Abstract
To compare the real-time reverse transcription quantitative polymerase chain reaction (RT-qPCR) assays used for the diagnosis of Porcine reproductive and respiratory syndrome virus (PRRSV), a Europe-wide interlaboratory ring trial was conducted. A variety of PRRSV strains including North American (NA) and European (EU) genotype isolates were analyzed by the participants. Great differences regarding qualitative diagnostics as well as analytical sensitivity were observed between the individual RT-qPCR systems, especially when investigating strains from the EU genotype. None of the assays or commercial kits used in the ring trial could identify all different PRRSV strains with an optimal analytical and diagnostic sensitivity. The genetic variability of the PRRSV strains, which is supposed to hinder the diagnostic of the RT-PCR because of mutations at the primer binding sites, was also confirmed by sequencing and subsequent phylogenetic analysis. In summary, a major problem in PRRSV diagnostics by RT-qPCR is false-negative results. To achieve maximum safety in the molecular diagnosis of PRRSV, the combined usage of different assays or kits is highly recommended.
Keywords
Introduction
Porcine reproductive and respiratory syndrome (PRRS), characterized by reproductive failure in pregnant sows and respiratory disease in piglets, 29 has a major economic impact on the pig industry worldwide. The causative agent is an enveloped positive-strand RNA virus (order Nidovirales, family Arteriviridae, genus Arterivirus 4 ) that was first isolated in the Netherlands. 38 The genome of the PRRS virus (PRRSV) is organized in 9 open reading frames (ORFs) coding for the viral replicase (ORF1a and 1b), 4 membrane-associated glycoproteins (ORFs 2a–5), 2 unglycosylated membrane proteins (ORF2b and ORF6), a polypeptide termed ORF5a protein, and the nucleocapsid protein (ORF7).6,14,19,20,40 PRRSV isolates are classified into 2 genotypes, the European (EU, genotype 1 [EU]) and the North American (NA, genotype 2 [NA]). 22 Although inducing the same disease symptoms, both genotypes are antigenically and genetically very different.16,28 Even within the EU genotype, originally thought to form a more homogeneous group, 38 a high genetic diversity was demonstrated.7,9,25,32,35 The classification of EU genotype PRRSV into 4 subtypes has been suggested recently.30,31
For the detection of the highly variable PRRSV genome, different commercial kits as well as a variety of published reverse transcription polymerase chain reactions (RT-PCR) and real-time RT quantitative PCR (RT-qPCR) systems are available. Previously described assays are either suitable for pan-PRRSV detection 2 or they are designed to differentiate between the EU and the NA genotype. This differentiation is possible in a single multiplex amplification8,17,26 or in 2 separate reactions. 27 Furthermore, there are RT-PCR assays for the specific genotype detection available.1,31 Beside these published RT-PCR assays, several commercial kits for the genome detection and differentiation of PRRSV are available.
The aim of the current study was a comparative assessment of the wide range of assays (in-house and commercial test systems) used in different European laboratories. Using a variety of PRRSV EU and NA strains, the suitability of these assays for a reliable PRRSV genome detection was investigated.
Materials and methods
PRRSV isolates and diagnostic samples
The PRRSV strains used in the current study were obtained from the virus collection of the Friedrich-Loeffler-Institut, Isle of Riems, Germany or were kindly provided by T. Stadejek, National Veterinary Research Institute, Poland. Designation, collection date, and country of the strains are summarized in Table 1. RNA was manually extracted a according to the manufacturer’s recommendations and diluted in RNA extracted from blood of PRRSV-negative swine. Two log10 dilution series of EU strains and 1 NA-HP (highly pathogenic) strain were prepared; 7 additional EU strains, at least 1 of each subtype, and 3 NA strains were also integrated in the panel. Finally, 3 negative samples were added. The 28 samples were randomly coded and sent to each of the 7 participating laboratories on dry ice. In addition to these ring trial samples, diagnostic lung specimens, kindly provided by German veterinary laboratories, were further studied using the commercial RT-qPCR kits.
