Abstract
We compared 2 novel loop-mediated isothermal amplification (LAMP) assays that target either the 16S ribosomal RNA (rrs) gene or the gene encoding a 32-kDa leptospiral lipoprotein (lipL32) in order to assess the effect of the target on the accuracy of the LAMP assays. The most sensitive assay was the rrs assay with a limit of detection (LOD) of 1.2 × 101 genome equivalents per reaction. The novel lipL32 assay showed an LOD of 1.2 × 102 genome equivalents per reaction. Both assays showed adequate specificity when tested against a collection of bacteria commonly found in voided canine urine. However, when field samples were assayed, the rrs assays gave many false-positive results and a poor positive predictive value of 8.33%. In conclusion, even if the LAMP assay is used in low prevalence areas, the lipL32 assay would be preferable. Conversely, the higher analytical sensitivity of the rrs assay could be effectively used as a screening test in endemic areas with high disease prevalence, followed by confirmation of the positive results using the lipL32 assay.
Leptospires are excreted in the urine of infected animals, and the detection of leptospiral DNA is advantageous for diagnosing leptospirosis.1,2 Unfortunately, leptospires lose their integrity rapidly in urine, and their number declines considerably over time. 14 Prompt analysis is crucial for detection. The detection of leptospiral nucleic acid is accomplished using molecular techniques, such as real-time polymerase chain reaction (PCR).4,12,23,24 However, veterinary clinics with basic laboratory facilities lack adequate equipment and skills for performing real-time PCR assays. Hence, urine needs to be shipped to reference laboratories for analysis, thus delaying diagnosis. Another critical consideration is the shipment of potentially hazardous samples.
Loop-mediated isothermal amplification (LAMP)15,17–19 is a nucleic acid amplification technique that has some advantages over more traditional techniques, such as PCR. LAMP reactions are run at fixed temperatures of 60–65°C, without sophisticated and expensive instrumentation such as thermocyclers, using heating blocks or water baths given that LAMP tolerates great temperature deviation.5,19 Furthermore, LAMP reactions amplify DNA more efficiently than PCR, allowing rapid amplification of the target without initial denaturation.17,20 The LAMP assay is also inherently insensitive to the principal PCR inhibitors, and it is strikingly robust across 2 pH units (7.3–9.3), allowing simplified DNA purification. 5
Nonetheless, LAMP assays require thorough validation in order to ensure performance comparable to other molecular techniques. Only a few studies have attempted to validate LAMP assays for the detection of leptospiral DNA in humans and animals. Two of the methods reported rely on the amplification of the 16S ribosomal RNA (rrs) gene11,25 and of the gene encoding a surface-exposed 41-kDa lipoprotein of Leptospira (lipL41). 13 The lipL41 gene was used to detect leptospiral DNA in rodent kidneys. 13 The 2 studies targeting rrs in both animals and humans reported an analytical sensitivity of 10–100 targets per reaction.11,25 However, concerns regarding the specificity of targeting rrs genes in assays relying on leptospiral nucleic acid amplification have emerged.4,7,8,26–28,30 In particular, two 2015 studies4,8 have investigated the detection of leptospiral DNA in canine urine. The studies showed that hydrolysis probe–based real-time PCR could give false-positive results when field samples were comprehensively analyzed without using appropriate cutoff values. Similar findings were also noted when the rrs gene was amplified by LAMP. 25 The cross-reacting interfering off-targets have been attributed to common environmental bacteria. 4
The suitability of the 32-kDa leptospiral lipoprotein gene (lipL32) as a real-time PCR target for the detection of leptospires has emerged in recent years. Importantly, LipL32 is a subsurface protein found exclusively in pathogenic leptospires.9,16,22 We compared the performance of 2 LAMP assays, one targeting the lipL32 gene and the other the rrs gene, by means of reference material and samples of DNA purified from canine urine.
