thomson’s gazelle that died after an acute neurologic illness characterized by mild nonsuppurative meningoencephalitis. Moreover, the case sequence was 97% similar to that of several EHV-1 strains that originated from horses (Equus caballus), and 91% similar to that of EHV-9 strain P19.
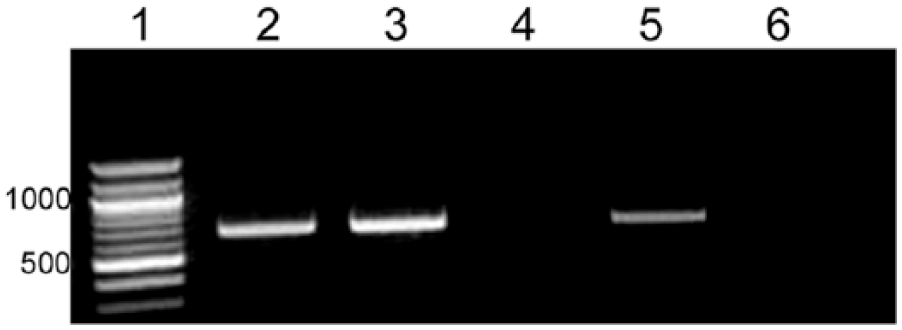
“Equid herpesvirus 9” (EHV-9) PCR amplifying a 679-bp region of the ORF59/60 gene. Lane 1: 100-bp DNA ladder; lanes 2, 3: gazelle sample; lane 4: negative extraction control; lane 5: EHV-9 positive control DNA; and lane 6: no template control (water).
To further substantiate the nature of the herpesvirus identified in this Thomson’s gazelle, designated henceforth as strain LM2014, partial sequencing of the herpesvirus glycoprotein G (gG) gene was performed. The gG gene has been commonly examined to assess the genetic relatedness of alphaherpesviruses.7,8,13,15 For primer design, the gG sequences of EHV-1 T-529 (GenBank accession KF644580) and EHV-9 P19 (accession AP010838) were aligned (Lasergene Biocomputing software, DNASTAR, Madison, WI). A 521-bp region at the 3’-end of the gene, including 33 bases after the stop codon, was identified as being a highly variable region between the 2 strains (nucleotide positions 126,375–126,895 for EHV-1 T-529, and 127,318–127,841 for EHV-9 P19). Specifically, strain T-529 has 46 scattered base mismatches and a three-base deletion compared to EHV-9 strain P19. PCR primers, gG forward primer (5’-GGATACCAGACCCAAACTCAT-3’) and gG reverse primer (5’-CCGGAATTGCCGCGATTTAGT-3), were designed from conserved sequences that flanked this 521-bp hypervariable region of gG that amplifies a 645-bp PCR target. The partial gG gene PCR was performed using a final primer concentration of 0.5 µm in a reaction volume of 50 µL, with the following cycling parameters: pre-denaturation at 94°C for 4 min, followed by 40 cycles of denaturation, annealing and elongation at 94°C for 30 s, 55°C for 30 s, and 72°C for 1 min, respectively. PCR products were visualized by agarose gel electrophoresis followed by ultraviolet transillumination. The amplicons were purified and submitted for sequencing at the Genomics Technology Support Facility at Michigan State University (East Lansing, Michigan). The 521-bp partial gG sequence of LM2014 was analyzed by BLAST against the GenBank database. The sequence showed 100% nucleotide sequence similarities to the corresponding sequences of onager- and Thomson’s gazelle–derived EHV-1 strains T-529 and 94-137, respectively, and 99% similarity to the zebra-derived EHV-1 strain T-616. Furthermore, the sequence was 97% similar to that of several horse-derived EHV-1 strains, but only 91% similar to that of EHV-9 strain P19. ClustalW alignment 26 of the 521-bp partial gG sequences of LM2014, zebra-borne EHV-1 strains T-529, 94-137, and T-616, horse-derived EHV-1 strains HH1 and 5586, EHV-9 strain P19, and EHV-4 as outgroup sequence, was performed with Lasergene software. Phylogenetic analysis of the aligned sequences was conducted with MEGA6, 24 using the neighbor-joining method,21,23 bootstrapping at 500 replicates, 6 and computation of evolutionary distances using the maximum composite likelihood method. 23 The phylogenetic tree in Figure 5 shows LM2014 falling within the clade formed by the 3 zebra-borne EHV-1 strains, distinct from the clade formed by the 2 horse-derived strains of EHV-1. All EHV-1 strains form a cluster that is clearly distinct from EHV-9.