Identity of the 28 Porcine reproductive and respiratory syndrome virus RNA samples used in the ring trial.*
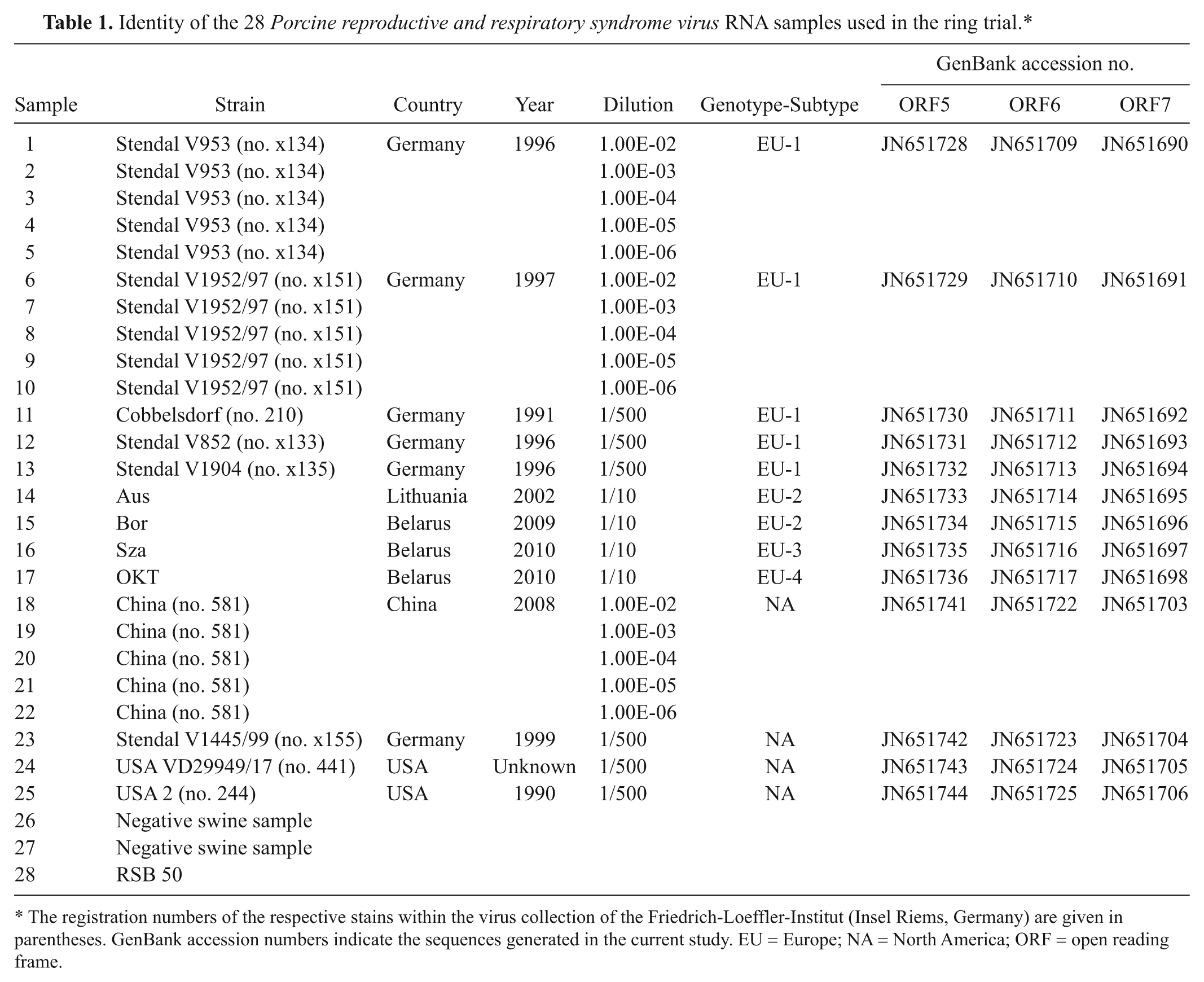
The registration numbers of the respective stains within the virus collection of the Friedrich-Loeffler-Institut (Insel Riems, Germany) are given in parentheses. GenBank accession numbers indicate the sequences generated in the current study. EU = Europe; NA = North America; ORF = open reading frame.
Real-time RT-PCR
In the 7 participating laboratories, a total of 4 different commercially available RT-qPCR kits (I–IV) and 8 in-house assays (Table 2) were used. The applied commercial kits varied regarding total reaction volume (kit I: 20 µl, kits II–IV: 25 µl) and template volume (IV: 4-µl kit, I+II: 5 µl, III: 8 µl). Kits I and II have been used by 1 of the participating laboratories and kits III and IV by 3 laboratories.
Sequences of primers and probes of the in-house assays used for Porcine reproductive and respiratory syndrome virus genome detection in the ring trial.*
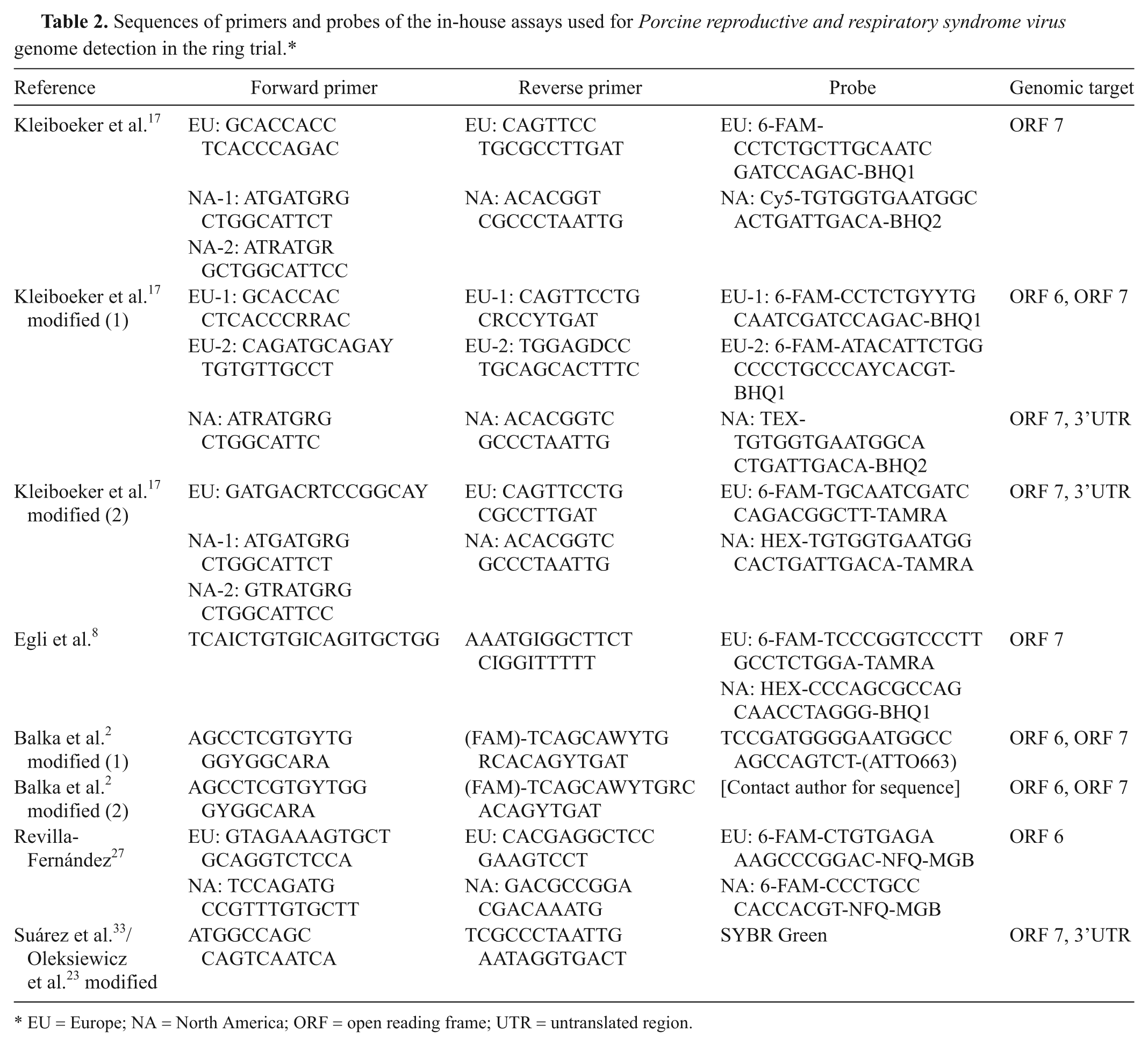
EU = Europe; NA = North America; ORF = open reading frame; UTR = untranslated region.
A part of the in-house RT-qPCR systems were used as described previously,8,17,27 while individual modifications were integrated in other assays.2,17,23,33 In 5 out of 7 laboratories, the samples were tested by more than 1 assay. Previously described systems8,17 as well as 3 of the commercial kits enable genotype differentiation in a single reaction; a previously described assay 27 and 1 of the commercial kits required for testing feature 2 separate reactions. One RT-qPCR system(23,33 modified; Table 2) allows genotyping only after gel electrophoresis.
