Abstract
We aimed to validate the use of 1) the modified agglutination test and a polymerase chain reaction (PCR) protocol in detecting Toxoplasma gondii infection in quenda (Isoodon obesulus) and brushtail possums (Trichosurus vulpecula); 2) immunofluorescence microscopy of feces and a PCR and sequencing protocol in detecting Giardia spp. infection in quenda; and 3) a fecal flotation protocol in detecting gastrointestinal helminth infections of quenda. Quenda and brushtail possum carcasses, and samples from trapped quenda, were tested with 2 parasite detection tests per parasite, and results were modeled using Bayesian latent class analysis to estimate test sensitivity and specificity. The modified agglutination test and the PCR protocol were highly specific at detecting T. gondii infections in quenda and brushtail possums (≥93%); however, data were insufficient to assess sensitivity with adequate precision. Immunofluorescence microscopy and the PCR and sequencing protocol were both highly specific at detecting Giardia spp. in quenda (≥96%), but the PCR and sequencing protocol was relatively insensitive (69%, 95% credible interval [CrI]: 60–77%) compared to the highly sensitive immunofluorescence microscopy (98%, 95% CrI: 93–99%). The fecal flotation protocol was generally highly specific in the detection of gastrointestinal helminth infections (≥94%, with the exception of Trichuris spp. (88%, 95% CrI: 71–99%). The fecal flotation protocol was moderately to highly sensitive (≥74%) in the detection of strongyles, Labiobulura spp., Linstowinema spp., and Trichuris spp. Sensitivity was poor for detection of the cestode genus Potorolepis (36%, 95% CrI: 14–67%).
Introduction
In epidemiologic investigations of infections in wildlife populations, validation of the test(s) used to detect the infectious agent in the host population under study is required for scientific rigor. Estimates of detection test sensitivity and specificity are essential for data interpretation and are valuable at the study design stage. A lack of validated detection tests is a substantial hindrance to the investigation of infectious disease epidemiology in wildlife worldwide, 38 and this is particularly true of parasitic infection epidemiology in Australian marsupial species. The lack of validated tests corresponds to a lack of “gold standard” tests (tests of 100% sensitivity and specificity) against which other tests can be compared in marsupial hosts. This deficiency may be overcome, however, by using Bayesian latent class analysis, which allows for estimation of detection test sensitivity and specificity in the absence of a gold standard. 18
The protozoan parasite Toxoplasma gondii is of particular epidemiologic interest in Australian marsupial species. 17 The marsupials quenda (syn. southern brown bandicoot, Isoodon obesulus) and brushtail possums (Trichosurus vulpecula) are susceptible to infection with T. gondii,6,30,31 and may be at particular risk of infection with this parasite given their common occurrence in urban environments.15,40 However, there is not yet a validated test for use in antemortem identification of T. gondii infection in quenda or brushtail possums or in any member of their respective order (Peramelemorphia) or suborder (Phalangeriformes). Similarly, quenda are susceptible to infection with parasites of the genus Giardia,1,41 which is of epidemiologic interest as a result of potential conservation and public health implications.40,41 However, to our knowledge, no studies have investigated the accuracy of screening tools for detection of Giardia spp. infection in this species nor in any other species in the order Peramelemorphia. Finally, although fecal flotations of helminth eggs are common methodologies used in identifying gastrointestinal (GI) helminths present in marsupials sampled antemortem for epidemiologic purposes,9,33,42 no fecal flotation protocol has been validated in any Australian marsupial species.
We used Bayesian latent class analysis to validate the use of the following parasite detection tests in quenda and brushtail possums: 1) the modified agglutination test and a polymerase chain reaction protocol (PCR) protocol in detecting T. gondii infection in both host species; 2) immunofluorescence microscopy and a PCR and sequencing protocol in detecting Giardia spp. infection in quenda; and 3) a fecal flotation protocol in detecting GI helminth infections in quenda.
Materials and methods
Sixty-two quenda and 22 brushtail possum carcasses originating from the Statistical Division of Perth (Western Australia) were obtained opportunistically across 2013–2014. Sources included a wildlife hospital and animals found dead (e.g., killed by vehicle strike). Seventeen quenda carcasses and 1 possum carcass were sampled within 72 h of being found, after storage at 4°C. Forty-five quenda carcasses and 21 possum carcasses were frozen prior to collection of samples. Frozen carcasses were left to defrost at 4°C for 12–36 h prior to sample collection (time dependent on body size). Likely cause of death was determined based on clinical signs prior to death (where carcasses were obtained from the wildlife hospital), circumstances surrounding the death, and gross findings on postmortem examination.
For each animal, sex and whether the animal was an adult or subadult were recorded. Male possums were classified as adult if at least 1 testis was >1 cm wide, or subadult if each testis was ≤1 cm wide. Male quenda were classified as adult if their scrotal width was ≥2 cm, or subadult if their scrotal width was <2 cm. Female possums and quenda were classified as adult if they had a parous pouch, or as subadult if they had a non-parous pouch. In adult females, pouch activity was noted, and adult females were classified as having an active pouch if lactating, or an inactive pouch if not lactating.
Where available, the following samples were collected from each carcass:
Hemorrhagic or serohemorrhagic chest cavity fluid was obtained, centrifuged at 314 × g for 10 min, then the serous top layer of the sample (if available) or the top half of the sample (if a serous layer was not available) was transferred to a storage tube and frozen at −20°C until analysis.
Organ samples: brain tissue was removed via the foramen magnum and stored in 70% ethanol (at a ratio of at most 1:4), and 2 mL of heart, liver, tongue, and diaphragm tissue (or the whole organ, if total volume was lower) was cut to ~1 cm width and added to separate tubes of 8 mL of 70% ethanol. All organ samples were stored at 4°C until analysis.
Feces from the large intestine (quenda only): 2 mL of feces were added to 8 mL of 10% buffered formalin and mixed thoroughly. One milliliter of feces was added to 8 mL of 70% ethanol and mixed thoroughly. Samples were stored at 4°C until analysis. Similarly, fecal samples were obtained from quenda trapped as part of a parasite survey in the Statistical Division of Perth. Trapped quenda had age, sex, and pouch activity recorded, as described for the quenda carcasses.
GI helminths (quenda only): the entire GI tract and its contents were screened for helminths under a dissection microscope. After the fecal flotations were completed, helminths were also recovered from the feces used for the flotation. All observed nematodes were removed and stored in 70% ethanol. Cestodes were removed and stored in 10% buffered formalin. Helminths were stored at 4°C until analysis.
All samples were obtained under Murdoch University Animal Ethics Permit R2530/12, and Department of Parks and Wildlife Regulation 17 (SF009640) and Regulation 4 (CE004287) permits.
Chest fluid samples from quenda and possum carcasses were tested using a commercial modified agglutination test kit, a according to the manufacturer’s instructions. The animal was considered to have tested positive on the modified agglutination test if it tested positive at titers of 1:40 and/or 1:4,000. The same observer performed all of these tests.
For the T. gondii PCR, DNA was extracted from tissue samples using a blood and tissue kit, b with some variations. A freeze–thaw step was conducted prior to addition of proteinase K, to facilitate disruption of T. gondii tissue cysts. Tissue was macerated in 50 µL of phosphate-buffered saline (PBS) in a screw-cap tube, 180 µL of buffer ATL was added, and tubes were subjected to 8 cycles of alternating between 1 min in liquid nitrogen and 1 min in a 95°C water bath. Proteinase K was added, samples were incubated overnight at 56°C, and the protocol was then continued according to the manufacturer’s instructions, with a final elution volume of 50 µL. Eluted DNA was stored at −20°C until required for PCR. A positive control (T. gondii–infected quenda liver) and negative control (PBS) were included in each batch of DNA extractions.
The target for PCR amplification of T. gondii DNA was the B1 gene, using a nested PCR with the following primers: external forward (5′-TGTTCTGTCCTATCGCAACG) and external reverse (5′-ACGGATGCAGTTCCTTTCTG); internal forward (5′-TCTTCCCAGACGTGGATTTC) and internal reverse (5′-CTCGACAATACGCTGCTTGA). 11 Each reaction of the nested PCR was performed in a 25-µL volume consisting of 1–2 µL of extracted DNA, 1.5 mM MgCl2, 1× reaction buffer, c 200 µM of each dNTP, 0.4 µM of each appropriate forward and reverse primer, and 0.2 units of Taq DNA polymerase c in a cresol loading dye solution. PCR was performed in a thermocycler, d with the following conditions: 95°C for 1 min, 35 cycles of 95°C for 30 s, 60°C for 30 s, and 72°C for 1 min, followed by a final extension of 72°C for 5 min. Each PCR plate contained a positive control (purified DNA from cultured T. gondii tachyzoites) and negative controls. PCR products were either used immediately, or stored at 4°C until required for either the internal PCR or visualization. After the internal PCR amplification, PCR products were separated by electrophoresis on 2% agarose gel containing DNA stain, e and gels were photographed under ultraviolet light. Carcasses were considered to have tested positive by PCR if specific DNA bands of the correct product size for T. gondii were amplified from any or all of the brain, heart, diaphragm, liver, or tongue. The same observer performed all PCR tests.
Of the nonspecific DNA bands that were sometimes amplified by PCR (faints bands that were not the correct product size for T. gondii), the 10 nonspecific bands that were closest to the size of T. gondii were sequenced to ensure that T. gondii had not been amplified. These DNA samples were purified using a purification system, f following the manufacturer’s instructions. Samples were sequenced in both directions using a sequencing kit g on a DNA analyzer. h Sequences were analyzed using sequencing software. i
Immunofluorescence microscopy was used to identify Giardia spp. cysts in fecal samples obtained from trapped quenda and quenda carcasses, using a commercial kit, j following the manufacturer’s instructions for a nonconcentrated sample. Samples were considered positive on immunofluorescence microscopy if at least 1 cyst of appropriate fluorescence and Giardia spp. morphology was identified. The same observer performed all immunofluorescence microscopy tests.
