Abstract
Despite the immunologic protection associated with routine vaccination protocols, Canine distemper virus (CDV) remains an important pathogen of dogs. Antemortem diagnosis of systemic CDV infection may be made by reverse transcription polymerase chain reaction (RT-PCR) and/or immunohistochemical testing for CDV antigen; central nervous system infection often requires postmortem confirmation via histopathology and immunohistochemistry. An 8-month-old intact male French Bulldog previously vaccinated for CDV presented with multifocal neurologic signs. Based on clinical and postmortem findings, the dog's disease was categorized as a meningoencephalitis of unknown etiology. Broadly reactive, pan-paramyxovirus RT-PCR using consensus-degenerate hybrid oligonucleotide primers, combined with sequence analysis, identified CDV amplicons in the dog's brain. Immunohistochemistry confirmed the presence of CDV antigens, and a specific CDV RT-PCR based on the phosphoprotein gene identified a wild-type versus vaccinal virus strain. This case illustrates the utility of broadly reactive PCR and sequence analysis for the identification of pathogens in diseases with unknown etiology.
Keywords
Canine distemper virus (CDV; family Paramyxoviridae, subfamily Paramyxovirinae, genus Morbillivirus) causes multisystemic disease in dogs. Infection is initiated in lymphoid tissue of the oropharynx, with subsequent spread to the respiratory, alimentary, urogenital, and central nervous systems. Central nervous system (CNS) lesions generally consist of varying degrees of lymphoplasmacytic inflammation, demyelination, and necrosis. 21
Although histopathology and immunohistochemistry (IHC) typically provide postmortem confirmation of CNS CDV infection, 15,21 antemortem diagnosis of CNS CDV infection is more challenging. Because cerebrospinal fluid (CSF) and magnetic resonance imaging (MRI) findings are highly variable with CDV infection, 2 diagnostic techniques with greater specificity must be used. Immunohistochemical testing for CDV antigen on biopsies of nasal mucosa, footpad epithelium, and the haired skin of the dorsal neck has been reported to be a relatively sensitive and specific test. 8 Similarly, reverse transcription polymerase chain reaction (RT-PCR) applied to RNA extracted from whole blood, urine, CSF, tonsilar, or conjunctival specimens are sensitive and specific assays. 1,5,6,10
The current investigators have reported a panel of 4 seminested RT-PCR assays that use consensus, degenerate primers for the diagnosis of paramyxovirus infections. 22 An advantage of pan-viral PCR is that it does not require individual testing for all known members of a viral family. Moreover, this technique has the potential to identify unsuspected or novel viruses affecting both people and animals.
An 8-month-old intact male French Bulldog was referred to the Texas A&M University Veterinary Teaching Hospital (College Station, TX) with a 7-day history of generalized seizures, circling, and falling to the right side. The dog had been vaccinated against infection with CDV, Canine adenovirus, Leptospira spp., Canine parvovirus, and Canine parainfluenza virus 3 months prior to referral. Based on the neurologic abnormalities, a multifocal neuroanatomic localization was made, with the cerebrum or thalamus and the hindbrain affected.
Magnetic resonance imaging of the brain was performed with a 1.0-T system a and an extremity coil. Multifocal, nonenhancing lesions were distributed asymmetrically in the cortical gray and subcortical white matter of the parietal, temporal, and occipital lobes of the cerebrum, as well as in the hypothalamus and the brainstem from the mesencephalon to the myelencephalon (Fig. 1, inset). Cerebrospinal fluid findings were moderate, mononuclear pleocytosis, with a total nucleated cell count of 103 cells/μl (reference [ref.] range: <5 cells/μl), 2 red blood cells (ref. range: <5 red blood cells/μl), negative pandy, and a microprotein of 35 mg/dl (ref. range: <30 mg/dl). A 100 nucleated cell differential included 71% large mononuclear cells, 27% lymphocytes, and 2% nondegenerate neutrophils. The large mononuclear cells were rarely vacuolated and contained a prominent Golgi zone around the nucleus. No microorganisms or neoplastic cells were identified.