RNA extracted from German field lung samples was tested by the 4 commercial kits according to the manufacturer’s recommendations within a single laboratory. The presence of PRRSV genome was confirmed by sequencing.
Sequencing and phylogenetic analysis
ORFs 5, 6, and 7 of the PRRSV strains were sequenced using published primers,10,23,28 based on data obtained hereby alternative primers were designed (Table 3). The RT-PCR was carried out using a commercial PCR kit, b and the amplified products were separated on 2.0% agarose gels, c visualized by staining with ethidium bromide, and subsequently excised and purified d according to the manufacturer’s recommendations. Sequencing reactions of both strands with the primers used for amplification were carried out. e Subsequent sequence determination was performed, f and the obtained sequence data was assembled. g Sequence alignments, identification of ORFs, and calculation of similarity of the used strains to their respective prototype strains were supported by BioEdit software version 7.0.9.0. 12 Sequences generated in the current study were submitted to GenBank (accession numbers are given in Tables 1 and 6, respectively). Neighbor-joining trees were generated using the Kimura 2-parameter model with 1,000 bootstrap replicates as implemented in Mega 5. 34 In addition to the PRRSV strains used in the ring trial, a selection of reference sequences representing the global diversity of PRRSV was used for phylogenetic analysis (Table S1 in the supplementary materials). These reference sequences were selected from GenBank and comprise strains from Europe, Asia, and North America.
