Abstract
A single nucleotide polymorphism–based PCR assay has been developed to differentiate the attenuated vaccine strain used in Japan from field isolates of Erysipelothrix rhusiopathiae found in pigs. However, this assay has been evaluated with only Japanese strains and isolates; therefore, it is unknown whether it could be used in other countries with E. rhusiopathiae strains and isolates of different genetic backgrounds. In our study, the PCR assay was evaluated using Chinese E. rhusiopathiae vaccine strains and field isolates. The PCR assay was able to differentiate the attenuated vaccine strains from the field isolates of E. rhusiopathiae in China but with a pattern different from that observed in Japan (only a single nucleotide polymorphism was detected in the Chinese vaccine strains compared with 5 in the Japanese vaccine strains). Importantly, either a DNA polymerase without 3′ to 5′ exonuclease activity or an exo+ polymerase with an antibody inhibiting the proofreading activity was required. In conclusion, after evaluation and improvement, this fast differentiation assay can be extended from Japan to China.
Erysipelothrix rhusiopathiae is a facultative anaerobic, small, slender, gram-positive bacterium, 4 which is ubiquitous in nature and has been isolated from many species of wild and domestic mammals, birds, reptiles, amphibians, and fish. 10 E. rhusiopathiae causes disease in domestic animals (erysipelas) and humans (erysipeloid).2,10 Many countries have developed their own vaccines to control E. rhusiopathiae diseases.1,5,8 These vaccine strains originate from different strains and different attenuation pathways. Thus, these strains may have different genetic backgrounds, including differences in single nucleotide polymorphisms (SNPs).
Reliable differentiation assays to discriminate live attenuated vaccine strains from field isolates play a critical role in disease control programs.7,9 In 2015, an assay was developed to differentiate the attenuated vaccine strain used in Japan from field isolates of E. rhusiopathiae found in pigs. 6 However, this assay has been evaluated with only Japanese strains and isolates; therefore, it is unknown whether this assay could be extended to other countries with vaccine strains and field isolates of different genetic backgrounds. In our study, we used Chinese vaccine strains and field isolates to evaluate and improve the assay.
Four standard strains and/or isolates were used: 2 live attenuated vaccine strains (GC42 and G4T10) and 2 field isolates (C43-5 and SE38). The 2 vaccine strains, which were developed in China in the 1970s, are the only E. rhusiopathiae live attenuated vaccine strains used in China.1,11 The GC42 vaccine strain was generated via passage of field isolate C43-5 through cavies and chickens, and G4T10 was generated via passage of field isolate C43-5 through cavies and acriflavine treatment. 13 Field isolate C43-5 was collected in the 1950s, whereas isolate SE38 was collected in 2014.
Genomic DNA was obtained from the E. rhusiopathiae strains as previously described. 14 Polymerase chain reaction (PCR) amplification was performed in a 50-μL reaction mixture consisting of 50 ng of genomic DNA, 1 U of DNA polymerase a (the same as that used in the original report), 6 and 1.5 μL each of 20 μM SNP-specific primers 6 in a thermal cycler. b The PCR conditions were: initial denaturation at 95°C for 5 min and 3 steps of amplification (35 cycles) at 95°C for 30 s, 67°C for 30 s, and 72°C for 60 s. Next, the SNP sites of the standard strains and isolates were determined. Then, the PCR assay was evaluated under the same conditions, except that a DNA polymerase c with 3′ to 5′ exonuclease activity or a DNA polymerase d without 3′ to 5′ exonuclease activity was used. Finally, the PCR assay was evaluated under the same conditions, except that an exo+ DNA polymerase e with an antibody inhibiting the proofreading activity was used.
Our results demonstrated that the PCR assay can be used to differentiate Chinese live vaccine strains from field isolates (Fig. 1). We observed 1 specific band for each of the Chinese live attenuated vaccine strains, whereas there was no specific band in the differentiation assay for either of the field isolates. Next, we sequenced the SNP sites of the 4 standard strains and/or isolates. The sequencing results (GenBank accession KT829597–KT829616; Table 1) further proved the validity of the PCR results. Only the 2 live attenuated vaccine strains had SNPs in the ERH_1398 locus (compared with Fujisawa strain), 6 the locus in which the differentiation assay amplified the specific bands. The differentiation assay results for the Chinese field isolates were identical to those for the Japanese field isolates—no band was produced. 6 However, each Chinese live attenuated vaccine strain had only 1 specific band, whereas the Japanese vaccine strain had 5 specific bands. 6 This difference reflects the different genetic backgrounds of the strains. Therefore, in countries with vaccine strains that differ from those used in Japan or China, an evaluation of this assay with relevant attenuated live vaccine strains and field isolates is required prior to any field application.