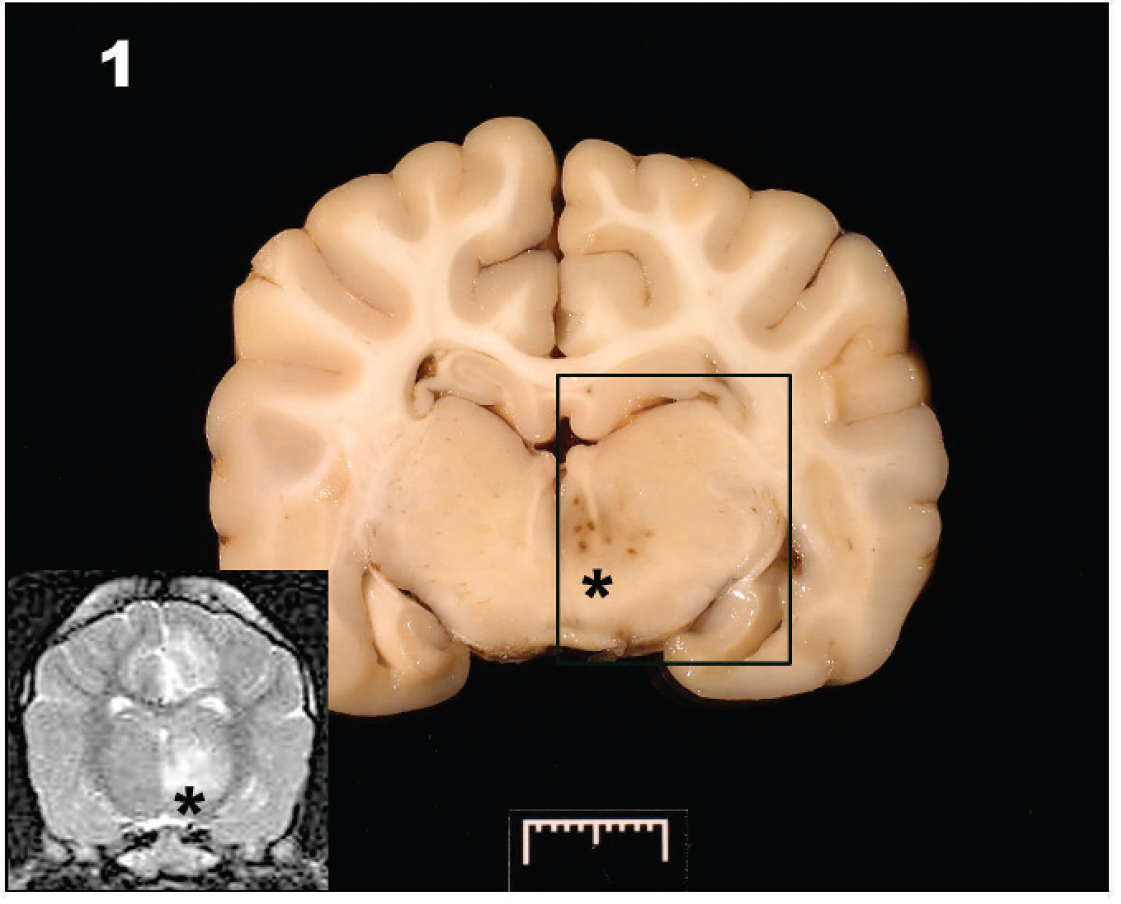
A discrete area of dark discoloration (*) is evident in the left thalamus. Inset: T2-weighted magnetic resonance imagery at the same level shows the hyperintense lesion (*) in the left thalamus and an additional hyperintense lesion associated with left cingulate gyrus.