The validation approach was carried out according to the methods used in our 2015 study of real-time PCR assays for leptospira. 8 The analytical sensitivity used as the lower limit of detection (LOD) was experimentally estimated as the last serial dilution investigated yielding positives on all 5 replicates of the standard. This translates into >50% probability of testing positive with a 95% level of confidence. 3 The validation experiments were carried out using the genomic (g)DNA of Leptospira interrogans serovar Canicola strain Alarik. Nineteen other leptospiral serovars were used to assess the ability of the assays to detect other pathogenic strains, namely serovars Australis strain Ballico, Bratislava strain Riccio 2, Lora strain Riccio 37, Autumnalis strain Akiyami A, Castellonis strain Castellon 3, Bataviae strain Pavia 1, Grippotyphosa strain Moskva V, Hebdomadis strain Hebdomadis, Icterohaemorrhagiae strain Bianchi I, Copenhageni strain Wijnberg, Poi strain Poi, serovar Mini strain Sari, Pomona strain Mezzano I, Zanoni strain Zanoni, Hardjo strain Hardjoprajitno, Hardjo strain Farina, Saxkoebing strain Mus 24, Sejroe strain Topo 1, and Tarassovi strain Mitis Johnson. Five other reference bacterial strains representative of common contaminants or infectious agents were used to evaluate specificity, namely Pseudomonas aeruginosa ATCC 27853, Proteus vulgaris ATCC 13315, Streptococcus canis ATCC 43496, Escherichia coli ATCC 25922, and Staphylococcus pseudintermedius DSM 21284.
The DNA from a 200-µL aliquot of leptospiral culture was purified by means of a silica-based column method a as recommended by the manufacturer and quantified using a fluorescence-based nucleic acid quantification method. b The genome equivalents (GE) were calculated, and the gDNA samples were serially diluted 1:10 in a solution of molecular biology–grade water containing 10 ng/µL of canine gDNA and the exogenous internal control (EIC) at equimolar concentration with respect to the urine samples. All of the DNA extracts of the reference bacterial strains were produced and used in a previous study 8 with the exception of the S. pseudintermedius extract. The EIC, already evaluated in a previous study, 8 confirmed the absence of inhibitors in all samples purified from urine. Detailed descriptions have been reported elsewhere. 29