Primers used in the current study for sequencing.*
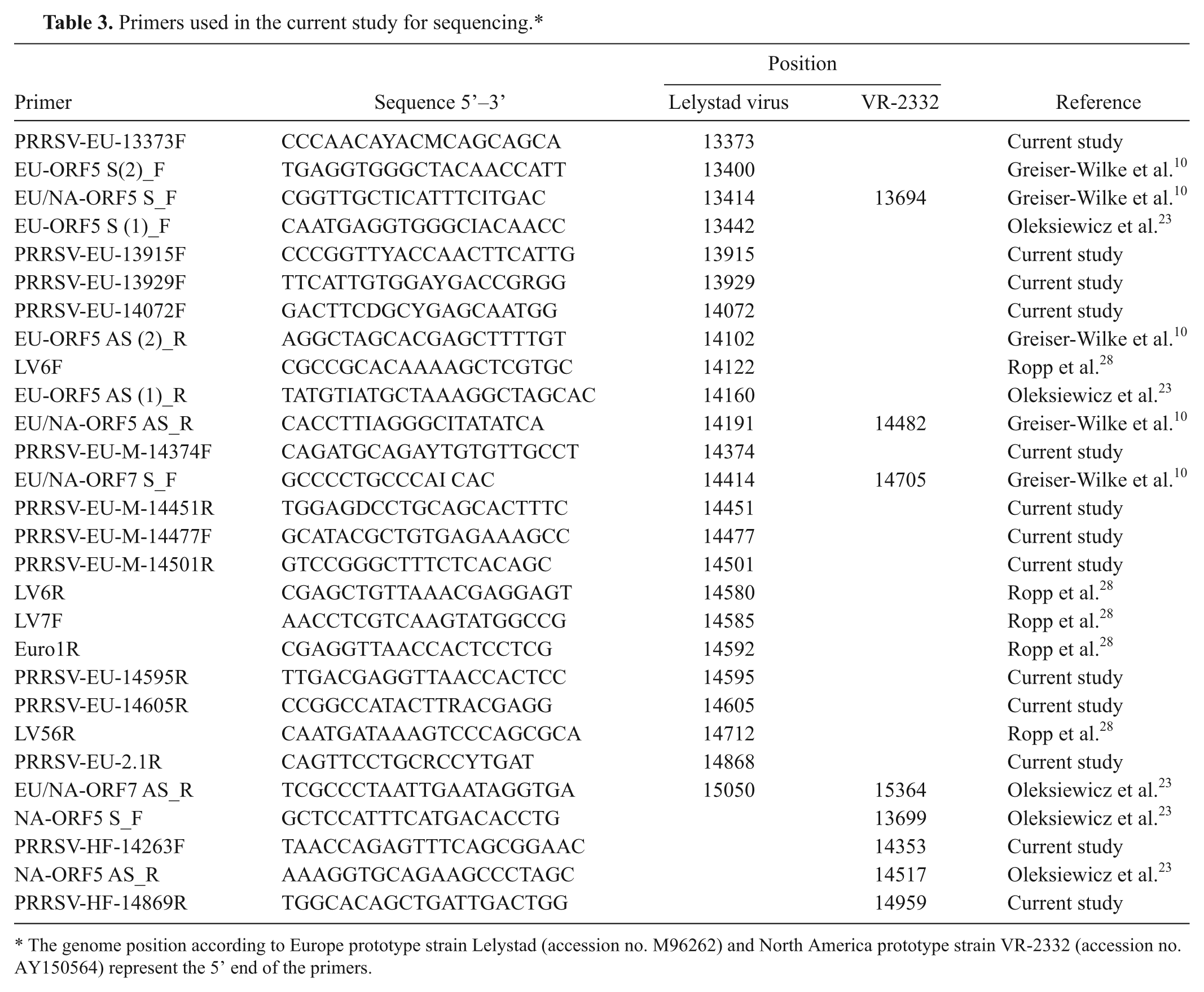
The genome position according to Europe prototype strain Lelystad (accession no. M96262) and North America prototype strain VR-2332 (accession no. AY150564) represent the 5’ end of the primers.
Results
Real-time RT-PCR
The results of the pan-PRRSV genome detection are outlined in Table 4 and of the genotype-specific assays in Table 5. Samples were defined as false negative if at least 50% of the assays produced a positive result, which means, for the pan-PRRSV genome detection, that false-negative results were postulated if 8 or more assays gave a positive result. For PRRSV genotype-specific RT-qPCR detection, 7 positive results were necessary to define negative results of this RNA dilution as false negative. The NA-HP strain used as dilution series was detected correctly by all assays; however, the analytical sensitivity ranged from dilution step 1.00E-03 to 1.00E-06. All of the other NA strains, with exception of “USA 2” in 1 assay, were also detected correctly. Every NA strain was classified into the NA genotype by the PRRSV-NA–specific PCR method, and only 1 assay produced an EU genotype-positive result incorrectly. When investigating the EU-PRRSV samples, the assay published previously 8 failed to detect 2 of the 5 EU-1 strains integrated in the ring trial panel. The other assays scored correctly positive, even though differences regarding the analytical sensitivity were observed. The 5 log 10 dilution steps of strain “Stendal V953” gave a positive result in 3 assays, 4 assays detected 4 dilution steps correctly, 7 methods gave a positive result for 3 dilution steps, and 1 assay detected PRRSV RNA in the 2 samples with the highest genome amount. The EU-2 strains integrated in the ring trial were tested positive by 81.25% and 68.75% of the used assays, the EU-3 strain by 93.75%, and the EU-4 strain OKT was detected correctly positive by 75.0% of the assays. When testing the EU-2 strains, kit I gave a negative result in the pan-assay, whereas the genotype-specific assay scored positive. Every PRRSV-EU strain was classified into the correct genotype by the assays used in the ring trial; the solitary exception was sample 9 investigated with 1 of the commercial kits.
Mean quantification cycle (Cq) values of pan–Porcine reproductive and respiratory syndrome virus (PRRSV) genome detection submitted by the 7 participating laboratories using commercial reverse transcription quantitative polymerase chain reaction kits and in-house assays.*

EU = Europe; NA = North America. False-negative results are depicted in bold; false-positive results are depicted in italic.
Only 1 of the duplicates scored positive.
Very flat curve.