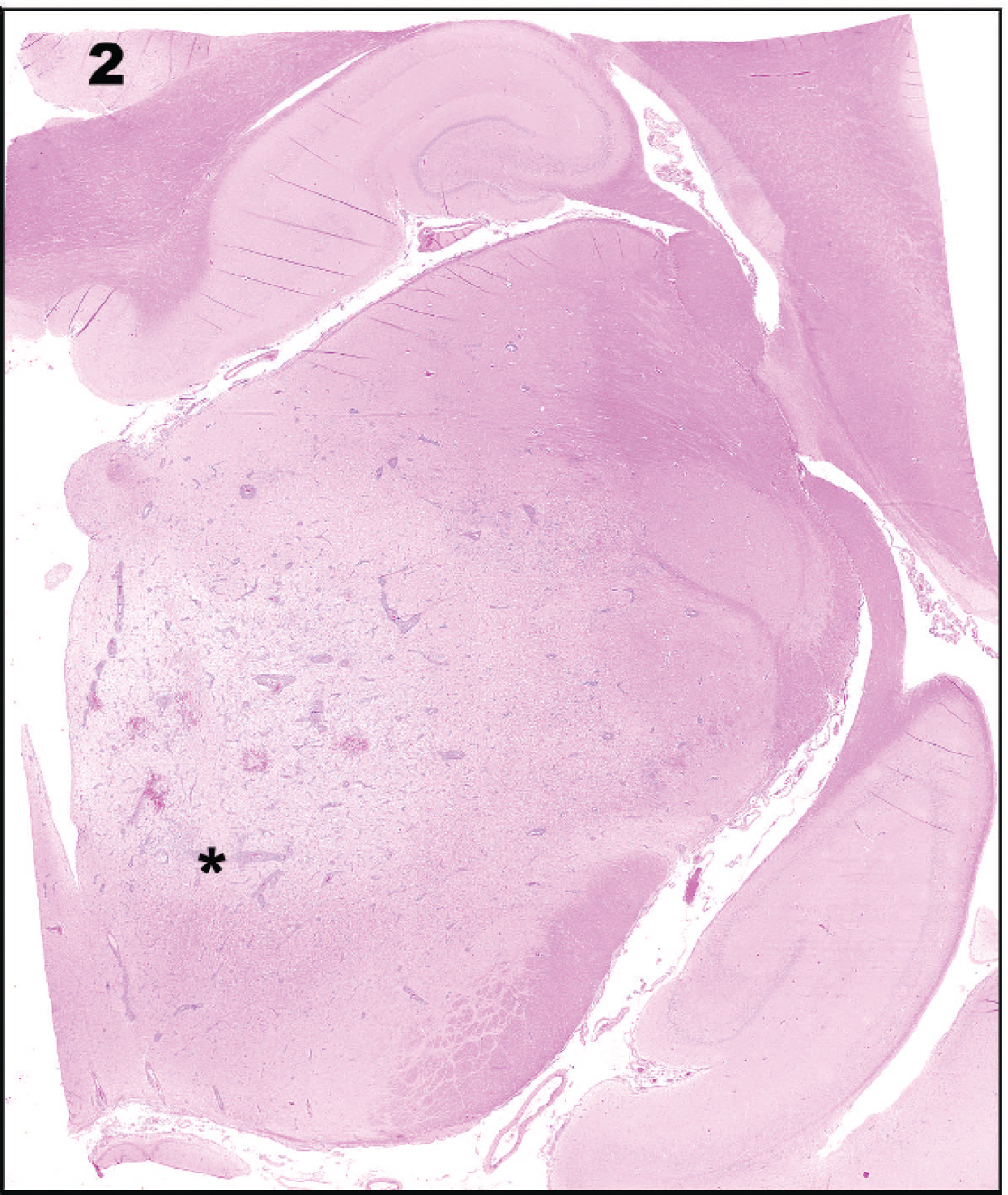
Histologic examination of the gross lesion (rectangle in Fig. 1 denotes region of this image) in the thalamus reveals an area of severe inflammation. Hematoxylin and eosin.
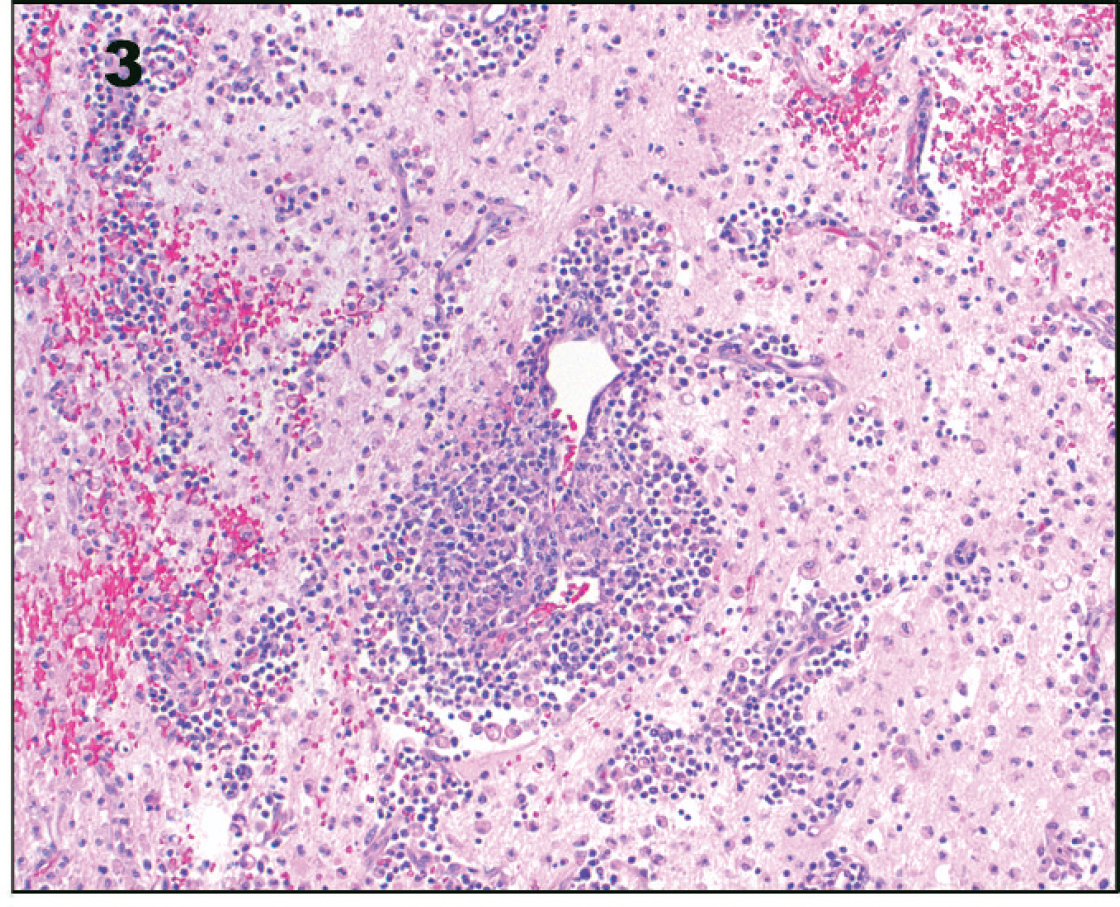
At higher magnification, prominent cuffs of lymphocytes, macrophages, and plasma