PCR and sequencing was undertaken on all immunofluorescence microscopy–positive samples and a random sample of 10 immunofluorescence microscopy–negative samples. Fecal samples that were preserved in 70% ethanol had DNA extracted for amplification of Giardia spp. at 3 loci (18S rRNA, ITS1-5.8s-ITS2, and gdh), and all amplified products on PCR were subject to sequencing, as described previously. 16 Quenda were considered positive for Giardia spp. via this PCR and sequencing protocol if genetic material was amplified and sequenced as Giardia spp., at 1 or more loci. The same observer performed all PCR work.
As an addendum to the Giardia spp. validation, during the fecal flotations for identification of GI helminth eggs (described below), the microscopist assessed the sample for the presence of Giardia spp. cysts.
For the fecal flotation protocol, each formalin-preserved quenda fecal sample originating from a carcass was centrifuged at 850 × g for 2 min, with the formalin supernatant discarded. The sample was then divided between two 10-mL centrifuge tubes. The samples were rehomogenized in 9 mL of distilled water, and centrifuged at 850 × g for 2 min, with the supernatant discarded.
One tube was analyzed using zinc sulfate flotation. Feces were thoroughly mixed with 9 mL of zinc sulfate solution (specific gravity [SG] 1.20), then centrifuged at 850 × g for 2 min. A flamed wire loop was used to transfer material from the surface of the flotation to a slide, and a coverslip was added. Slides were systematically scanned at 200× magnification via a light microscope. k The second tube was analyzed using sodium nitrate flotation. Feces were mixed thoroughly with 9 mL of sodium nitrate solution (SG 1.37) and centrifuged at 850 × g for 2 min. The tube was then topped up with sodium nitrate solution to create a meniscus. A coverslip was placed on top of the tube and left for 10 min, then the coverslip was placed on a slide and systematically scanned at 200× magnification via a light microscope. k Before the first sodium nitrate slide was examined, the sodium nitrate tube was topped up with sodium nitrate solution again to create another meniscus, and a second coverslip was placed on top. This was transferred to a slide and examined at 100× magnification via a light microscope, k directly after the first sodium nitrate flotation slide had been examined (a pilot study indicated that examining a second slide from the sodium nitrate tube substantially increased the likelihood of detecting Trichuris spp. eggs in quenda feces, AE Hillman, unpublished data).
Helminth eggs were identified morphologically, using features and size ranges described in the literature22,36,43 and in microscopic photography records collated from prior quenda fecal analyses (AD Elliot, unpublished data). Quenda were considered positive for the respective helminth taxon by the fecal flotation protocol if at least 1 egg of appropriate morphologic features was identified on any or all of the 3 fecal flotations performed as part of the protocol. All fecal flotations were undertaken by the same observer. To aid observers using this fecal flotation protocol in the future, photographic records of eggs of the respective helminth taxa, and ranges of egg length and width, were obtained. Of all helminth taxa eggs observed by fecal flotation, at least 1 egg per taxon per host was measured microscopically at 400× magnification to include in reported ranges of egg length and width. However, the reported ranges of Trichuris spp. egg length and width were restricted to dimensions of eggs obtained from the feces of quenda from which Trichuris spp. worms were obtained from the GI tract. Photographs and measurements were undertaken at 400× magnification.
The GI helminths obtained from the quenda carcasses via the microscopic GI tract helminth screen were differentiated to genus, family, or suborder level by light microscopy k (at 4× to 200× magnification, depending on the worm genus/family/suborder), using published quenda helminth descriptions.22,36,43 Where cestodes were too degraded to have retained identifying features (particularly rostellar hooks), the morphologic features of eggs obtained from gravid proglottids were used to identify the genus of cestode. If gravid proglottids were not present, and thus the cestode was not identifiable morphologically, the sample was excluded from cestode sensitivity and specificity calculations. Quenda were considered to have tested positive for the respective nematode taxon on microscopic GI tract examination if at least 1 nematode of the taxon was obtained on microscopic GI tract examination and differentiated as such morphologically. Quenda were considered to have tested positive for a cestode genus on microscopic GI tract examination if at least 1 cestode of that genus was obtained and differentiated as such morphologically, or if eggs of that cestode genus were obtained and identified from gravid proglottids of a cestode that was not otherwise morphologically identifiable. All GI helminth identifications were undertaken by the same observer.
The sensitivity and specificity of the parasite detection tests were calculated via Bayesian latent class analysis, using a software package l with a software interface. m Briefly, in the Bayesian latent class analyses used in our study (where 2 detection test results are available per animal, with tested animals obtained from 1 population), 5 parameters require consideration: infection prevalence, and the sensitivity and specificity of each test. Parameters are entered as either informative or noninformative distributions. As the number of parameters exceeds the 3 degrees of freedom provided by the data, informative prior distribution inputs are required for at least 2 of the 5 model parameters for model identifiability. Published data (where available) and/or expert knowledge can be used to guide informative (beta) prior distribution inputs. Noninformative (uniform) prior distributions can be used where no information is available to guide prior distribution estimations. The prior distribution inputs are then modeled with the detection test data obtained from the study and the unknown latent variable (representing the true infection status of the host) using the iterative Gibbs sampler Markov chain Monte Carlo method, to obtain posterior distribution estimates of the sensitivity and specificity of each detection test and of the infection prevalence 18 (although in this study, prevalence is not of interest as an output).
For the Bayesian latent class analysis of the T. gondii tests, tests were modeled as conditionally independent. The prior distribution inputs were identical for the quenda and brushtail possum models (Table 1). The relatively conservative prior distribution for modified agglutination test specificity (0.50–0.99) was based on findings from several macropod species (taxonomic family Macropodidae). An enzyme-linked immunosorbent assay (ELISA) was validated as highly specific, and was shown to have excellent agreement with the modified agglutination test 28 (see also Parameswaran N. Toxoplasma gondii in Australian marsupials [PhD thesis]. Perth, Australia: Murdoch University; 2008:52). The more precise prior distribution used for PCR specificity was based on previous research findings at the B1 locus (Table 1).
Prior distribution inputs for Bayesian latent class analysis of parasite detection tests in quenda (Isoodon obesulus) and brushtail possums (Trichosurus vulpecula).*
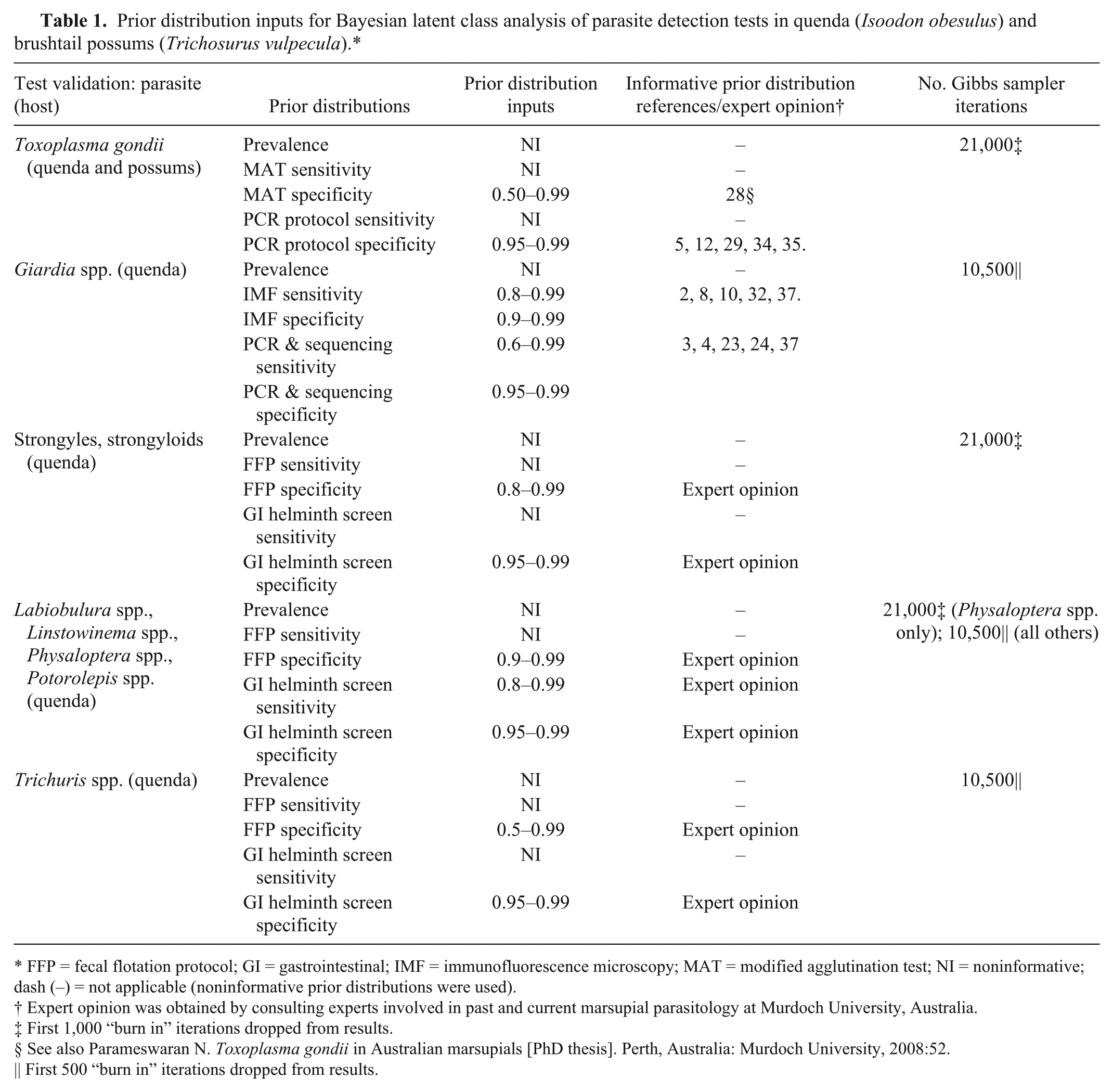
FFP = fecal flotation protocol; GI = gastrointestinal; IMF = immunofluorescence microscopy; MAT = modified agglutination test; NI = noninformative; dash (–) = not applicable (noninformative prior distributions were used).