Mean quantification cycle (Cq) values of genotype-specific genome detection using commercial reverse transcription quantitative polymerase chain reaction kits and in-house assays.*
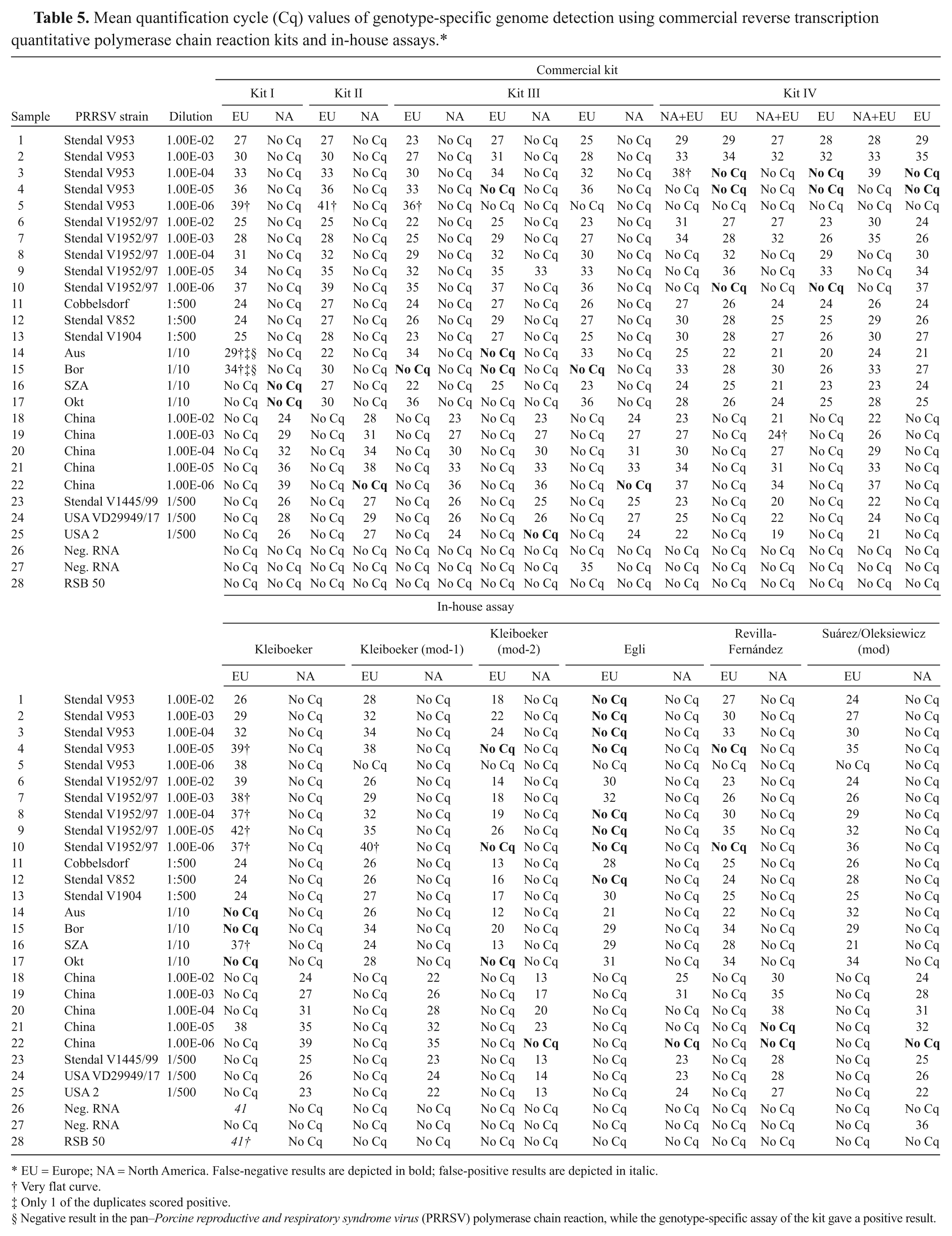
EU = Europe; NA = North America. False-negative results are depicted in bold; false-positive results are depicted in italic.
Very flat curve.
Only 1 of the duplicates scored positive.
Negative result in the pan–Porcine reproductive and respiratory syndrome virus (PRRSV) polymerase chain reaction, while the genotype-specific assay of the kit gave a positive result.
For further validation of the assays, lung samples were tested by the different commercial kits. The results obtained with those samples using the pan-PRRSV as well as the genotype-specific commercial kits are shown in Table 6. In 1 sample, the EU genotype was detected consistently, while in the 2 other samples containing only EU-PRRSV, 1 kit failed to detect the genome. All 4 kits gave a correct positive result when investigating the sample containing NA-PRRSV, whereas in 1 case (kit IV), an incorrect EU-PRRSV–positive result was obtained. Double infection with both genotypes in sample “BH 95/10-8” was detected by all kits used in the current study; however, kit IV did not enable a robust differentiation between infection with both genotypes and NA-PRRSV.