cells surround blood vessels. The parenchyma is edematous with necrosis, multifocal hemorrhage, and numerous gitter cell macrophages. Hematoxylin and eosin. 10× magnification.
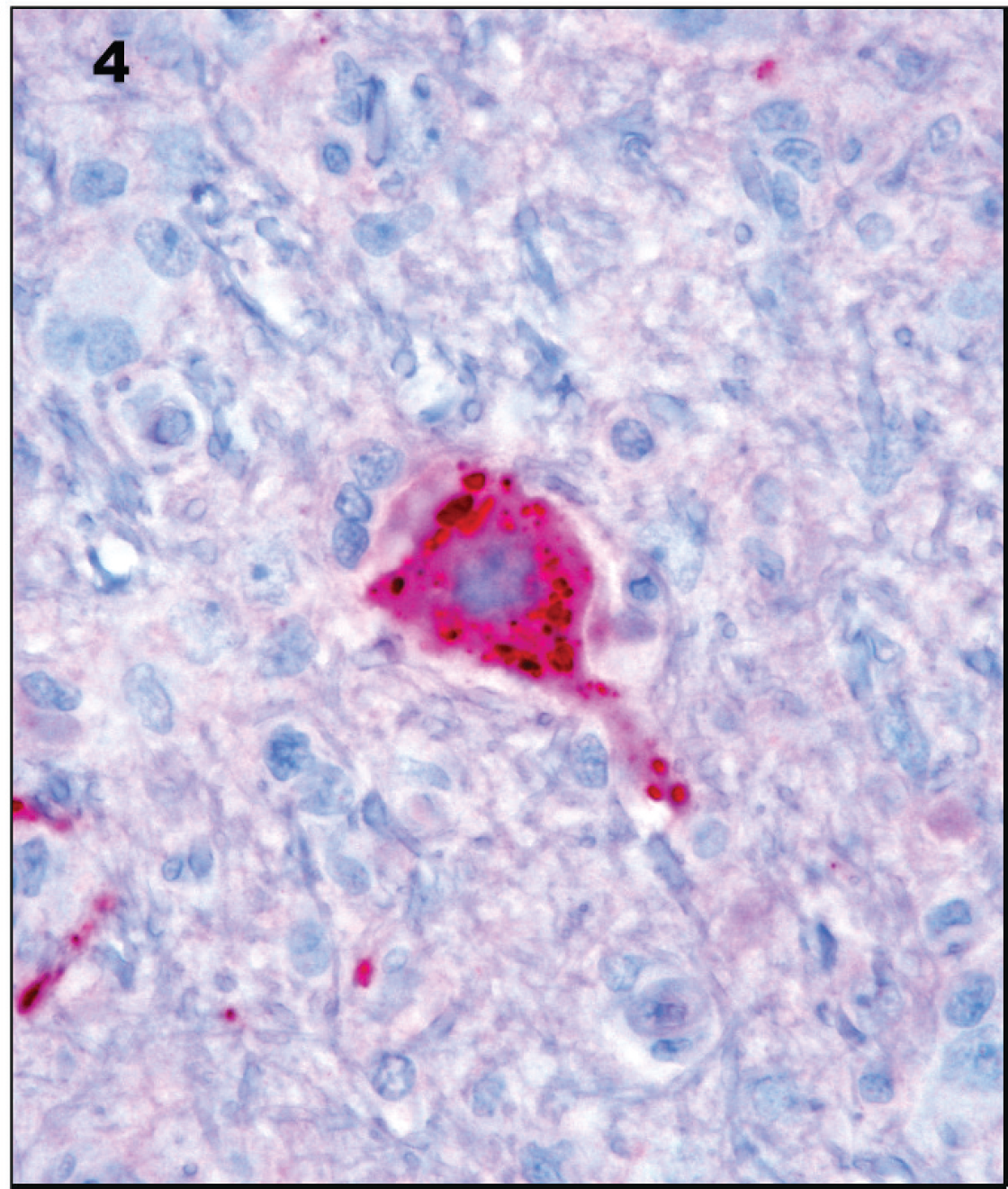
Immunohistochemistry for Canine distemper virus discloses intense intraneuronal staining. 60× magnification.