Expert opinion was obtained by consulting experts involved in past and current marsupial parasitology at Murdoch University, Australia.
First 1,000 “burn in” iterations dropped from results.
See also Parameswaran N. Toxoplasma gondii in Australian marsupials [PhD thesis]. Perth, Australia: Murdoch University, 2008:52.
First 500 “burn in” iterations dropped from results.
For the Bayesian latent class analysis of the Giardia spp. tests, tests were modeled as conditionally independent, as the widely reported high sensitivity and specificity of Giardia spp. detection via immunofluorescence microscopy of feces in other species indicated that conditional dependence between tests was unlikely to affect these models (Table 1). The relatively conservative informative prior distribution used for PCR and sequencing sensitivity was chosen in view of the variable results observed via PCR in different studies of Giardia spp. that used various protocols on a variety of host species. 39 The high and narrow prior distribution used for PCR and sequencing specificity was selected, given that the use of sequencing in our case definition made false positive results highly unlikely.
For the Bayesian latent class analysis of the GI helminth tests, tests were modeled as conditionally independent (Table 1). All informative priors were based on expert opinion, obtained in consulting expert personnel involved in past and current marsupial parasitology at Murdoch University. The informative prior distributions used for the specificity of the fecal flotation protocol in detection of Labiobulura spp., Linstowinema spp., Physaloptera spp., and Potorolepis spp. were recommended based on the distinctive morphologic features of these helminth eggs, rendering false positives unlikely. The prior distribution for the fecal flotation protocol specificity for the detection of strongyles (suborder Rhabditina; likely representing family Trichostrongylidae, although other families cannot be excluded) and strongyloids (family Strongyloididae) was recommended in consideration of the possibility that parasite eggs of quenda prey items may be present in the quenda feces; there are strongyle and strongyloid helminths with similar eggs parasitic in a wide range of species. The prior distribution for the fecal flotation protocol specificity for the detection of Trichuris spp. infection was recommended in consideration of the risk of eggs of other capillarids (superfamily Trichinelloidea, e.g., Capillaria spp.) being mistaken for those of Trichuris spp. The prior distributions for the sensitivities of the observer finding Labiobulura spp., Linstowinema spp., Physaloptera spp., and Potorolepis spp. helminths in the GI tract of infected quenda and correctly differentiating them to genus level were recommended based on the relatively large size of adult helminths, and the reasonably uncomplicated taxonomic differentiation required, rendering false negatives relatively unlikely. Uninformative prior distributions were used for sensitivity of detection and differentiation of strongyles, strongyloids, and Trichuris spp., based on the tiny size (strongyles and strongyloids) and the tiny (coiled) size and apparent fragility of Trichuris spp. worms after recovery from frozen carcasses, causing uncertainty about the potential for false negatives. The informative prior distribution inputs for the specificity of the observer finding helminths in the quenda GI tract and correctly differentiating to a genus/family/suborder level were recommended based on the distinctiveness of the helminth morphology at the taxonomic level of differentiation required, rendering false positives unlikely.
The number of Gibbs sampler iterations for each model started at 10,500 iterations, with the first 500 “burn in” iterations dropped from the results. The number of iterations was doubled if the model did not adequately converge using the lower number of iterations (Table 1).
Model validity for each infection type was established in a number of ways. First, Gibbs sampler trace plots were evaluated to ensure convergence, and models were rerun 5 times with different starting values to ensure consistency of output. Second, the observed versus the median predicted test profiles were compared to ensure they were similar. Finally, the observed and expected probabilities of agreement between the tests were compared to ensure that values were similar and that the associated probability statistic was between 0.05 and 0.95.
The median value of the modeled test sensitivities and specificities were reported as the point estimates, with 95% credible intervals (CrI; Bayesian analogues of frequentist confidence intervals). Where fewer than 6 animals tested positive or negative for a parasite taxon by 1 or both tests, data were considered insufficient for sensitivity or specificity calculations (respectively), and were not reported in results. The prevalence outputs, and the output of the sensitivity and specificity of the observer finding and correctly identifying helminths to a genus/family/suborder level were also not reported. This was because our study was not designed to measure infection prevalences, and although the data provided by the observer obtaining and differentiating helminths were valuable components of the latent class analyses, the sensitivity and specificity of this test is not of practical value to future research.
Sensitivity analyses of all models were undertaken, by varying each informative prior distribution separately, while all other model inputs were held constant. The credible limits of the informative priors were decreased by 10% and 20% (gross), respectively. A change in the sensitivity and/or specificity point estimate by 5% (gross) or more was considered “substantial,” and reported in Results.
Results
Of quenda carcasses used in our study, 54 of 62 (87%) were judged to have been killed by vehicle strike, attack by another animal, household poisons (such as rat and snail baits), or drowning in backyard pools or ponds. Of possum carcasses used in our study, 15 of 22 (68%) were judged to have been killed by traumatic injuries (particularly vehicle strike or attack by another animal), electrocution, or household poisons (such as rat and snail baits).
Chest fluid and body tissues were available from 56 quenda carcasses and 22 brushtail possum carcasses. The quenda carcasses comprised 6 subadult females, 12 adult females, 21 subadult males, 16 adult males, and an additional subadult quenda of undetermined sex. Of the adult female quenda, 9 had an active pouch and 3 had an inactive pouch. The possum carcasses comprised 2 subadult females, 9 adult females, 4 subadult males, and 7 adult males. Of the adult female possums, 2 had an active pouch and 7 had an inactive pouch.
All 5 targeted body tissues (brain, heart, liver, diaphragm, and tongue) were available from 51 quenda carcasses and all brushtail possum carcasses. Because of traumatic injuries, diaphragmatic tissue was not recoverable from 3 quenda, heart tissue was not recoverable from 1 quenda, and sampling of brain tissue was severely compromised in 1 quenda. All carcasses positive on the modified agglutination test had all 5 tissues available for testing.
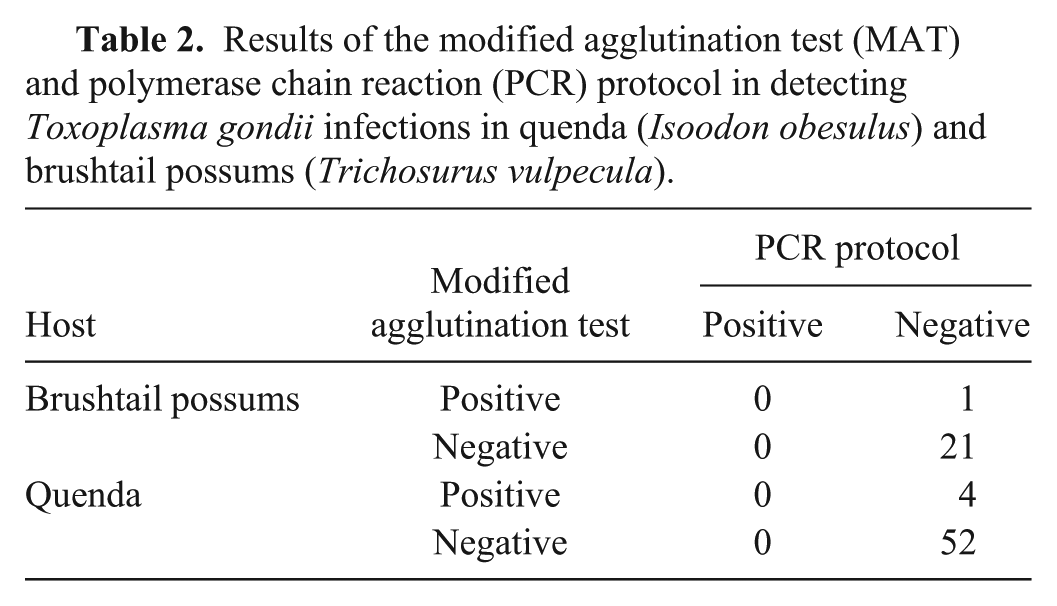
The T. gondii tests in brushtail possums were 96% concordant (21 of 22 possums). One possum was positive on the modified agglutination test; bands specific for T. gondii were not amplified from any brushtail possum tissues. In quenda, the T. gondii tests were 93% concordant (52 of 56 quenda). Four quenda were positive on the modified agglutination test; bands specific for T. gondii were not amplified from any quenda tissues (Table 2). On sensitivity analysis, no substantial changes occurred to the point estimates of the specificity of either T. gondii test in either species (Table 3).
Results of the modified agglutination test (MAT) and polymerase chain reaction (PCR) protocol in detecting Toxoplasma gondii infections in quenda (Isoodon obesulus) and brushtail possums (Trichosurus vulpecula).