Identity of lung samples and mean quantification cycle (Cq) values of Porcine reproductive and respiratory syndrome virus (PRRSV) genome detection using the commercial reverse transcription quantitative polymerase chain reaction kits.*
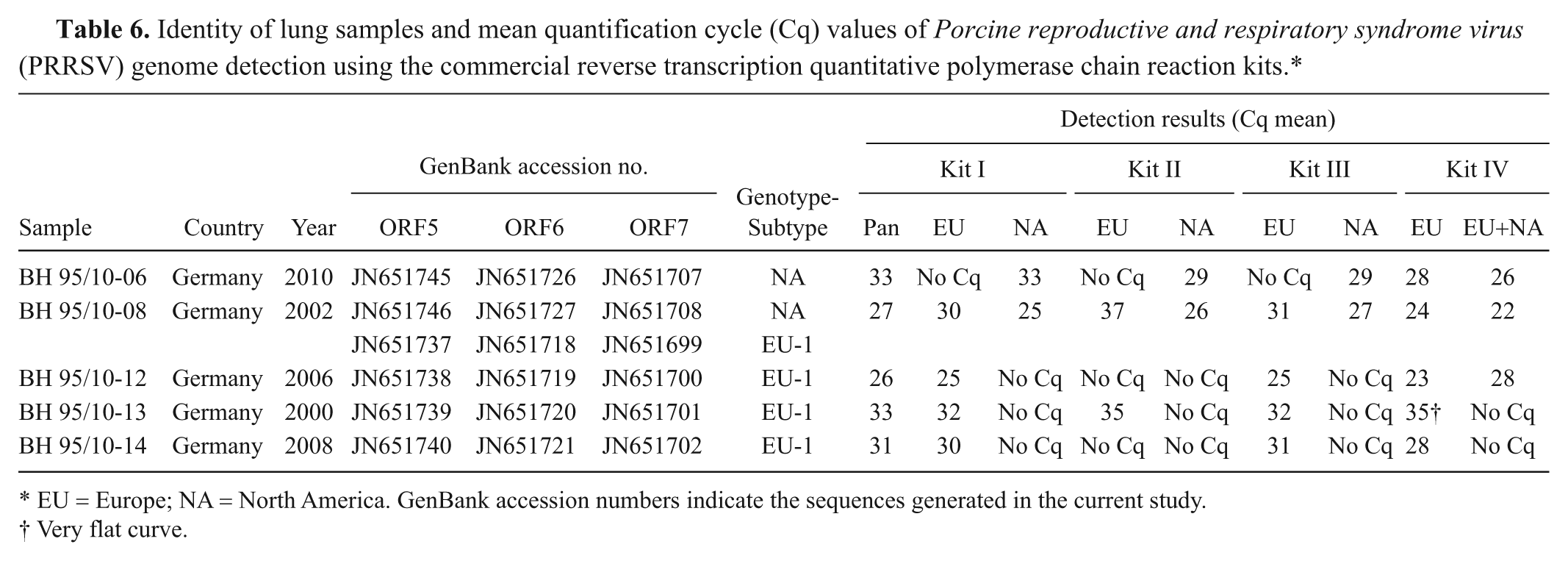
EU = Europe; NA = North America. GenBank accession numbers indicate the sequences generated in the current study.
Very flat curve.
Sequencing and phylogenetic analysis
The phylograms (Fig. 1a–c) of the sequences generated in the current study compared to the reference strains selected from GenBank illustrate the genetic diversity of the PRRSV strains used in this ring trial. The European strains can be separated into 4 genetic subtypes. When analyzing ORF5 sequences of the ring trial samples, 5 of the EU genotype strains are classified into subtype 1, 2 strains into subtype 2, 1 strain into subtype 3, and strain “Okt” into subtype 4. The isolates “China,” “Stendal V1445/99,” “USA VD29949/17,” and “USA 2” were clearly assigned to the NA genotype. The classification observed in ORF5 correlates with the phylogenetic grouping of ORF6 as well as ORF7 nucleotide sequences.
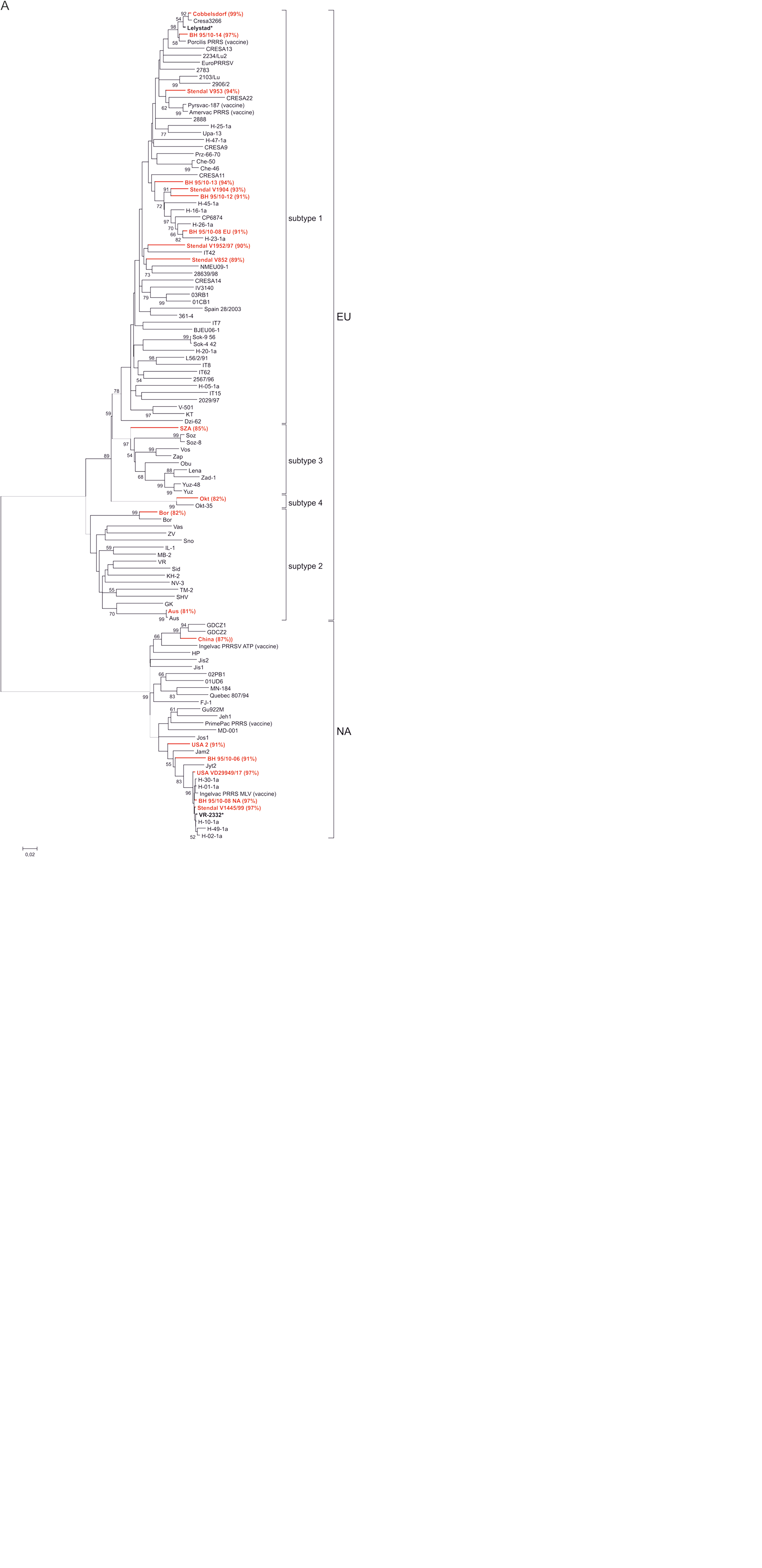
Phylogenetic analysis of ring trial samples and reference strains based on nucleotide sequences of open reading frame (ORF)5 (A), ORF6 (B), and ORF7 (C). Each isolate is indicated by virus name; ring trial samples are depicted in red. Statistical support for nodes was obtained by bootstrapping (1,000 replicates); only values ≥50% are shown. Scale bar indicates the estimated number of nucleotide substitutions per site. Sequence identity of the ring trial samples to their respective prototype strain is depicted in parentheses. The prototype strains are Lelystad for Europe (EU) and VR-2332 for North America (NA) genotype (both are marked by asterisk).
In 1 of the lung samples collected in Germany, the NA genotype was detected, 1 further sample contained both genotypes (mixed infection), clearly separated by sequencing, and in the 3 additional samples, EU-PRRSV was found. None of them belonged to the Eastern European subtype.
The similarity of the NA strains to their prototype strain “VR2332” ranged between 87% and 97% for complete ORF5 sequences, 95% and 99% for ORF6, and 94% and 100% for ORF7 sequences. The EU strains used are 81–99% (ORF5), 86–99% (ORF6), and 83–100% (ORF7) identical to their prototype strain “Lelystad.” Major length differences were found between the ORF7 of EU-1 isolates, which were 387 nucleotides (nt), with exception of BH 95/10-12, which had a length of 381 nt.