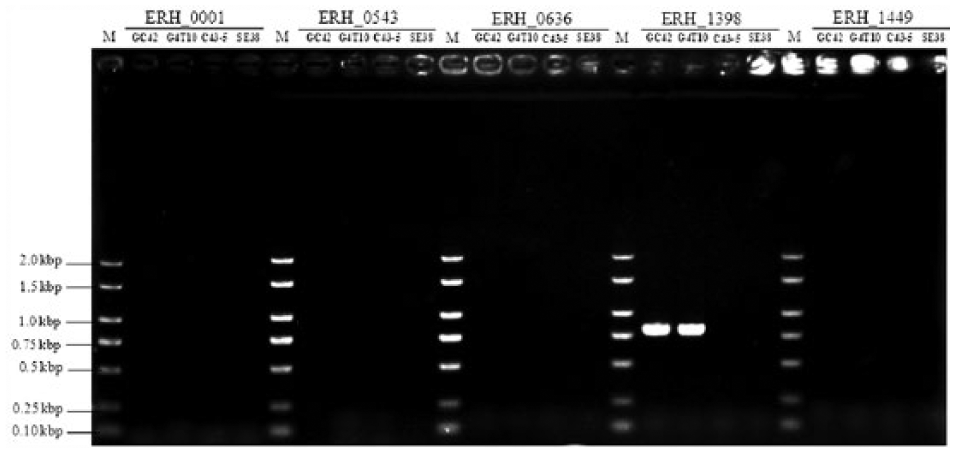
Differentiation assay results for the 4 standard Chinese Erysipelothrix rhusiopathiae strains and field isolates. ERH_0001, ERH_0543, ERH_0636, ERH_1398, and ERH_1449, gene locus tags on the Fujisawa strain genome 6 ; GC42, an attenuated live vaccine strain used in China; G4T10, another attenuated live vaccine strain used in China; C43-5, a field isolate obtained in the 1950s; SE38, a field isolate obtained in 2014. M = molecular size marker.
Single nucleotide polymorphisms (SNPs) detected for the 4 standard Chinese Erysipelothrix rhusiopathiae strains compared with the Fujisawa strain. 6
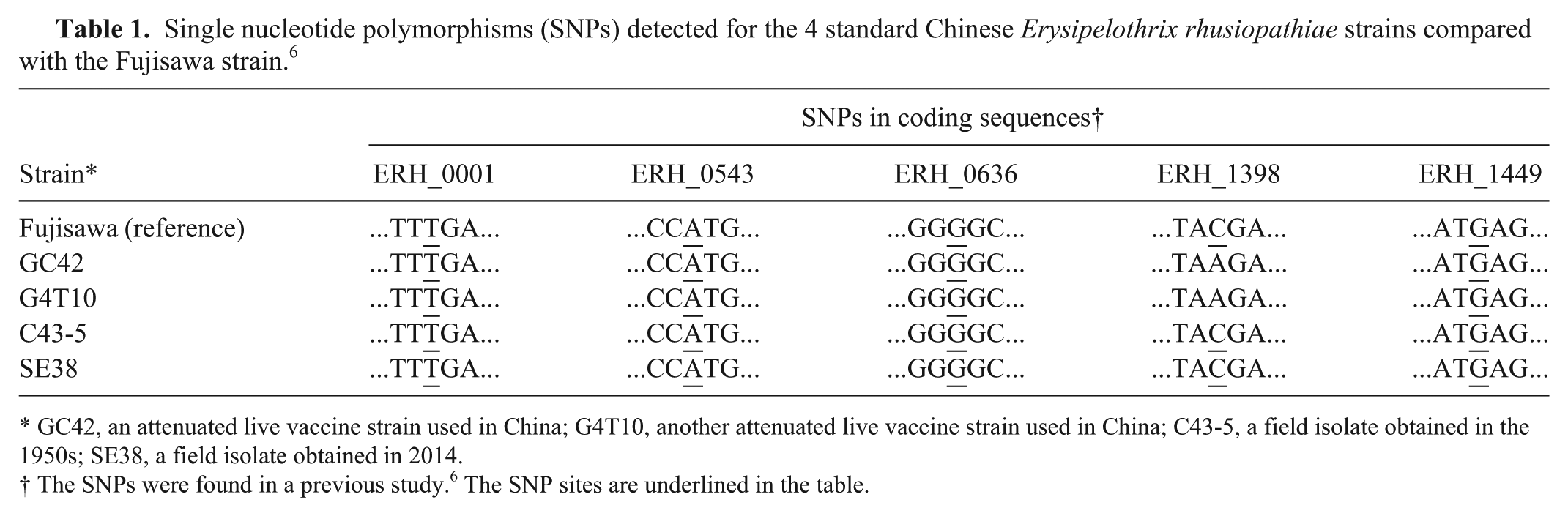
GC42, an attenuated live vaccine strain used in China; G4T10, another attenuated live vaccine strain used in China; C43-5, a field isolate obtained in the 1950s; SE38, a field isolate obtained in 2014.