Primary differentials for the neurologic signs, MRI, and CSF results included necrotizing meningoencephalitis (NME), granulomatous meningoencephalitis (GME), and infectious meningoencephalitis (protozoal, fungal, algal, rickettsial, bacterial, or viral). Serum antibody reactivity was not found for Ehrlichia canis (immunoglobulin G [IgG] indirect immunofluorescent antibody test [IFAT]), Borrelia burgdorferi (IgG enzyme-linked immunosorbent assay [ELISA]), Rickettsia rickettsii (IgG IFAT), Bartonella vinsonii (IgG IFAT), Babesia canis (IgG IFAT), Bartonella henselae (IgG IFAT), Toxoplasma gondii (IgG and IgM ELISA), or Neospora caninum (IgG IFAT). Agar gel immunodiffusion fungal panels also yielded negative results for serum antibody to Cryptococcus, Blastomyces, Coccidioides, and Histoplasma spp. Specific immunologic testing for viruses was not performed. The presumptive diagnosis was autoimmune CNS inflammation (e.g., NME or GME), although infectious meningoencephalitis could not be ruled out entirely.
The day after neurodiagnostic testing, the dog deteriorated and was euthanized. At necropsy, gross lesions were confined to the brain. A 2.0 cm × 0.5 cm area of hemorrhage was located along the ventral aspect of the left medulla oblongata. On sectioning of the fixed brain, a 6.0 mm × 5.0 mm area of discoloration was found along the medial aspect of the left thalamus, corresponding to the thalamic lesion identified on MRI (Fig. 1). Histologically, the thalamic lesion was characterized by severe necrosis and inflammation, with edema and hemorrhage (Fig. 2). The inflammatory infiltrate was composed of lymphocytes, plasma cells, macrophages, and fewer neutrophils, which formed thick perivascular cuffs and extended into the surrounding edematous and necrotic neuropil (Fig. 3). Numerous gitter cell macrophages were present in this area. Swollen axons were common, and an increased number of microglia and astrocytes were present. Similar inflammatory foci with a milder degree of necrosis were present in a multifocal to coalescing pattern throughout the mesencephalon, ventral metencephalon, myelencephalon, and cerebrum. Locally extensive areas of the cerebral cortex had severe neuronal necrosis with vacuolation and prominent capillaries with hypertrophied endothelial cells. Microorganisms or viral inclusions were not observed. The lesions were thought to resemble NME, although the degree of thalamic and brain stem involvement was considered unusual for the disease.
The brain from this French Bulldog was included in a large, blinded pan-viral PCR study of canine meningoencephalitis of unknown etiology (MUE). RNA was extracted from 16 brains (12 NME and 4 GME) stored at −80°C, by using a commercial kit b per the manufacturer's protocol. Samples were tested by the panel of panparamyxovirus RT-PCR assays using a commercial 1-step RT-PCR kit. c 22 The RT-PCR mixtures contained 50 pmol each of forward and reverse primers (pan-PAR-F1/PAR-R [round 1], pan-PAR-F2/PAR-R [round 2]; pan-RES-MOR-HEN-F1/RES-MOR-HEN-R [round 1], pan-RES-MOR-HEN-F2/RES-MOR-HEN-R [round 2]; and pan-AVU-RUB-F1/AVU-RUB-R [round 1], AVU-RUB-F2/AVU-RUB-R [round 2]), 22 1 × buffer with a final concentration of 2.0 mM MgSO4, 20 units of RNase inhibitor, a 5-μl aliquot of RNA extracts, and 1 unit of an RT-Taq mix. c Water was added to achieve a final volume of 50 μl. The RT-PCR reaction mixture was incubated at 60°C for 1 min for denaturing, 44–50°C for 30 min (for RT), 94°C for 2 min (for hot start), then 40 cycles at 94°C for 15 sec, 48–50°C for 30 sec, 72°C for 30 sec, and a final extension at 72°C for 7 min. The final PCR products were visualized by ultraviolet light after electrophoresis on a 2% agarose gel containing 0.5 μg/ml ethidium bromide in 0.5× Tris-borate buffer (pH 8.0).