A total of 184 urine samples, obtained from dogs undergoing routine urinalysis at the Veterinary Clinical Pathology Service of the University of Bologna (Italy), and already used in another study, 8 were analyzed. Urine samples were included in the study regardless of the method of collection (free-catch, cystocentesis, or catheterization).
The oligonucleotides for the LAMP assays were designed with software c using a consensus sequence obtained by aligning the available Leptospira spp. lipL32 gene partial sequences obtained from GenBank. Those sets of primers, with some mismatches in the 3’ moiety of the oligonucleotides, were excluded. The oligonucleotides were purchased from a manufacturer, d and their sequences are reported in Table 1.
Primers used in loop-mediated isothermal amplification assays in the current study.
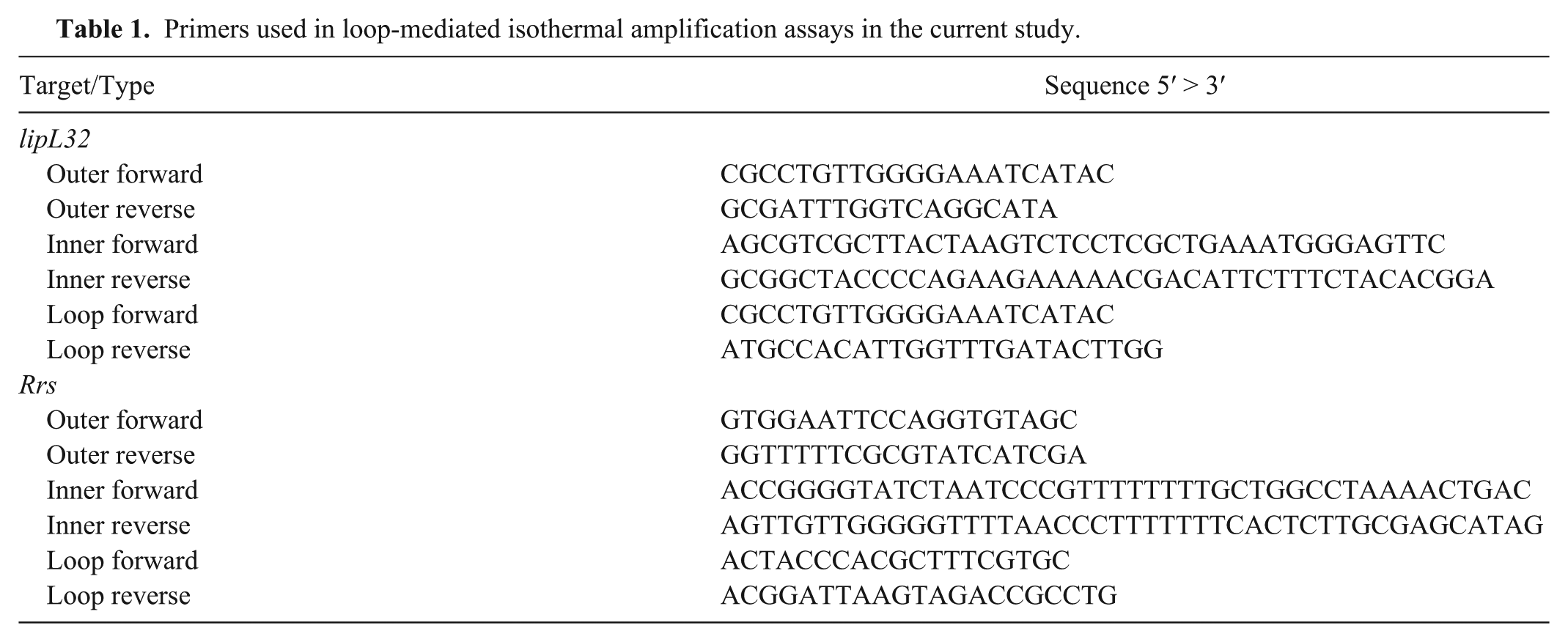

The lipL32 LAMP assays were carried out with a mixture of 1× LAMP buffer containing 2 mM MgSO4, e additional 3 mM MgSO4, 2 mM deoxynucleotide triphosphates, 1 M betaine, 1.6 mM of inner primers, 0.5 mM of loop primers, 0.2 mM of outer primers, 8 U of Bst DNA polymerase large fragment, e 1× of a commercial dye, f 2.5 μL of template, and molecular biology–grade water sufficient for a final volume of 25 μL. The tubes were incubated in a thermocycler g at a fixed temperature of 65°C for 1 h using a program of 60 cycles of 1 min with signal acquisition in the SYBR channel.
The lipL32 LAMP assay consistently amplified 1.2 × 102 Leptospira serovar Canicola GE per reaction for all 5 replicates (Fig. 1). Although the lipL32 LAMP assay is not intended to be quantitative, the assay showed a broad dynamic range with an almost linear range from 1.2 × 107 to 1.2 × 102 GE per reaction. The R2 coefficient was 0.977 (Fig. 2). The LOD experiments with the rrs LAMP assay gave, to some extent, even better results (Fig. 3), 1.2 × 101 copies per reaction were positive in all replicates.