Discussion
To compare the wide range of RT-qPCR systems for PRRSV genome detection, a variety of EU and NA strains were sent to 7 European laboratories. The presence of PRRSV genome was confirmed by sequencing and phylogenetic analysis. A total of 4 different commercially available RT-qPCR kits and 8 in-house assays were used to analyze the samples. Altogether, huge differences were observed between the individual RT-qPCR systems. Based on the selected PRRSV strains used in the ring trial, no assay or commercial kit could be identified with optimal analytical and diagnostic sensitivity. In general, inconsistencies were mainly observed in the detection of the EU genotypes compared to the PRRSV-NA strain–specific sequences. This observation could be explained by the large genetic diversity of the viruses within the EU genotype.7,25,32 Atypically diverse EU genotype PRRSV strains have been reported in several European countries, 18 for instance in Denmark, 24 Spain, 7 Germany, and the Netherlands, 25 as well as in Italy. 9 The global dissemination of pan-European subtype 1 is assumed, as strains classified into this subtype have been recently reported in Asian countries such as China, 5 Korea, 21 and Thailand. 37 Subtypes 2, 3, and 4, whose existence was solely reported from Eastern Europe so far, exceed the diversity of subtype 1.15,30,31 Four of the Eastern European, non–EU-1 strains were integrated in the ring trial, and these strains caused major difficulties concerning molecular detection. The strain “Bor,” for example, was only detected by 68.75% of the RT-qPCR assays.
Contrary to the EU strains, the NA strains used in the current study were rather similar. As a result, the actual performance capability of the qPCR assays regarding NA strains is not reflected in its entirety.
Analytical sensitivity of the applied assays varied in a range of up to 3 log 10 dilution steps for the NA as well as for the EU strains. However, an insufficient diagnostic sensitivity has to be assessed more critically than a reduced analytical sensitivity. In summary, commercial kit II and the in-house assay using a combination of a modified published assay 17 and an additional PRRSV-EU assay based on ORF6 (modification 1) revealed the best results when investigating the virus strains sent to the participating laboratories. However, commercial kit II failed to correctly detect 2 of the strains isolated from German lung samples. In contrast, the Kleiboeker-modification 1 assay had no problems detecting these strains (data not shown).