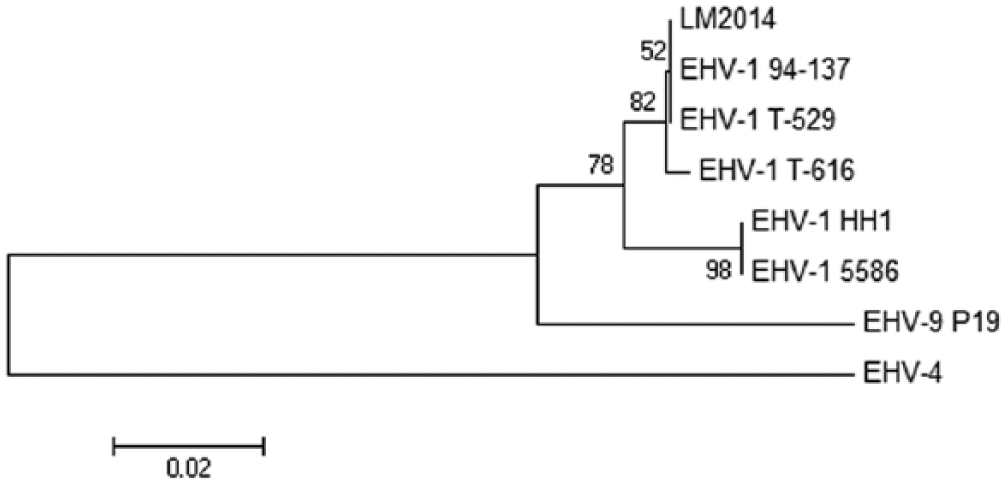
Phylogenetic tree based on partial nucleotide sequences of the gG genes of LM2014 (this study), zebra-borne EHV-1 strains 94-137 (GenBank accession KF644575), T-529 (KF644580), and T-616 (KF644574), horse-derived EHV-1 strains HH1 (AB992258) and 5586 (AP012321), EHV-9 strain P19 (AP010838), and EHV-4 (S44796) as the outgroup sequence.
LM2014 was also tested for the presence or absence of the EHV-1–specific neuropathogenicity marker characterized by a non-synonymous nucleotide substitution (A-to-G) at a specific position in the polymerase gene (ORF30). 22 Similar to the reported findings for all 3 zebra-born EHV-1 strains (T-529, 94-137, and T-616), 12 LM2014 was found to bear that same marker.
The sequence information derived from partial sequencing of ORF59/60 and a hypervariable region of the gG gene of the Thomson’s gazelle–derived herpesvirus, LM2014, indicate that this virus is zebra-borne EHV-1–like and not EHV-9. This finding is further confirmed by phylogenetic analysis of the hypervariable region of the gG gene. The zebra-borne EHV-1 strains are known to form a group of EHV-1 viruses, closely similar but distinct from horse-derived type EHV-1 strains.1,10–12,15,17 They are sometimes referred to as “variants” or a “subtype” of horse-derived EHV-1. The full genome sequences of the 3 zebra-borne EHV-1 strains have been determined and compared to the full genome sequences of horse-derived type 1 EHVs and of EHV-9. 12 The zebra-borne EHV-1 strains shared 99% identities with each other, and 98% and 95% identities with horse-derived EHV-1 and EHV-9, respectively. The phylogenetic tree derived from the full genome sequence analysis 12 mirrors the pattern of clades derived in the analysis of the hypervariable region of the gG gene (Fig. 5).