Sensitivity and specificity of various parasite detection tests in quenda (Isoodon obesulus) and brushtail possums (Trichosurus vulpecula).*
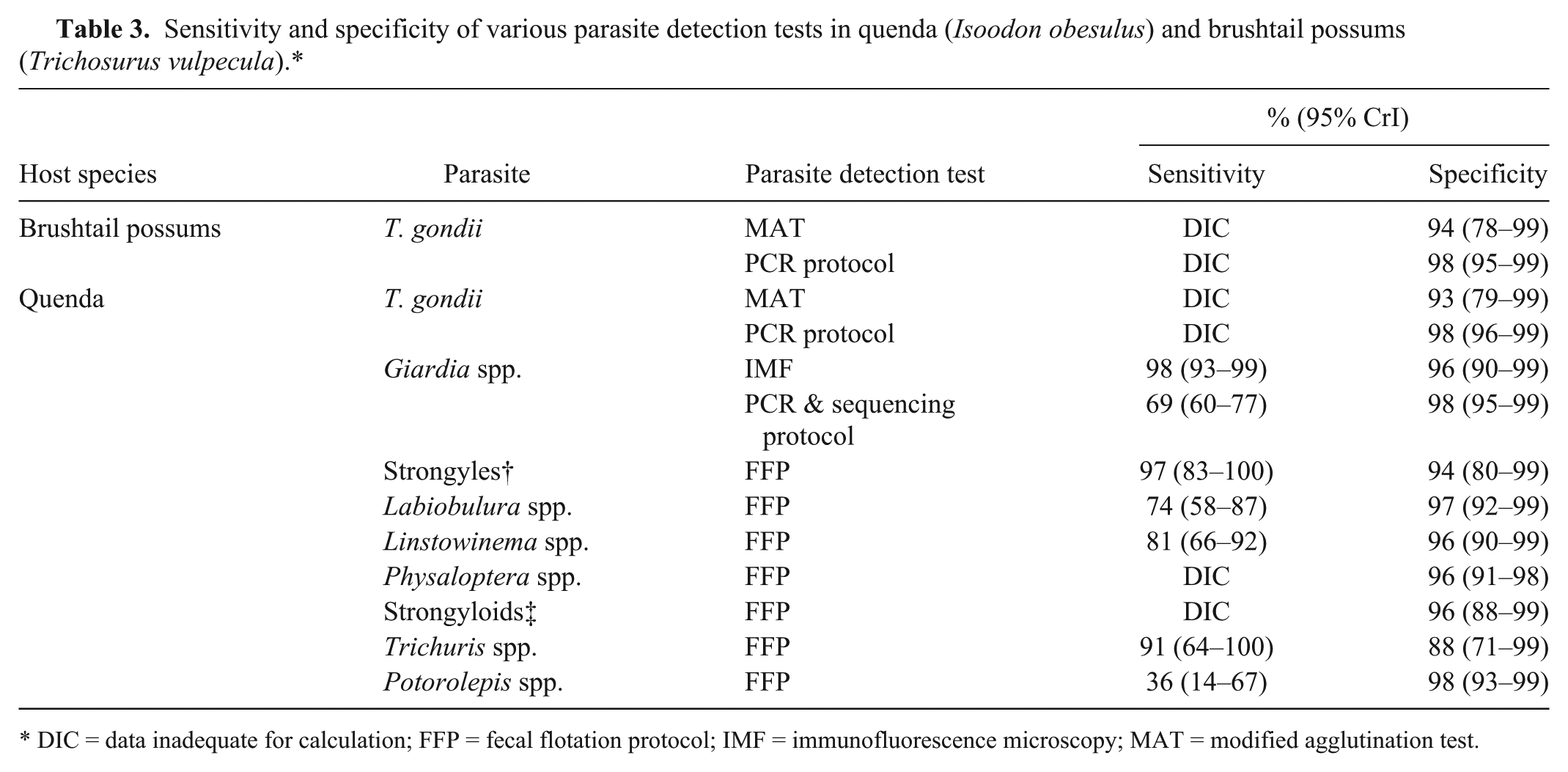
DIC = data inadequate for calculation; FFP = fecal flotation protocol; IMF = immunofluorescence microscopy; MAT = modified agglutination test.
The 10 faint, nonspecific bands amplified by T. gondii PCR that were closest to the expected size for T. gondii (from 10 tissues obtained from 6 quenda) did not sequence as T. gondii. One of the 22 possums and 1 of 56 quenda were known to have nonspecific neurologic signs prior to death. Both animals were negative on both the modified agglutination test and PCR.
Fecal samples available for the Giardia spp. test validation were from 110 immunofluorescence microscopy–positive quenda and 10 immunofluorescence microscopy–negative quenda. The samples were obtained from 6 subadult females, 42 adult females, 11 subadult males, and 61 adult males. Of the adult females, 33 had an active pouch and 9 had an inactive pouch. Eleven samples were from quenda carcasses; 109 samples were obtained from trapped quenda.
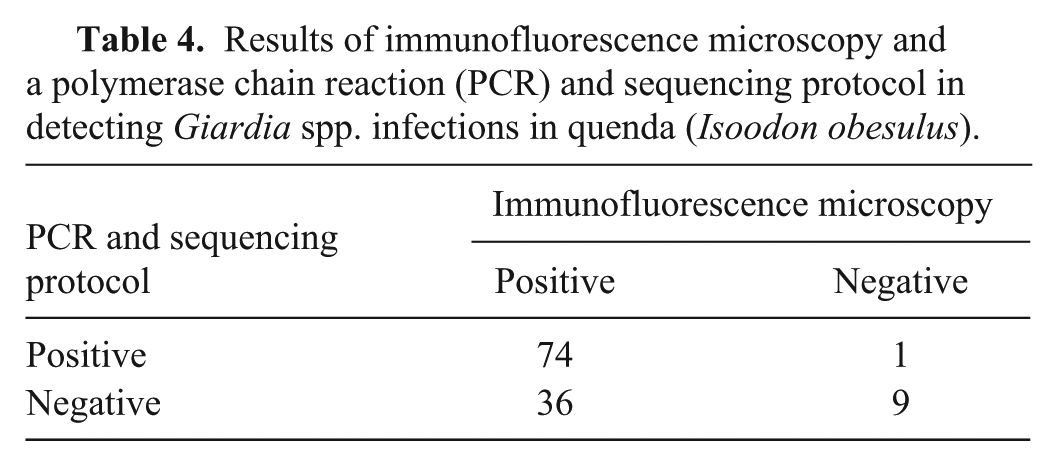
The Giardia spp. infection detection test results were concordant in 83 of 120 (69%) quenda (Table 4). Thirty-six quenda were positive for Giardia spp. infection on immunofluorescence microscopy, but negative by PCR and sequencing. Giardia spp. cysts were identified in 22 (61%) of these quenda, by light microscopy during the fecal flotations.
Results of immunofluorescence microscopy and a polymerase chain reaction (PCR) and sequencing protocol in detecting Giardia spp. infections in quenda (Isoodon obesulus).

On sensitivity analysis, no substantial changes occurred to the point estimates of sensitivity and specificity of both Giardia spp. tests (Table 3).
GI tract helminth screens and fecal flotation analyses were available from 49 quenda carcasses. This comprised 4 subadult females, 11 adult females, 19 subadult males, and 15 adult males. Of the adult female quenda, 2 had an inactive pouch and 9 had an active pouch.
Six nematode taxa, consisting of strongyles, Labiobulura spp., Linstowinema spp., Physaloptera spp., strongyloids, and Trichuris spp., were detected by fecal flotation (Figs. 1–7) and/or the microscopic GI helminth screen. The cestode genus Potorolepis was detected by fecal flotation (Figs. 8, 9) and the microscopic GI helminth screen. Three quenda carcasses contained unidentifiable cestode segments and were thus excluded from Potorolepis spp. results, and from the Potorolepis spp. sensitivity and specificity calculations. Cestode eggs were not identified in the feces of any of these 3 quenda.
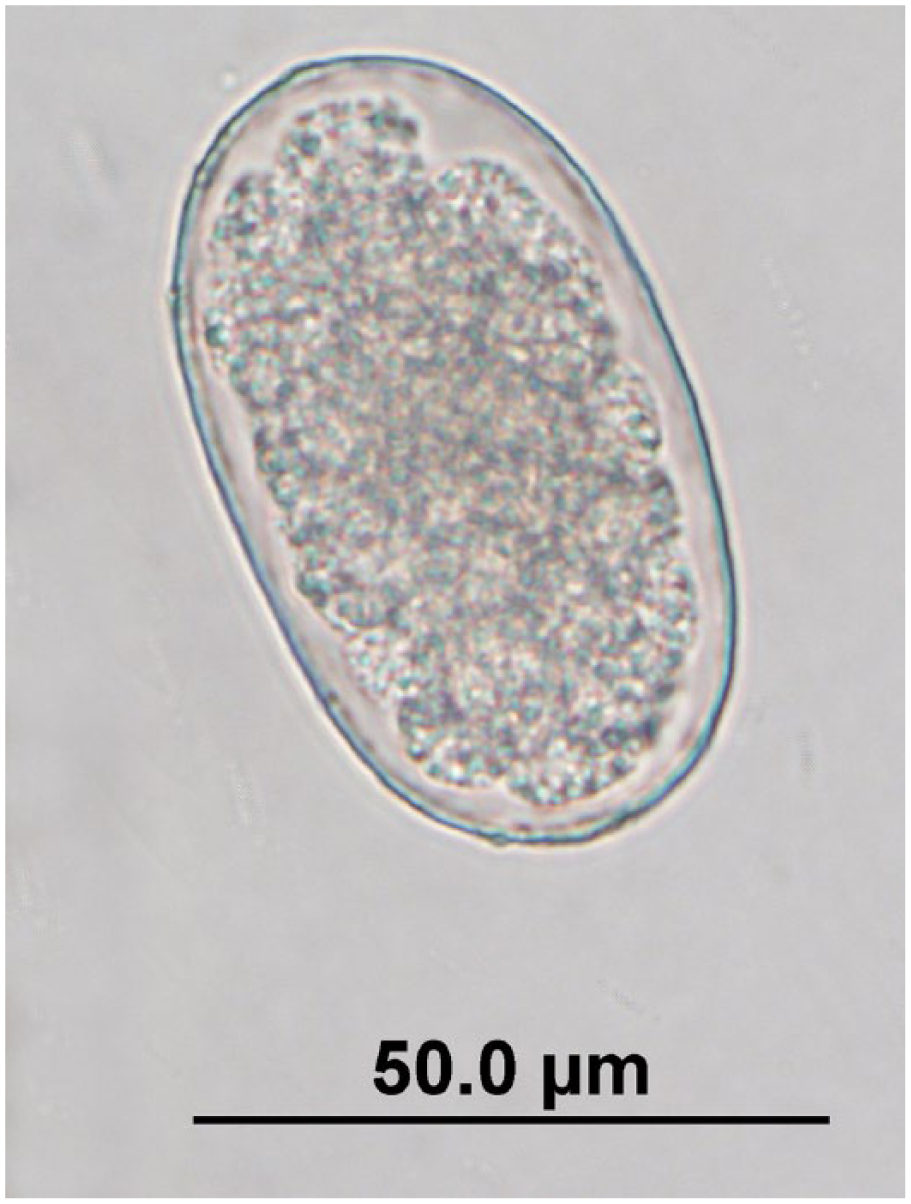
Unembryonated strongyle egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 52.5–72.5 µm; width: 30–45 µm.