The pan-paramyxovirus RT-PCR tests on the French Bulldog brain generated intense staining amplicons of the expected size (563 bp) for both Paramyxovirinae subfamily RT-PCRs and pan-Morbillivirus-Respirovirus-Henipavirus genera-subgroup RT-PCRs compared with their positive controls (Fig. 5); water was extracted in parallel to clinical samples and non-template, water controls generated no amplicons. The positive controls for Paramyxovirinae sub-family RT-PCRs and pan-Morbillivirus-Respirovirus-Henipavirus genera-subgroup RT-PCRs are the synthesized RNA fragments that have engineered multiple mutations from the L polymerase gene of Human parainfluenza virus 2 and CDV, respectively. A total of 8 of 15 additional NME and GME brains generated faint to mildly intense staining of PCR amplicons of the expected size for pan-Paramyxovirinae PCR only (Fig. 5). All positive PCR products were gel purified, d and both strands of the amplicons were sequenced with a ready reaction cycle sequencing kit e using the corresponding PCR primers. Sequencing of the amplicons from the French Bulldog brain generated sequences with 98.7% homology to the CDV L polymerase gene fragment. Sequencing of the 8 fainter PCR amplicons (from cases of canine NME and GME) generated random canine sequences, suggesting mispriming of the pan-Paramyxovirinae primers to canine DNA.
Subsequent to reevaluation of the French Bulldog brain (which failed to disclose viral inclusion bodies), IHC was performed for CDV. Briefly, formalin-fixed tissues were trimmed, embedded in paraffin, sectioned at 5 μm, and processed for IHC using a previously described protocol. 20 An alkaline phosphatase red detection system, f as well as bulk buffers specifically designed for use on the BenchMark Automated Staining System, f were used for immunolabeling and visualization. Slides were baked in a drying oven at 60°C for 30 min. The slides were then barcode labeled and placed in the BenchMark for deparaffinization and heat-induced epitope retrieval. Antigen retrieval was performed using retrieval solution CC1 standard. g A mouse monoclonal anti-CDV antibody g at a concentration of 1:500 for 32 min was used as primary antibody. The slides were counterstained, f and after bluing for 2 min each, the slides were dehydrated, cleared, and mounted. Formalin-fixed, paraffin-embedded sections of lung and bladder from a dog that had tested positive previously for CDV infection by RT-PCR were used as positive controls. For negative controls, the primary antibody was replaced with homologous nonimmune sera. Positive IHC staining was present in neurons in the brain of the French Bulldog, confirming the pan-paramyxovirus RT-PCR results (Fig. 4).
The IHC confirmation of active viral replication raises questions about the CDV pathogenesis in the present case. The dog received a complete series of (nonexpired) modified live (Rockborn strain) CDV vaccinations. One possibility is the development of postvaccinal CDV caused by an incompletely attenuated viral strain. This pathogenesis seems unlikely because of the several-month interval between the dog's final vaccination and the development of meningoencephalitis; clinical signs with postvaccinal CDV typically are seen within 2 weeks of vaccination. 4 Moreover, the severe necrotizing inflammation and lesion distribution are not consistent with previous descriptions of postvaccinal CDV infections. 3,9 The latter also has a predilection for ventral pons, and intranuclear and intracytoplasmic inclusion bodies typically are present in many neuronal cells, whereas inclusions were not present in the current case. 9 The more likely explanation is that the dog had an incomplete immune response upon vaccination and developed a natural CNS infection upon exposure to a wild strain of CDV.