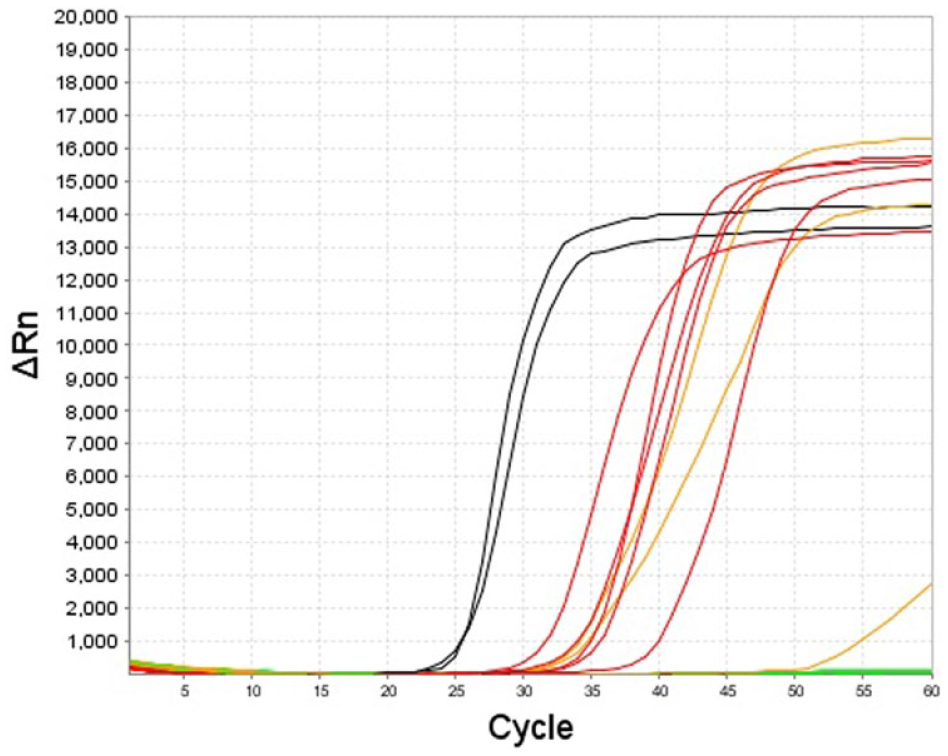
Analytical sensitivity experiments of the lipL32 loop-mediated isothermal amplification assay. The black lines represent the 2 replicates of the positive control containing 1.2 × 104 genome equivalents (GE) per reaction; the red lines represent the 5 replicates of 1.2 × 102 GE per reaction; and the orange lines represent the 1.2 × 101 GE per reaction.
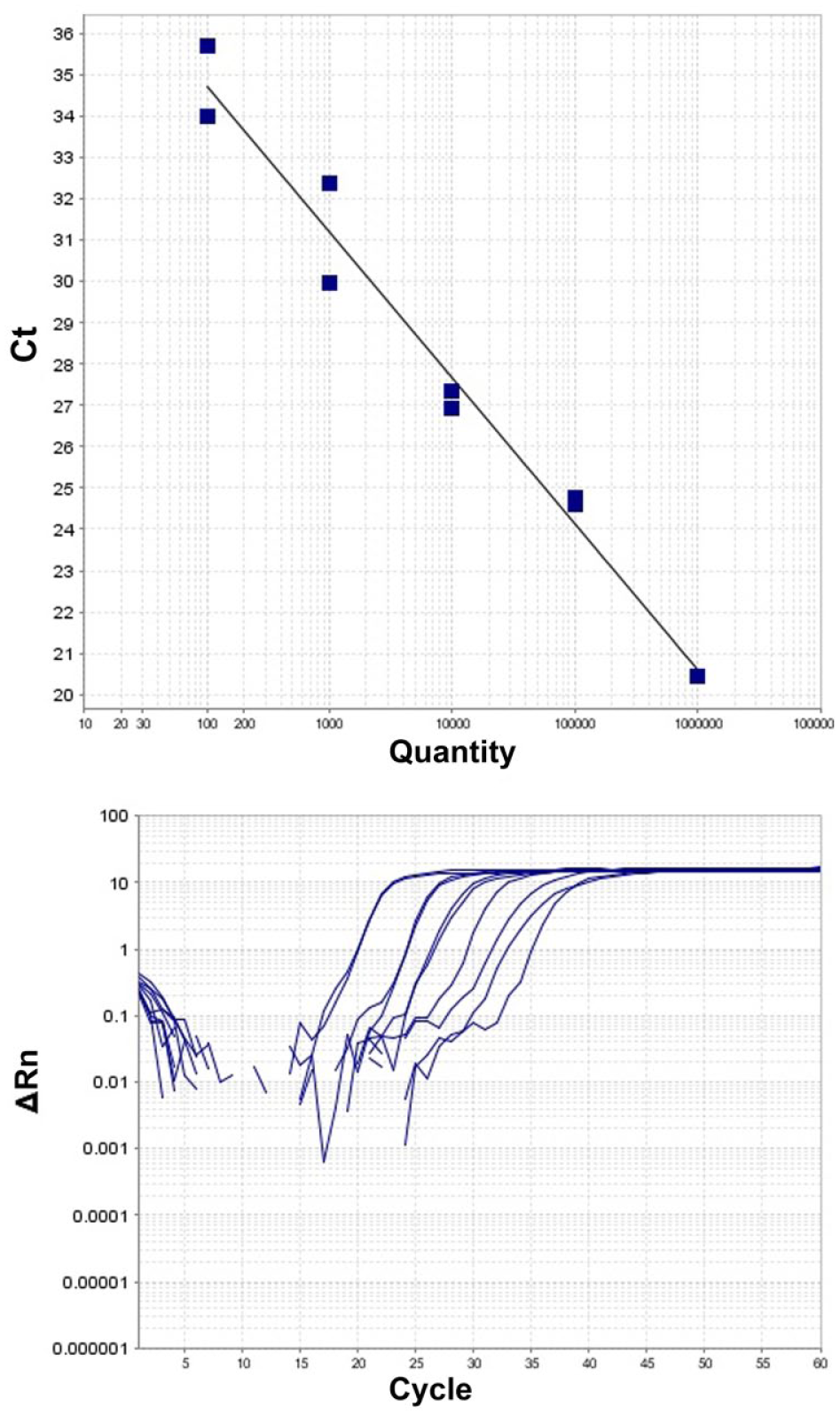