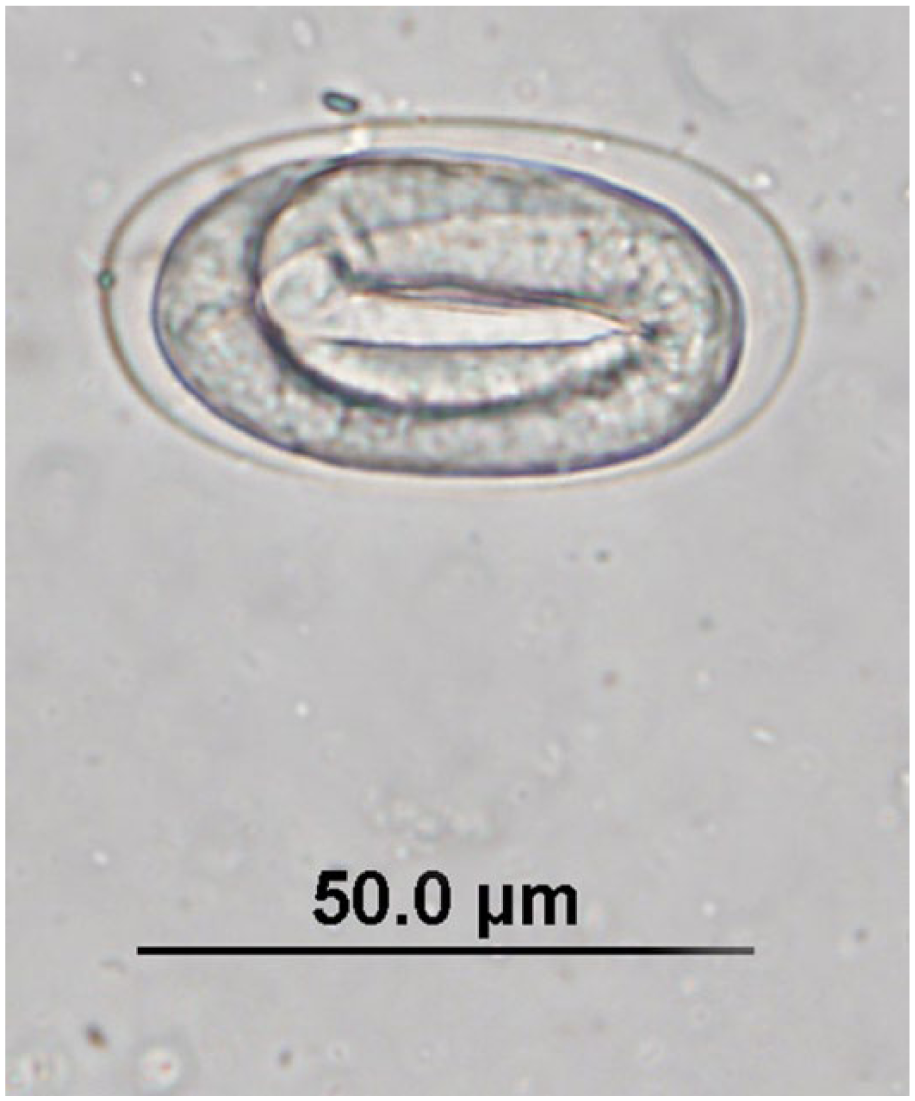
Embryonated strongyle egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 52.5–72.5 µm; width: 30–45 µm.
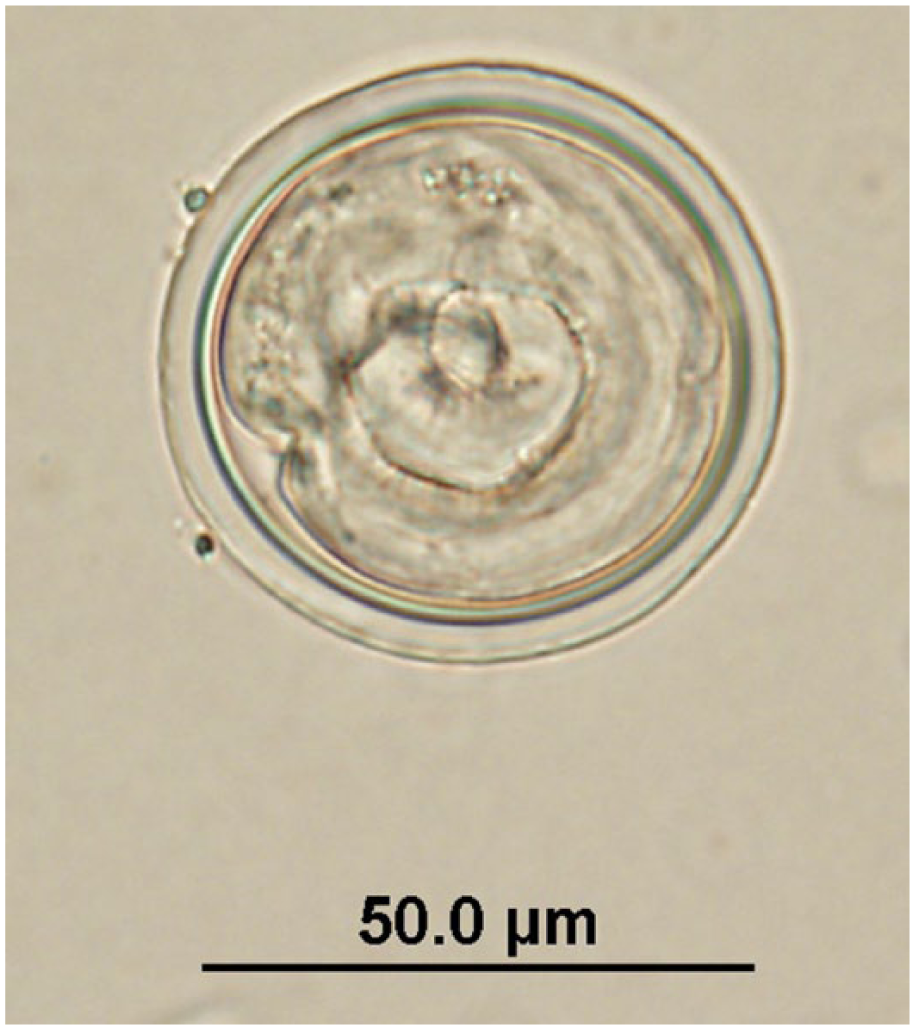
Labiobulura sp. egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 52.5–65 µm; width: 45–60 µm.
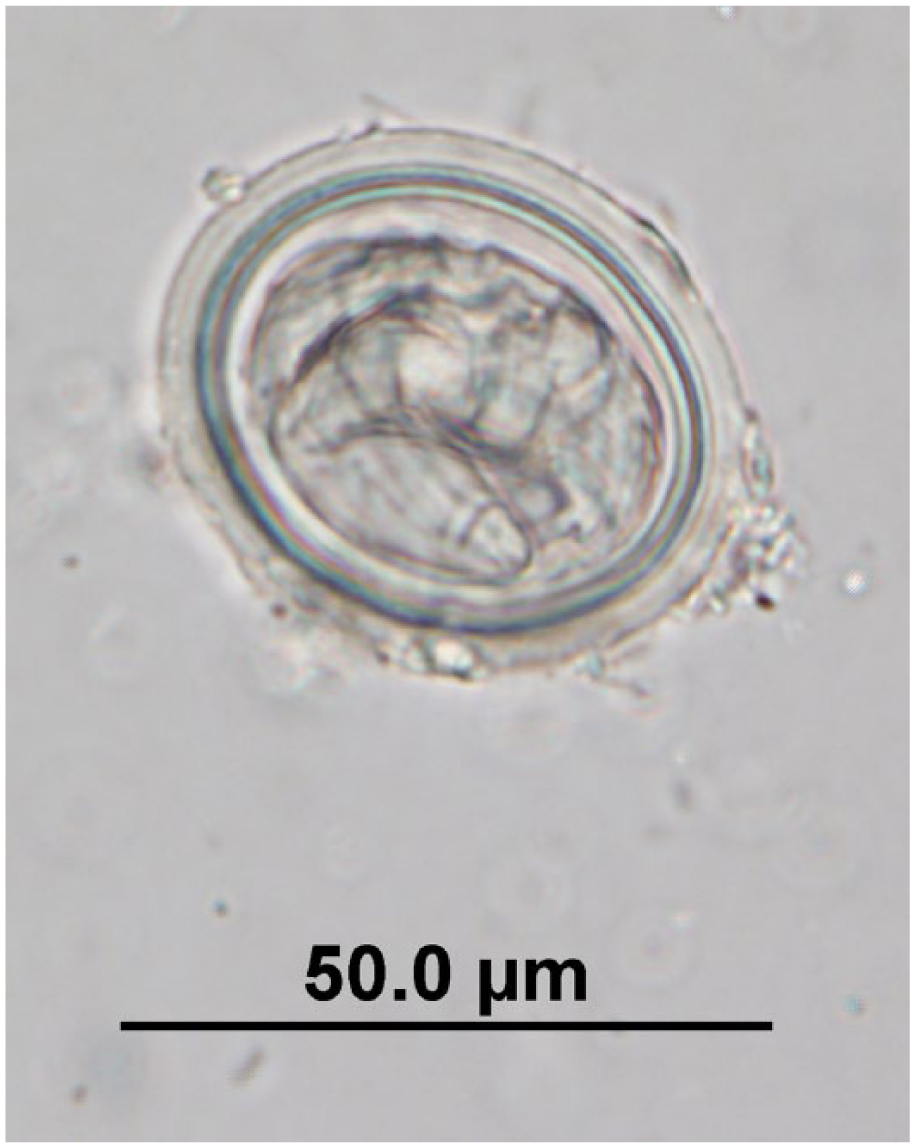
Linstowinema sp. egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 35–45 µm; width: 32.5–45 µm.