The SNPs were found in a previous study. 6 The SNP sites are underlined in the table.
In the original report describing the assay, the authors speculated that PCR enzymes with excellent 3′ to 5′ exonuclease activity are a critical determinant of differentiation. 6 We compared the PCR using a DNA polymerase c with excellent 3′ to 5′ exonuclease activity with the PCR using a DNA polymerase d without 3′ to 5′ exonuclease activity. The results indicated a gain of specific bands in the ERH_0543, ERH_0636, and ERH_1398 loci from all strains in the assays using the DNA polymerase c with excellent 3′ to 5′ exonuclease activity (Fig. 2A). In contrast, the assay using the DNA polymerase d without 3′ to 5′ exonuclease activity yielded identical results (Fig. 2B) to those in the assay using DNA polymerase a (Fig. 1). The 3′-terminal mismatched nucleotides of the primers were removed through the proofreading function in the PCR assay using an exo+ polymerase 12 ; therefore, we speculate that the PCR amplifications with the exo+ polymerase c are promoted in the ERH_0543, ERH_0636, and ERH_1398 loci. Thus, PCR enzymes without 3′ to 5′ exonuclease activity are recommended for the differentiation assay.
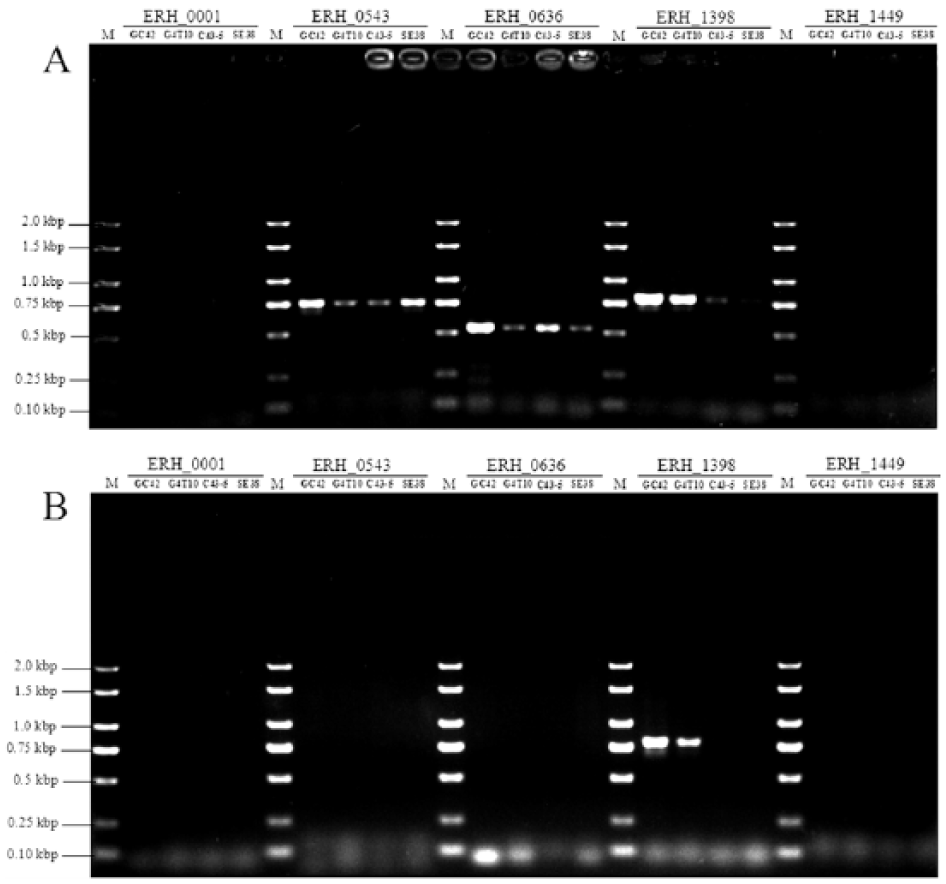
Differentiation assay results of the PCR using DNA polymerases with or without 3′ to 5′ exonuclease activity.