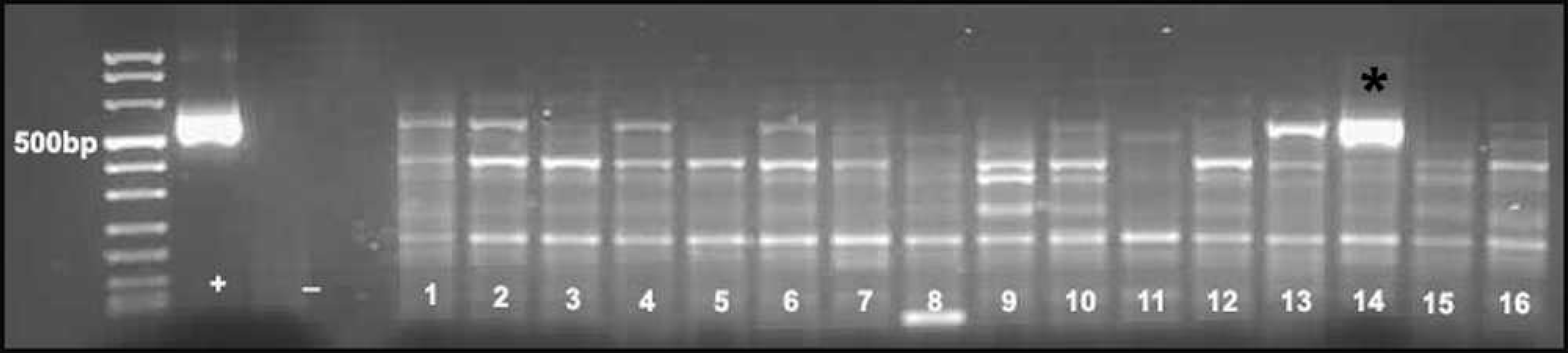
Pan-paramyxovirus polymerase chain reaction on total RNA extracted from 16 freshly frozen canine brains with necrotizing meningoencephalitis (NME) and granulomatous meningoencephalitis. Sample 14 is from the French Bulldog with NME and has a strong amplicon (*) of the expected size (563 bp). Fainter amplicons also are present for samples 1, 2, 4, 5, 7, 10, 13, and 16. Sequencing of all bands, except sample 14, generated nonspecific canine sequences. Sequencing of sample 14 disclosed Canine distemper virus. + = positive control; - = negative control.
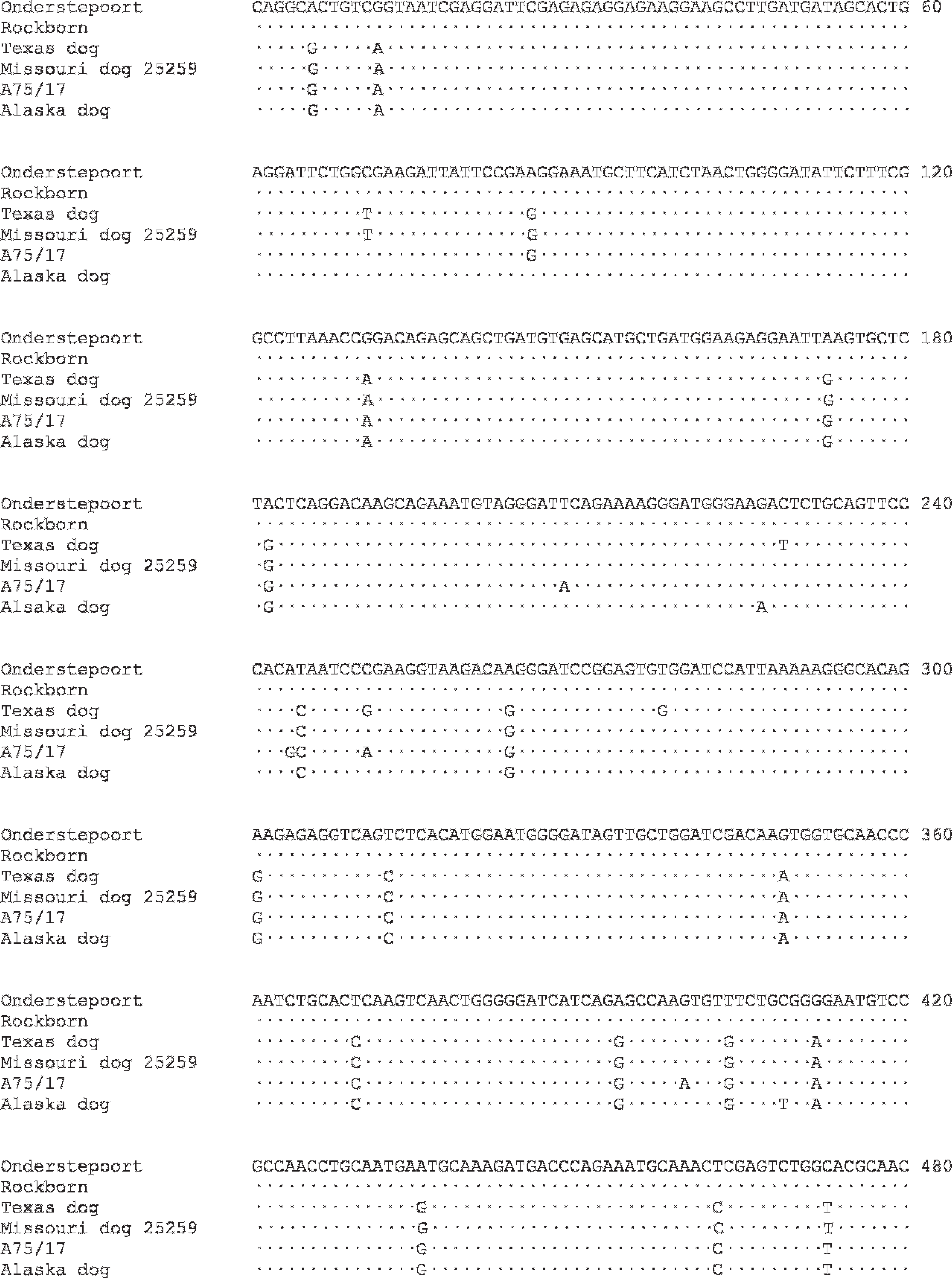

Nucleotide sequence alignment of 480-bp segment of P gene vaccine and wild Canine distemper virus strains. Differences in nucleotide sequence from the Onderstepoort vaccine strain are shown; dots represent sequences identical to Onderstepoort. GenBank accession numbers are in parentheses: Onderstepoort (AF014953), Rockborn (AF181446), Missouri dog 25259 (AY964113), A75/17 (AF164967), Alaska dog. 14 The Texas dog is the case of meningoencephalitis of unknown etiology reported here.