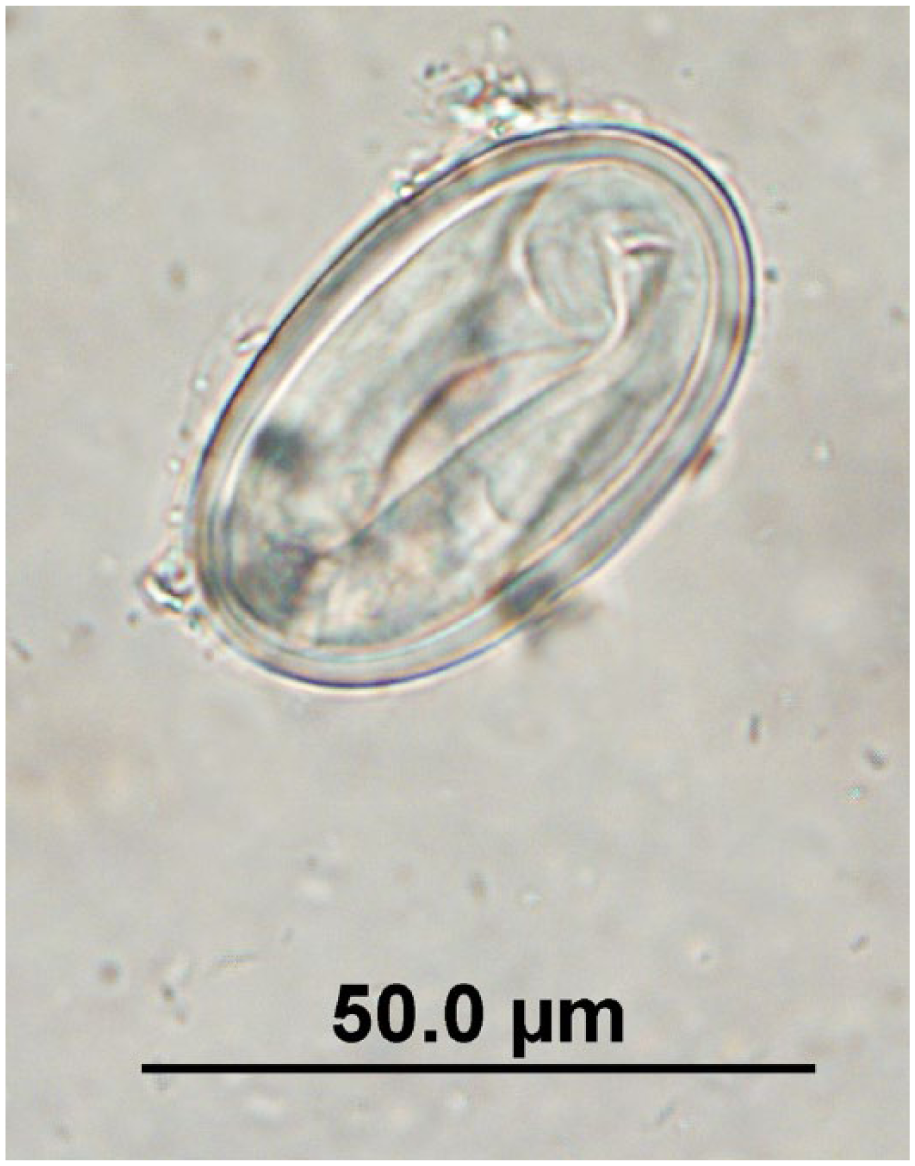
Physaloptera sp. egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 45–55 µm; width: 27–32.5 µm.
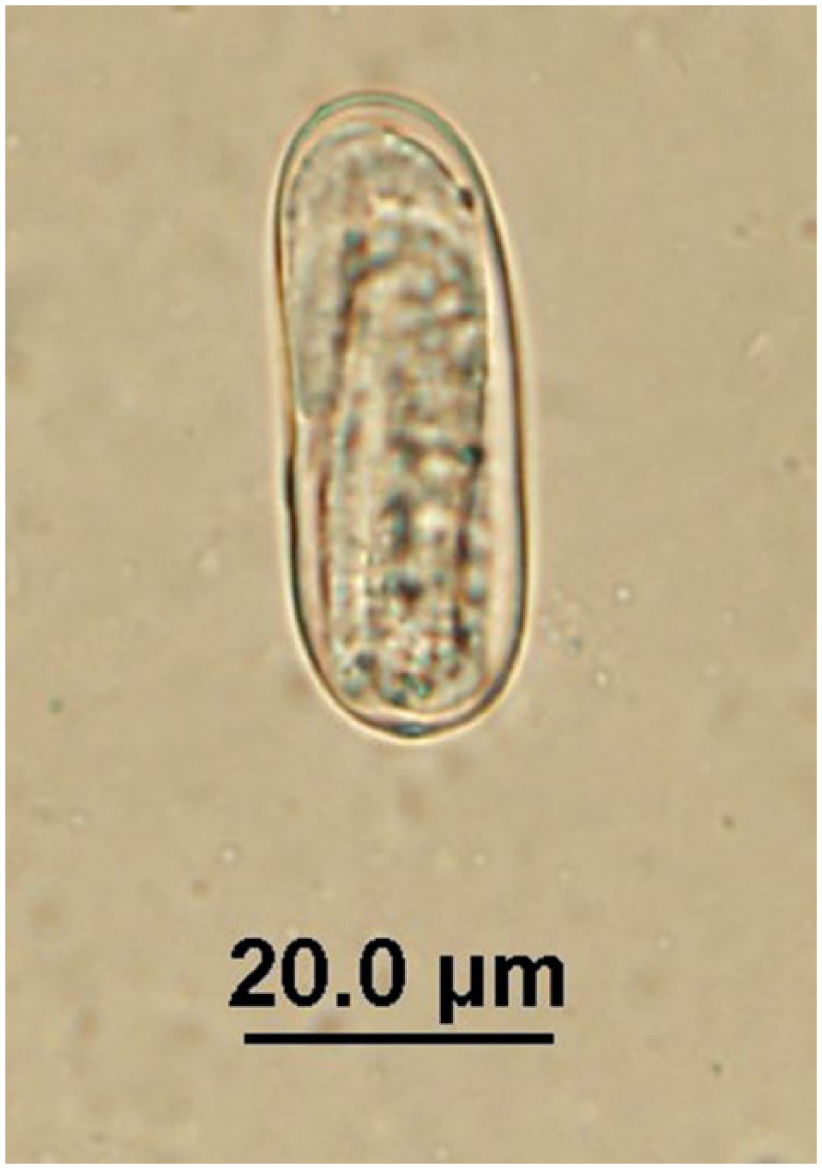
Strongyloid egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 40–45 µm; width: 15–20 µm.
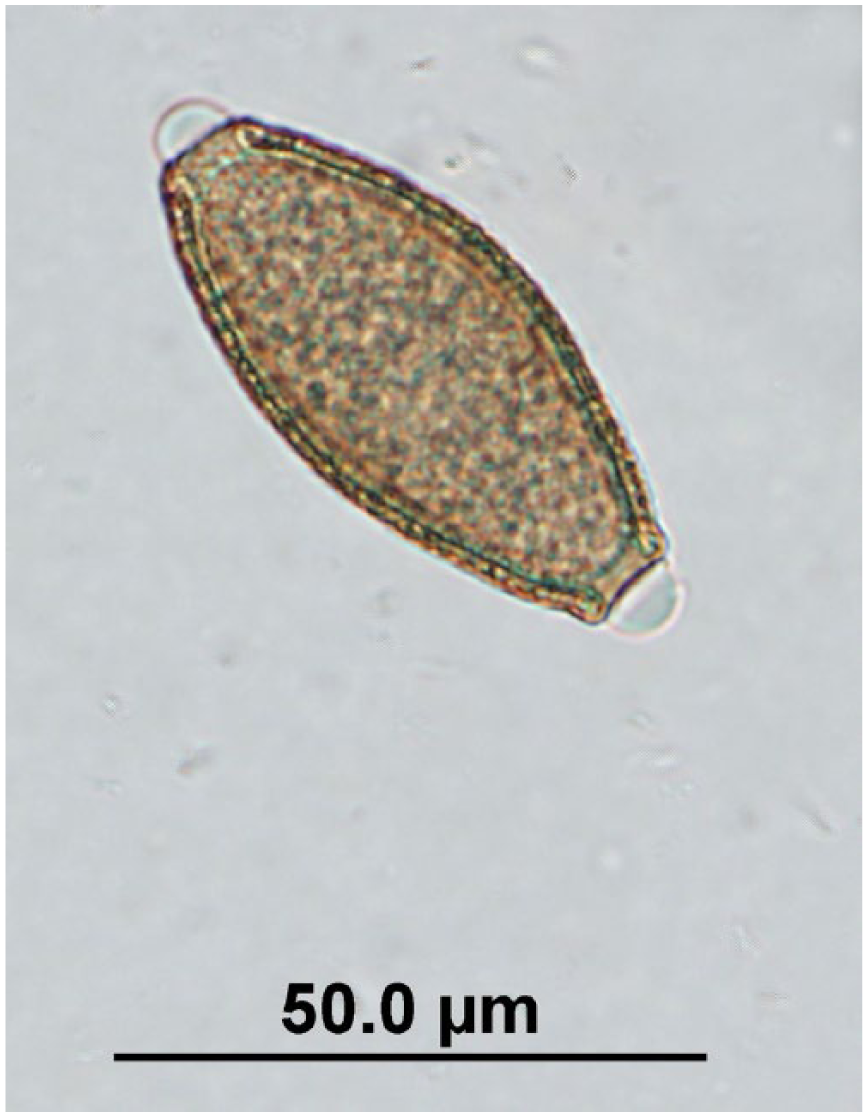
Trichuris sp. egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 47.5–60 µm; width: 22.5–27.5 µm.