The histologic lesions in the brain of the Thomson’s gazelle in our study were characterized by nonsuppurative meningoencephalitis with intranuclear inclusion bodies in degenerate and necrotic neurons. The distribution of the neuronal degeneration and necrosis with intranuclear inclusion bodies was similar to that previously described in Thomson’s gazelles and a polar bear naturally infected with EHV-9 and other animals experimentally infected with EHV-9 (Table 1).4,9,30 Similar lesions that locally affected the olfactory bulb and the cerebral cortex of the frontal lobe were also observed in outbreaks of neuropathogenic EHV-1 infections of black bears, Thomson’s gazelle, and guinea pigs from 2 zoological collections in Germany (Table 1). 28 The isolates from these cases were genetically close to zebra-borne strains T-529 and T-616. 28 Thus, in several non-equid species, the neurologic disorders caused by both EHV-9 and zebra-borne EHV-1 are characterized by nonsuppurative meningoencephalitis with neuronal necrosis and intranuclear inclusion bodies. Therefore, differentiation between these viral infections should not be solely based on histopathology. Interestingly, other neuropathogenic zebra-borne EHV-1 infections in a Thomson’s gazelle, a polar bear, and hamsters were characterized by nonsuppurative meningoencephalitis with no viral inclusions (Table 1).11,15,16 This variation in neuropathologic findings between several zebra-borne EHV-1 infections might be associated with the virulence characteristics of the specific viral strain, the host’s immune status, and the animal species affected. Horse-derived EHV-1 causing EHM is vasculotropic and causes endothelial necrosis, thrombosis, perivascular lymphocytic cuffing, and hemorrhage without intranuclear inclusion bodies and neuron localization. Similar to animals naturally infected with EHV-9 and zebra-borne EHV-1, no vascular lesions were evident in the brain and cervical spinal cord of the present Thomson’s gazelle.4,9,28,30
Signalment, clinical information, and distribution of lesions in animals naturally and experimentally infected with EHV-9 and neuropathogenic zebra-borne equid herpesvirus 1 (EHV-1).
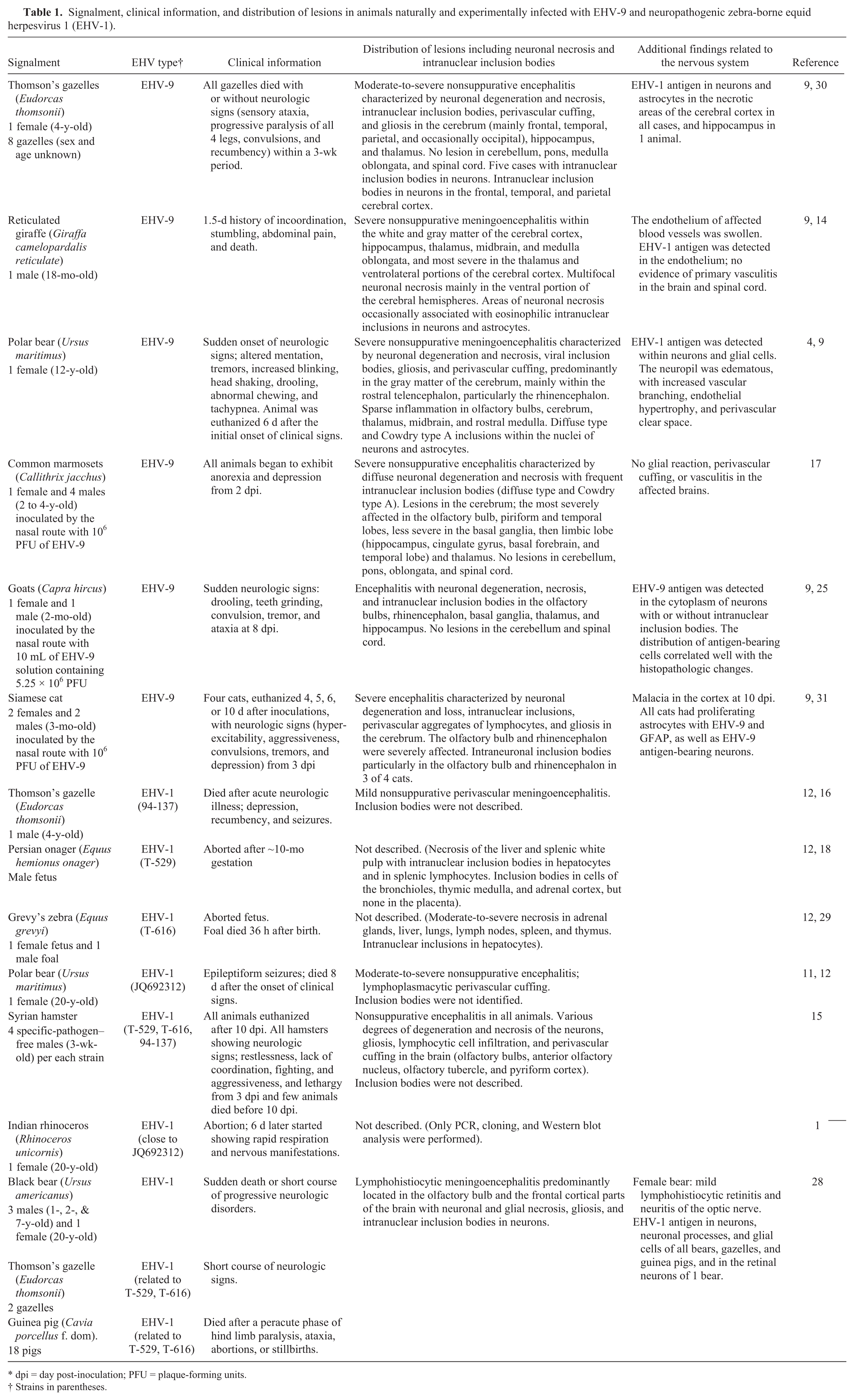
dpi = day post-inoculation; PFU = plaque-forming units.
Strains in parentheses.