Linearity of the lipL32 loop-mediated isothermal amplification assay using genomic DNA of Leptospira interrogans serovar Canicola as a template. The plot of the 2 replicates for each standard dilution is shown.
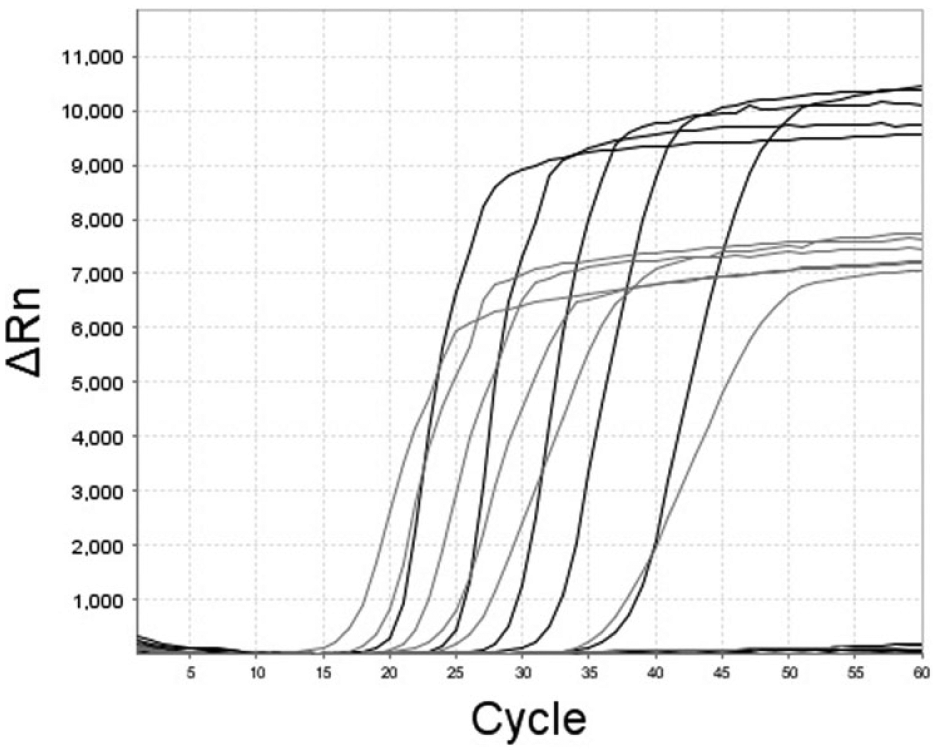
Comparison of the amplification curves of a loop-mediated isothermal amplification assay targeting lipL32 (black lines) and rrs (gray lines) using genomic DNA containing 1.2 × 106 to 1.2 × 101 genome equivalents of Leptospira interrogans serovar Canicola as templates.