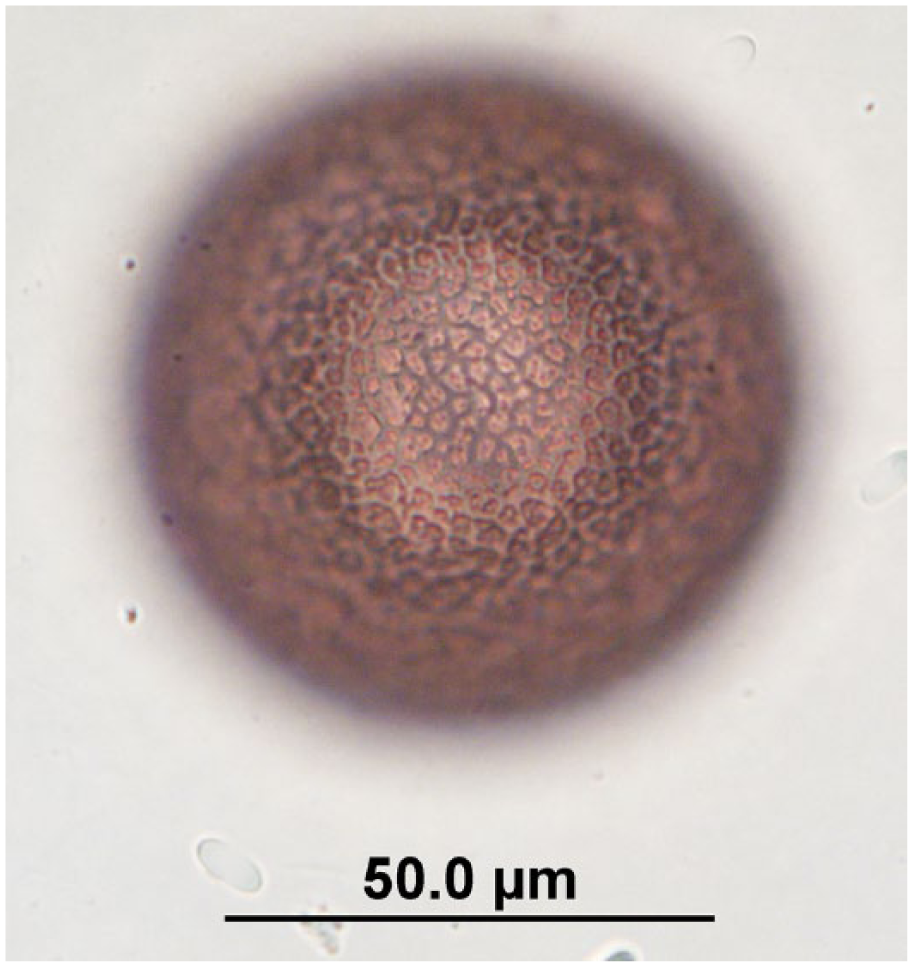
Potorolepis sp. egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 65–102 µm; width: 65–80 µm. Image is focused on the egg wall.
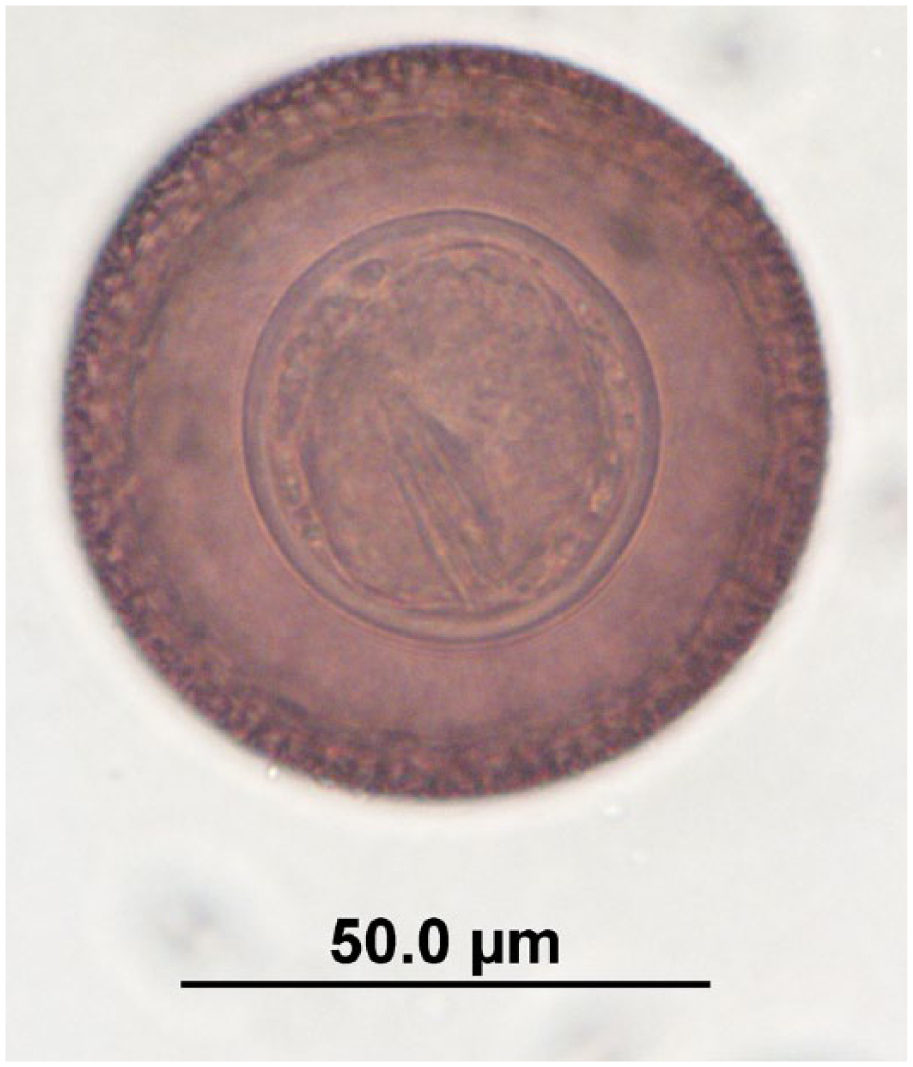

Potorolepis sp. egg observed in quenda feces by fecal flotation microscopy at 400× magnification; egg dimensions: length: 65–102 µm; width: 65–80 µm. Image is focused on the hexacanth embryo.
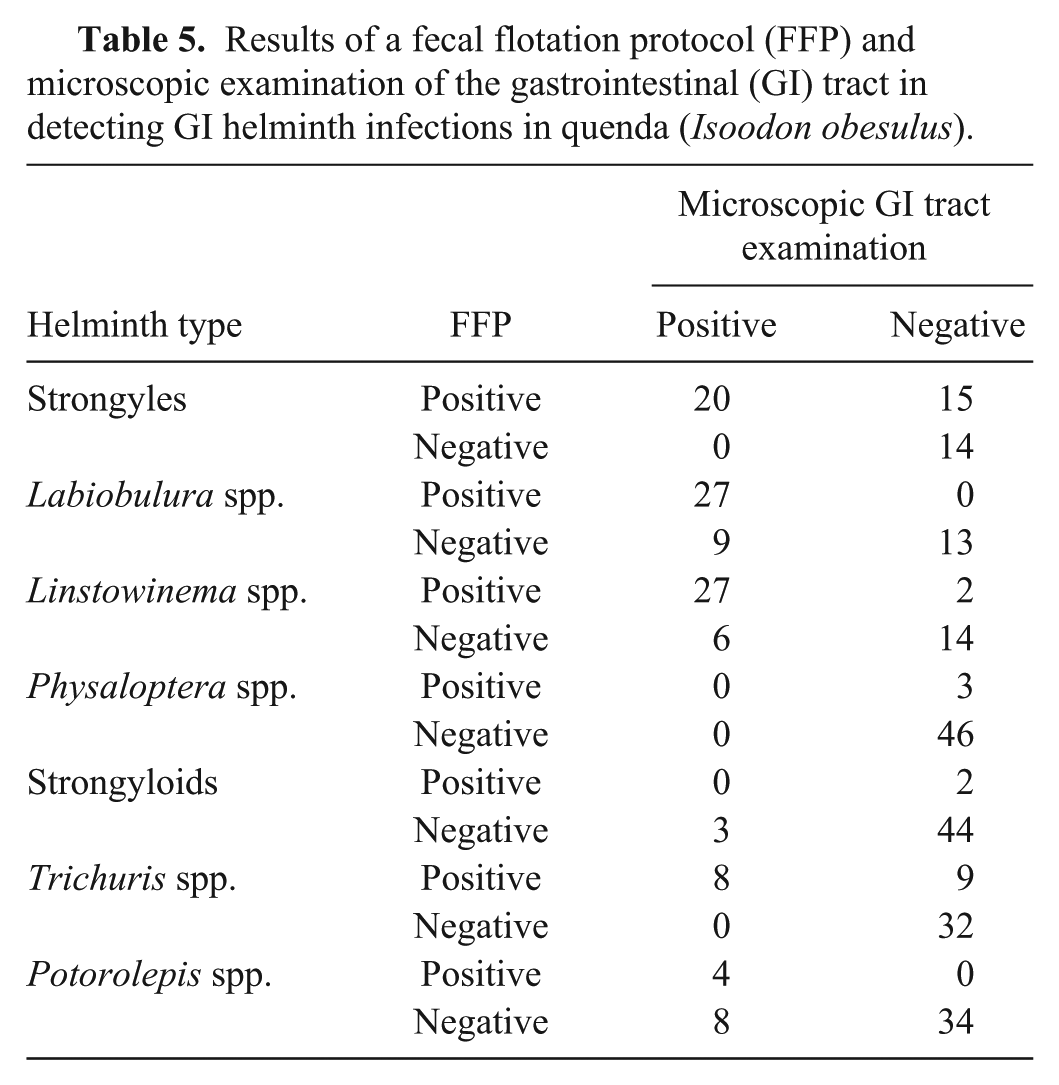
The concordances of the GI helminth detection tests were as follows: 34 of 49 (69%) for strongyles; 40 of 49 (82%) for Labiobulura spp.; 41 of 49 (84%) for Linstowinema spp.; 46 of 49 (94%) for Physaloptera spp.; 44 of 49 (90%) for strongyloids; 40 of 49 (82%) for Trichuris spp.; and 38 of 46 (83%) for Potorolepis spp. (Table 5).