The use of monoclonal antibodies against Taq DNA polymerase in a hot-start PCR is an important method of improving PCR specificity. 3 Therefore, the performance of a PCR using an exo+ polymerase e with an antibody that inhibits proofreading activity was also evaluated. We observed results (Fig. 3) that were identical to those in the assay using the DNA polymerase a (Fig. 1). The DNA polymerase in the original report 6 was an exo+ polymerase with an antibody that inhibits proofreading activity. a From our results, we suggest that the critical determinant of successful differentiation is not the 3′ to 5′ exonuclease activity but, instead, is the antibody that inhibits proofreading activity. Therefore, if an exo+ polymerase is used in the differentiation assay, it is recommended that the enzyme include an antibody that inhibits the enzyme’s proofreading activity.
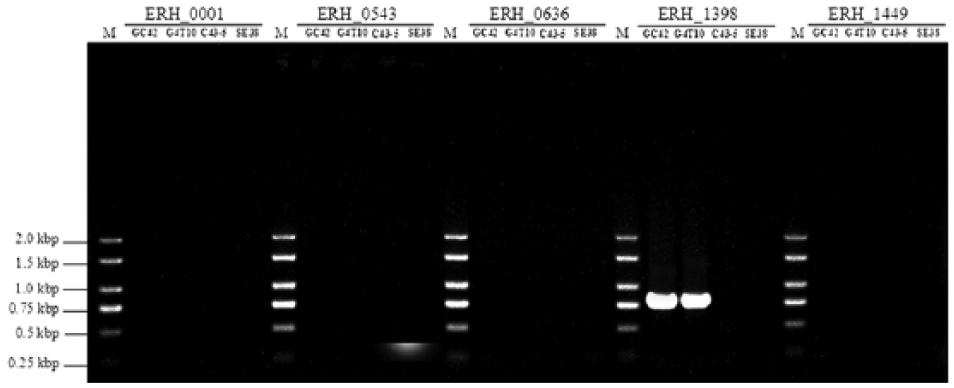
Differentiation assay results of the PCR using an exo+ polymerase e with an antibody inhibiting the proofreading activity. ERH_0001, ERH_0543, ERH_0636, ERH_1398, and ERH_1449, gene locus tags on the Fujisawa strain genome 6 ; GC42, an attenuated live vaccine strain used in China; G4T10, another attenuated live vaccine strain used in China; C43-5, a field isolate obtained in the 1950s; SE38, a field isolate obtained in 2014. M = molecular size marker.
Footnotes
Acknowledgements
We thank Chao Wu (Cooperative Innovation Center for Sustainable Pig Production) for technical assistance.
Author’s contributions
W Zhu contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and critically revised the manuscript. J Li contributed to conception and design of the study, and contributed to acquisition, analysis, and interpretation of data. Y Wang contributed to conception and design of the study, and critically revised the manuscript. C Kang contributed to acquisition, analysis, and interpretation of data. M Jin critically revised the manuscript, and gave final approval. C Kang, M Jin, and H Chen agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
KOD FX DNA polymerase, TOYOBO Co. Ltd., Osaka, Japan.
b.
ETC811 thermal cycler, Eastwin Scientific Equipments Inc., Beijing, China.
c.
LA Taq polymerase, Takara Bio Inc., Shiga, Japan.
d.
rTaq DNA polymerase, TOYOBO Co. Ltd., Osaka, Japan.
e.
KOD Plus DNA polymerase, TOYOBO Co. Ltd., Osaka, Japan.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by funding from the National Natural Science Fund of China (31472220).