All of the leptospiral strains tested gave positive signals with both LAMP assays. Neither the lipL32 nor the rrs LAMP assays gave positive results when the non-target reference bacterial strains were used as templates (all tested in replicate).
The 184 field samples had already been adequately purified and assayed with 2 hydrolysis probe real-time PCR assays in a previous study, 8 and only 1 sample had been found positive. The same sample was also found to be positive by both LAMP assays in the current study (observed prevalence 0.5%; 95% confidence interval (CI): 0.1–3.0%). However, the rrs assay had an additional 11 positive samples. These latter samples were also re-tested in a repeat of the rrs LAMP assay, and 10 of the 11 cases were positive in the re-run. To ascertain that those signals did not originate from leptospiral DNA, 2 cases were examined more in depth using target-specific restriction endonuclease digestion. For this purpose, the LAMP amplification products were incubated at 37°C for 60 min using HpaII restriction endonuclease, which was expected to cut the LAMP-specific rrs concatemers, and then electrophoresed in 1.5% agarose gel. However, no difference in the ladder pattern of the amplification products before and after the digestion could be detected. This indicated that HpaII did not find the recognition site specific for the leptospiral rrs concatemers. Although the false-positive findings were not significantly associated with the method of urine collection, 8 of 10 were attributed to free-catch urine, and they were most likely caused by commensal and/or environmental bacteria as has been suggested previously for other false-positive molecular results. 4
When the LODs reported in the current study were converted into leptospires/mL of urine assuming 100 µL of elution volume, the lipL32 LAMP assay showed an analytical sensitivity higher than that reported for an endpoint PCR and slightly less than that reported for a real-time PCR. 4 The rrs assay showed even higher LODs but conversely revealed a very high rate of false-positives and a very low positive predictive value (PPV) of 8.33% (95% CI: 0.21–38.5%). This may have been caused by the fact that the target was well-conserved among the many bacterial species, and spurious amplification may have occurred as a result of cross-reactivity with bacteria present in the lower urinary tract and contaminating only the free voided urine.
It should be noted that a higher analytical sensitivity was attained by the rrs LAMP assay as compared to the lipL32 assay, with LODs comparable to the most sensitive PCR techniques using hydrolysis probes. In consideration of the zoonotic nature of leptospirosis, it is also possible to consider the rrs LAMP assay as a valuable screening test in endemic areas with positive results needing to be confirmed by a more specific assay. Indeed, assuming a disease prevalence of ~13.5% (25 true positives out of the 184 samples), the same proportion of false-positives would have an acceptable 70% PPV. Based on the above considerations, the use of a LAMP screening assay using the rrs as a target and the confirmation of the positive sample by means of the lipL32 assay could be suggested as a suitable approach for identifying animals shedding leptospires in their urine.
The sensitivity of the LAMP assay to inhibitors possibly present in urine samples, which only underwent lysing procedures without complex DNA purification steps, was evaluated. The same volume of leptospiral gDNA diluted in molecular biology–grade water and in urine diluted 1:5 with molecular biology–grade water was added to a reaction mixture and amplified. The results confirmed that the addition of urine to the reaction mix had only a minimal impact on the LAMP assay (Fig. 4).