Results of a fecal flotation protocol (FFP) and microscopic examination of the gastrointestinal (GI) tract in detecting GI helminth infections in quenda (Isoodon obesulus).
On sensitivity analysis, no substantial changes occurred to the point estimates of: the sensitivity and specificity of the fecal flotation protocol in detecting Labiobulura spp., Linstowinema spp., and Trichuris spp. infections; the sensitivity of the fecal flotation protocol in detecting strongyle infections; and the specificity of the fecal flotation protocol in detecting Physaloptera spp., strongyloids, and Potorolepis spp. infections (Table 3). When the lower credible limit of the informative prior distribution for the specificity of the fecal flotation protocol in identifying strongyle infections was extended to 0.6, the point estimate of specificity of fecal flotation in detecting strongyle infections decreased to 88% (95% CrI: 57–99%). When the lower credible limit of the informative prior distribution of the specificity of the microscopic GI tract exam in identifying Potorolepis spp. infections in quenda was widened to 0.75, the point estimate for the sensitivity of detecting Potorolepis spp. by the fecal flotation protocol increased to 45% (95% CrI: 16–92%).
Discussion
Our study presents the first validation data regarding parasite tests for use in the detection of Giardia spp. and GI helminth infections in any Australian marsupial species (superorder Australidelphia), and the first validation data for tests to detect T. gondii infection in any species of order Peramelemorphia and suborder Phalangeriformes. The robustness of the Bayesian models on sensitivity analysis suggests that the results were primarily driven by the data, rather than unduly influenced by the choice of credible limits for the informative prior distributions. On sensitivity analysis, substantial changes to the point estimates of the specificity of the fecal flotation protocol in detecting strongyle infections, and the sensitivity of fecal flotation in detecting Potorolepis spp. infections, only occurred when informative prior distribution limits that were considered highly implausible were used. Therefore, we do not consider these sensitivity analysis findings to invalidate the results presented herein.
The apparently accidental cause of death of most of the animals used in the study may have avoided substantial bias of results toward that of “sick” animals, therefore maintaining the applicability of the results to epidemiologic studies in the broader host populations.
Our findings suggest that both the modified agglutination test (at the titers used in this study) and the PCR protocol are highly specific in the detection of T. gondii infection in quenda and brushtail possums. Our study used chest fluid for the modified agglutination test, rather than serum, which may bias findings if extrapolating these results to the use of serum. However, findings of a study comparing T. gondii ELISA results in tissue fluid versus serum did not suggest ELISA specificity was substantially affected when using tissue fluid. 14 The finding of high modified agglutination test specificity in quenda and brushtail possums concurs with findings in other host species.7,13,20
The data were insufficient to assess the sensitivity of these tests in an adequately precise manner, as the apparent prevalence (using both tests) of T. gondii was low in both the quenda and possum study samples. The nonspecific bands amplified from some quenda tissues on T. gondii PCR were faint and not the correct product size for T. gondii. The 10 bands that were closest in size to T. gondii did not sequence as T. gondii, which supports our interpretation of such bands as negative results.
Our study found both immunofluorescence microscopy and the PCR and sequencing protocol to be highly specific in the detection of Giardia spp. infections in quenda, which concurs with high specificities observed in other host species.8,10,25 Immunofluorescence microscopy was highly sensitive in detecting Giardia spp. infection in quenda feces, and substantially more so than the PCR and sequencing protocol. This reflects other research findings, in which amplification by PCR was not successful in all Giardia-positive fecal samples tested.23,27 The identification of Giardia spp. cysts by fecal flotations in the majority of the immunofluorescence microscopy–positive, PCR and sequencing–negative fecal samples provided reassurance that the discrepancy between the number of samples positive by immunofluorescence microscopy versus those positive by the PCR and sequencing protocol was not because of lack of specificity by immunofluorescence microscopy. As Giardia spp. detection by fecal flotations has been shown to lack sensitivity in other species,10,32 the fact that Giardia spp. cysts were not identified in all fecal flotations is not unexpected. The sensitivity of both immunofluorescence microscopy and PCR and sequencing in detecting Giardia spp. infection in quenda may have been overestimated if there are commonly periods in which infected quenda do not shed cysts in their feces.
Given that only 2 fecal samples tested positive for Giardia spp. by PCR and sequencing at the gdh locus, and that these samples also tested positive for Giardia spp. at the 2 other loci tested, the gdh locus could be excluded from the PCR and sequencing protocol with no change in protocol sensitivity or specificity (further detail of the PCR and sequencing results at the individual loci are described previously). 16
The specificities of detection of various GI helminth infections by the fecal flotation protocol were generally high. Fecal flotation specificities for Labiobulura spp., Linstowinema spp., Physaloptera spp., and Potorolepis spp. may have been biased downward as we chose to be relatively conservative in inputting prior distributions, despite the reasonable expectation of a very high degree of specificity in the protocol for these helminths (distinctive helminth egg morphology, with all flotations performed by an experienced observer). However, we cannot exclude the possibility that these specificities reflect a small degree of misclassification. The relatively lower specificity for Trichuris spp. may have been attributable to eggs of other capillarids being mistaken for those of Trichuris spp. If this is the case, our observations on microscopy suggest that there is very little difference in the morphology or size range of eggs between Trichuris spp. and other capillarid genera in quenda. The published description of smooth-shelled eggs of a size range of 65 × 35 µm for T. peramelis 22 was not commonly reflected in the Trichuris spp. eggs frequently observed in the feces of quenda in which Trichuris spp. worms were obtained from their GI tract, and in the Trichuris spp. worms themselves. Eggs observed in this study did not have smooth shells and were typically 55–57.5 × 22.5–25 µm (only occasionally larger or smaller, AE Hillman, unpublished data).
The fecal flotation protocol detected the helminths Labiobulura spp., Linstowinema spp., Trichuris spp., and strongyles in quenda with a moderate-to-high degree of sensitivity, although results are limited by a lack of precision. The imperfect sensitivity of fecal flotation in detecting GI helminth infections may be influenced by factors such as prepatent infections (particularly given the relatively large number of subadult quenda in the study sample) and intermittent shedding of eggs. Although also limited by a lack of precision, the relatively low sensitivity of the fecal flotation protocol in detecting Potorolepis spp. reflects a lack of sensitivity in the detection of other intestinal cestodes via similar methods in other host species.19,21,26 Our point estimate of 36% sensitivity may have been an overestimate if any or all of the 3 unidentifiable cestode-positive quenda were infected with Potorolepis spp., which is considered highly likely. Physaloptera spp. and strongyloids were not sufficiently prevalent in our study to provide an adequately precise estimate of fecal flotation protocol sensitivity.
Compared with our findings, both the sensitivity and specificity of fecal flotations may be influenced through the use of different observers, or if the particular fecal flotation protocol varies from that used in our study. However, as quenda helminth eggs are relatively large and have distinct morphologic features, we believe that observers with prior experience undertaking fecal flotation microscopy (in any species) would achieve similar sensitivity and specificity of parasite detection using this described fecal flotation protocol.
Footnotes
Acknowledgements
We thank Louise Pallant, Russ Hobbs, Nandini Dendukuri, and Crystal Cooper; Kanyana Wildlife Rehabilitation Centre; associates of this project who collected and forwarded quenda and possum carcasses; the Department of Parks and Wildlife; Whiteman Park, Perth Airport, the Botanic Gardens and Parks Authority, John Forrest National Park and the Cities of Joondalup, Cockburn, Armadale, and Perth; WWF-Australia; participating property owners, managers, and staff; and volunteer field assistants.
Authors’ contributions
AE Hillman contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; and drafted the manuscript. AL Ash and AD Elliot contributed to design of the study; contributed to acquisition of data; and critically revised the manuscript. AR Kristancic contributed to design of the study; contributed to acquisition of data; drafted the manuscript; and critically revised the manuscript. AJ Lymbery and ID Robertson contributed to design of the study, and critically revised the manuscript. RCA Thompson contributed to conception and design of the study, and critically revised the manuscript. All authors gave final approval and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Toxo-Screen DA, bioMérieux, Mercy l’Etoile, France.
b.
Qiagen, Hilden, Germany.
c.
Fisher Biotec, Perth, Australia.
d.
Bio-Rad thermocycler, Bio-Rad Laboratories, Gladesville, New South Wales, Australia.
e.
SYBR safe DNA stain, Molecular Probes, Eugene, OR.
f.
Agencourt AMPure XP, Beckman Coulter, Brea, CA.
g.
Prism terminator cycle sequencing kit, Applied Biosystems, Foster City, CA.
h.
Applied Biosystems 3730 DNA analyzer, Thermo Fisher Scientific, Waltham, MA.
i.
Geneious 7.2, Biomatters, Auckland, New Zealand.
j.
Merifluor cryptosporidium/giardia, Meridian Bioscience, Cincinnati, OH.
k.
BX50 microscope, Olympus, Tokyo, Japan
l.
R version 3.1.1, R Foundation for Statistical Computing, Vienna, Austria.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by The Holsworth Wildlife Research Endowment–Equity Trustees Charitable Foundation, the Weston Fernie Research Fund, and the Australian Research Council (LP130101073). AE Hillman was supported by an Australian Postgraduate Award.