Similar to the current study, in a ring trial conducted in Germany, 36 false-negative results were found to be the most common pitfall of the RT-PCR assays used. The presence of inhibitors within the reaction, not fully optimized PCR systems, unsuited primer sequences, and not standardized RNA preparation methods were suspected to be responsible for this observation. The latter can be excluded in the current Europe-wide ring trial because identical RNA samples were sent to the participating laboratories. To ensure adequate extraction of nucleic acids, especially when using potentially difficult biological matrices, as well as to confirm the absence of PCR inhibiting factors, the coamplification of an internal control is a valuable tool.3,13
To avoid mismatches in the primer and probe binding sites, the sequences should be controlled continuously by sequence analysis of emerging strains. In addition to ORF5, whose sequences were commonly used for phylogenetic analysis, sequencing of ORF6 and 7 was conducted, because most of the RT-qPCR assays used in the ring trial target this genomic region (see Table 2). Phylogenetic grouping of ORF5, 6, and 7 showed a good correlation for the selected strains. In agreement with previous studies,30,31 the division of EU isolates into 4 subtypes is observed in ORF5 and 7 phylograms. In the ORF6 phylogenetic tree, subtype 1 isolates can be separated from other subtypes, and further division is not possible due to the lack of ORF6 sequence information of additional Eastern European strains.
For EU isolates, concordance between the size of ORF7 and the phylogenetic grouping has been reported 31 ; subtype 1 contained viruses with an ORF7 size of 387 nt. However, 1 of the lung samples collected in Germany, which clearly could be classified as subtype 1, exhibited an ORF7 containing only 381 nt, thus size polymorphism is observed within subtype 1. Consequently, for subtyping of EU PRRSV isolates, the sequence analysis is most suitable. Furthermore, sequencing is essential to notice mismatches in the primer and probe binding sites. However, identity of the complete ORF sequences of the used strains to their respective prototype strain did not correlate in full with the real-time PCR detection results. As an example, lung sample “BH 95/10-14” with an identity of 99% (ORF6, 7) and 97% (ORF5), respectively, to “Lelystad” was not detected by 1 of the commercial kits. The same result was obtained when investigating subtype 3 strain “SZA,” whose identity to the EU prototype strain ranged between 83% and 86% for the sequenced ORFs 5, 6, and 7. This observation emphasizes the importance of the combined application of several, independent real-time detection systems, which is possible either by usage of different assays or kits or by combination of 2 or more qPCR systems within a single multiplex reaction. The latter is realized in the Kleiboeker-modification 1 assay. For the detection of EU genotype strains, 2 systems are used that target ORFs 6 and 7 sequences, respectively.
In 1 of the samples isolated from lung material, EU and NA genotype PRRSV was independently detected by 3 commercial kits and confirmed by sequencing. In Europe, coexistence of both genotypes within the same swine herd has been documented.10,23 The EU genotype PRRSV isolates have been introduced into North American swine herds 28 as well. To investigate the spread of both genotypes, the classification of new isolates is highly recommended.
Genotyping was possible by most of the RT-qPCR assays used in the ring trial, either in 2 separated or in a single multiplex reaction. Multiplex approaches, where 2 or more targets are amplified in the same tube and detected in parallel, 39 enable cost-effective and fast answers to several diagnostic questions 11 (e.g., in the case of PRRSV, the detection and genotyping of a sample in a single reaction). The selected PRRSV strains used in the ring trial were correctly assigned to the appropriate genotype by almost all assays. However, kit IV is, in contrast to manufacturer’s information, not useful for genotyping. Based on the fact that one probe is specific for EU and another for EU and NA, a differentiation between a double infection with both genotypes and NA-PRRSV is not possible. As an example, the lung sample “BH 95/10-08” would not have been found as being double infected, as the results of the other kits and the sequencing of the sample had suggested. But on the other hand, PRRSV genome was detected in every lung sample by this kit, even though genotyping is not possible. In contrast, 2 out of 5 of these lung samples were tested negative by kit II. This observation emphasizes that the greatest problem in PRRSV genome detection is the false-negative results. Because of the great genetic diversity of PRRSV strains, even affecting binding sites for diagnostic primers and probes, the combined usage of different assays or kits is highly recommended to achieve a maximum of diagnostic safety.
Footnotes
Acknowledgements
The authors thank Christian Korthase and Patrick Zitzow for excellent technical assistance, and Sabine Bilk (Landeslabor Berlin Brandenburg) and Tilman Kühn (synlab, Leipzig) for providing diagnostic lung samples.
a.
QIAamp® Viral RNA Mini Kit, Qiagen GmbH, Hilden, Germany.
b.
Super Script® III One-Step RT-PCR with Platinum® Taq, Invitrogen, Carlsbad, CA.
c.
Roche Diagnostics Deutschland GmbH, Mannheim, Germany.
d.
QIAquick® Gel Extraction Kit, Qiagen GmbH, Hilden, Germany.
e.
BigDye® Terminator Cycle Sequencing Kit Version 1.1, Applied Biosystems, Germany.
f.
ABI 3130 Genetic Analyzer, Life Technologies GmbH, Darmstadt, Germany.
g.
CodonCode Aligner, CodonCode Corp., Dedham, MA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by EPIZONE, the EU Network of Excellence for Epizootic Disease Diagnosis and Control (contract FOOD- CT-2006-016236).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