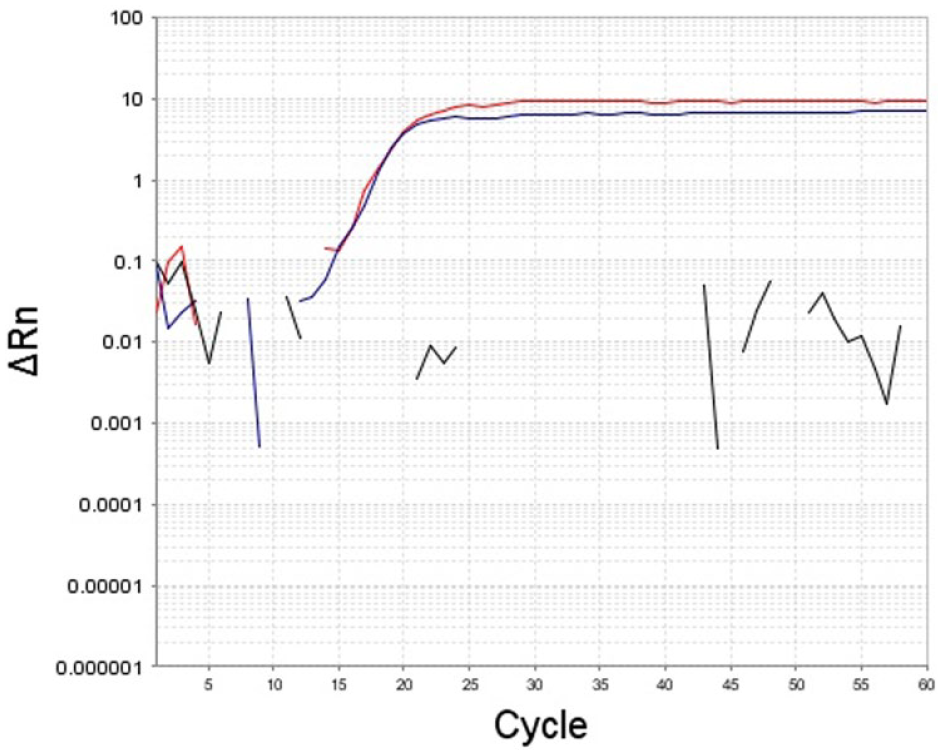
Effect of the addition of diluted urine in the loop-mediated isothermal amplification assay; in red, the amplification curve obtained using genomic (g)DNA diluted in molecular biology–grade water (MBGW) as a template and, in blue, the amplification curve obtained using the same amount of gDNA diluted in urine diluted 1:5 in MBGW as a template.
Even though the microscopic agglutination test (MAT) is considered the gold standard for diagnosing leptospirosis, 6 a combination of MAT and PCR is recommended for reaching a diagnosis with a higher level of confidence.10,21,26 The LAMP assay could be considered a suitable ancillary tool. MAT is a complicated test to carry out, control, and interpret; it requires the maintenance of living cultures of all the serovars, and is available in only a few laboratories. 14 Furthermore, MAT antibodies are not detectable in blood until ~7 d after the onset of clinical signs. 6 Likewise, the detection of DNA in blood, cerebrospinal fluid, and urine using PCR can be carried out in only a few reference laboratories because PCR is a technically demanding and costly technique. In contrast, the features of a LAMP assay could favor the use of this molecular technique in less specialized laboratories. However, it should be emphasized that, in many circumstances, the detection of Leptospira nucleic acids in canine urine can be problematic. Antibiotic treatment, frequently administered to the dog before the first visit to a clinic, rapidly reduces the presence of Leptospira in biological fluids to below the threshold of detection. This occurrence is a real concern in veterinary medicine with regard to achieving the correct diagnosis. 6
Footnotes
Acknowledgements
We thank Dr. Silvia Tagliabue for providing the Leptospira interrogans serovar Pomona strain Mezzano I and L. interrogans Canicola strain Alarik, and Dr. Francesca Brunello for providing all of the other leptospiral strains.
Authors’ contributions
F Gentilini contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and critically revised the manuscript. RG Zanoni contributed to analysis and interpretation of data; drafted the manuscript; and critically revised the manuscript. E Zambon contributed to analysis of data and drafted the manuscript. ME Turba contributed to conception and design of the study; contributed to interpretation of data; drafted the manuscript; and critically revised the manuscript. All authors gave final approval and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
NucleoSpin Tissue, Macherey-Nagel, Düren, Germany.
b.
QuantiFluor dsDNA System, Promega Italia, Milan, Italy.
c.
PrimerExplorer version 4, Eiken Chemical, Tokyo, Japan.
d.
Integrated DNA Technologies, Leuven, Belgium.
e.
New England Biolabs, Ipswich, MA.
f.
SYBR Safe, Life Technologies, Milan, Italy.
g.
StepOne, Life Technologies, Milan, Italy.
Declaration of conflicting interests
The author(s) declared no potential conflict of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