EHV-9–labeled neurons and glial cells were observed particularly in the gray matter and white matter of the cerebral cortex and in the malacic areas that had significant infiltration of gitter cells and their adjacent areas throughout the brainstem. EHV-9–labeled neurons were more often observed in the areas of the gray matter that had extensive neuronal necrosis with inclusion bodies. Although neuronophagia was not evident in the examined sections of our case, immunohistochemical stain with anti–EHV-9 antibody revealed viral neurotropism and not endotheliotropism. Although we did not see evidence of vascular damage in any of the examined sections, considering the significant infiltration of gitter cells and neuronal necrosis observed in the brainstem, ischemic damage could not be ruled out completely.3,20 Therefore, we believe that the pathogenesis of these malacic lesions characterized by significant infiltration of gitter cells could be the result of both ischemic and viral cytopathic effect.
Our study indicated that the anti–EHV-9-specific polyclonal antibody can cross-react with zebra-borne EHV-1, whereas no EHV-1 antigen was detected by a specific monoclonal anti–horse-derived EHV-1 envelope glycoprotein C antibody. This could be explained by the much broader antigenic specificity of the polyclonal anti–EHV-9 versus the single epitope detection limits of the EHV-1–specific monoclonal antibody used in our study. Further evaluation with polyclonal anti–horse-derived EHV-1 antibodies may clarify this aspect. The neuronal viral localization also indicated strong neurotropism of this zebra-borne EHV-1 strain LM2014 instead of the vasculotropism seen in EHM caused by horse-derived EHV-1.
The pathologic findings of the zebra-borne EHV-1 meningoencephalitis in this Thomson’s gazelle indicate strong viral neurotropism with neuronal necrosis and intranuclear inclusion bodies similar to that seen in EHV-9 and other zebra-borne EHV-1–related neurologic disease in non-equid species. Vasculotropism, as seen with EHM associated with horse-derived EHV-1 infections, was not observed in our case. These histopathologic differences might be the result of several factors, including the species of the affected animals and the viral strains. In order to accurately detect zebra-borne EHV-1 infection and distinguish it from EHV-9 infection, it is therefore necessary to combine histologic, immunohistochemical, and molecular techniques.
Footnotes
Declaration of conflicting interests
The authors declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The authors received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