To more definitively characterize the infection as secondary to a vaccinal or wild strain of CDV, specific RT-PCR and sequence analyses of the CDV phosphoprotein (P) gene were performed. 13 Subsequent to RT-PCR, a 585-bp amplicon was gel purified d and sequenced e bidirectionally using the CDV-specific PCR primers. Sequence data for a 480-bp segment of the P gene were aligned with previously published sequences from vaccine and wild strains of CDV using ClustalW2 (Fig. 6). 12,13,16 Sequence alignment revealed significant disparity in the CDV strain detected from the current case when compared with the Onderstepoort and Rockborn vaccine strains, with a total of 22 nucleotide differences. Seventeen of these nucleotide differences were mutual among all of the aligned wild CDV strains. There was 99.4% homology to a wild strain isolated from a Missouri dog in 2004, as well as a 98.5% homology to wild strain A75/17 and an Alaskan dog isolate. 13,16 Although mutation of the vaccinal strain cannot be excluded, sequence analysis of the P gene strongly suggests that the dog was infected with a wild strain of CDV.
In the current investigation, the pan-paramyxovirus approach included Paramyxovirinae subfamily PCR, pan-Morbillivirus-Respirovirus-Henipavirus genera-subgroup PCR, pan-Rubulavirus-Avulavirus genera-subgroup PCR, and pan-Pneumovirinae subfamily PCR. 22 This group of paramyxovirus PCR primers collectively target highly conserved amino acid sequences within each of 2 subfamilies and each of 2 genera subgroups, based on the multiple alignments of 33 deduced, nonredundant paramyxovirus L polymerase genes. This broadly reactive RT-PCR, coupled with sequence analysis and IHC, identified CDV in this canine case of MUE. Specific RT-PCR and IHC testing for CDV was not pursued initially because CNS CDV infection was not a primary antemortem or histopathologic differential.
Although CDV is a known cause of canine encephalitis, the present case demonstrates the utility of pan-viral PCR in situations where specific microbial etiologies are considered unlikely. Similar strategies have been used to identify and characterize a number of novel human viruses, including Severe acute respiratory syndrome coronavirus, 11 Hepatitis G virus, 19 Sin Nombre virus, 14 Human retrovirus-5, 7 and novel animal viruses, such as the Macaque gamma herpesvirus and Pig endogenous retrovirus. 17,18
In conclusion, the current case report illustrates the utility of pan-paramyxovirus RT-PCR and sequence analysis in a case of canine MUE. In light of the CDV sequence that was identified, IHC investigations for CDV were pursued and confirmed active viral replication. Future studies will use pan-viral PCR approaches to evaluate various canine MUEs.
Footnotes
a.
Siemens Magnetom Impact Expert, Siemens Medical Solutions USA Inc., Malvern, PA.
b.
RNeasy® Mini Kit, Qiagen Inc., Valencia, CA.
c.
SuperScript™ III Platinum® One-Step RT-PCR System, SuperScript™ III RT/Platinum® Taq Mix, Invitrogen Corp., Carlsbad, CA.
d.
MinElute® Gel Extraction Kit, Qiagen Inc., Valencia, CA.
e.
Applied Biosystems, Foster City, CA.
f.
Enhanced V Red (Alkaline Phosphatase Red) Detection System, Ventana hematoxylin; Ventana Medical Systems Inc., Tucson, AZ.
g.
DV2-12, Custom Monoclonals International, West Sacramento, CA.
