Abstract
Equine recurrent uveitis (ERU) is the most frequent cause of blindness in horses worldwide. Leptospira has been implicated as an etiologic agent in some cases of ERU and has been detected in fresh ocular tissues of affected horses. The objective of this study was to determine the presence of Leptospira antigen and DNA in fixed equine ocular tissues affected with end-stage ERU. Sections of eyes from 30 horses were obtained. Controls included 1) 10 normal equine eyes and 2) 10 equine eyes with a nonrecurrent form of uveitis. The experimental group consisted of 10 eyes diagnosed with ERU based on clinical signs and histologic lesions. Sections were subjected to immunohistochemical staining with an array of rabbit anti-Leptospira polyclonal antibodies. DNA extractions were performed by using a commercial kit designed for fixed tissue. Real-time PCR analysis was completed on extracted DNA. The target sequence for PCR was designed from alignments of available Leptospira 16S rDNA partial sequences obtained from GenBank. Two of 10 test samples were positive for Leptospira antigen by immunohistochemical assay. Zero of 20 controls were positive for Leptospira antigen. All test samples and controls were negative for Leptospira DNA by real-time PCR analysis. Leptospira was detected at a lower frequency than that previously reported for fresh ERU-affected aqueous humor and vitreous samples. Leptospira is not frequently detectable in fixed ocular tissues of horses affected with ERU when using traditional immunohistochemical and real-time PCR techniques.
Keywords
Equine recurrent uveitis (ERU) is the most frequent cause of blindness in horses worldwide, and economic losses associated with ERU are significant. 7 The disease is initiated by a single uveitis episode, which is followed by waxing and waning episodes of uveitis. It is the chronic and recurrent nature of the disease that differentiates ERU from the primary ocular and systemic forms of equine uveitis. Leptospira interrogans has been implicated as an etiologic agent in some cases of ERU. Horses experimentally or naturally infected with L. interrogans were shown to develop clinical uveitis 1 to 2 years after infection. 1,14 Numerous studies directly connected L. interrogans and ERU by identifying the bacteria in fresh aqueous and vitreous humor samples of affected horses. A recent study performed in Western Europe revealed 52.8% of equine vitreous humor samples with ERU were positive via culture for Leptospira and 71% were positive by PCR. 15 Although extensive studies were performed in Western Europe, only 1 study that confirmed the presence of Leptospira organisms in ERU affected equine ocular tissue was performed in the United States. In that California study, Leptospira DNA was detected by PCR in fresh samples of aqueous humor from 70% of horses with clinical ERU and in 6% of normal control horses. 5 Horses with ERU also have increased antibodies to Leptospira in vitreous humor compared with serum antibodies; this suggests that there is a persistent ocular infection with Leptospira. 16 Cross-reactivity among antigens of equine cornea, lens, and leptospires has been documented by using immunoblotting techniques. 12 Research results suggest that local immune mechanisms also contribute to the pathogenesis of equine recurrent uveitis. 9 The findings of predominantly CD4+ lymphocytes, increased interleukin (IL) 2 and interferon-γ, and decreased IL-4 are suggestive of a cell mediated (Th1) immune response. 6 The autoimmune nature of ERU was demonstrated experimentally by the induction of recurrent uveitis in horses after injection of interphotoreceptor retinoid-binding protein and S-angiten. 3,4 The purpose of this study was to determine the frequency of detection of L. interrogans DNA and antigen in fixed equine ocular tissues affected with end-stage ERU by using real-time PCR and immunohistochemistry. Our hypothesis was that Leptospira DNA and antigen would be detectable in fixed archival equine ocular tissue affected with ERU.
Samples were fixed in 10% neutral buffered formaldehyde, dehydrated in ascending concentrations of ethanol and xylene, and routinely embedded in paraffin. Sections of 5-micron thickness were prepared from tissue blocks. Thirty globes were obtained from 30 separate horses. Controls included 1) 10 normal equine eyes and 2) 10 equine eyes with a nonrecurrent form of uveitis. The experimental group consisted of 10 eyes with ERU. A positive control sample of L. interrogans-infected porcine kidney was also included. Ocular histopathologic evaluation to confirm the presence or absence of ERU in experimental and control groups was performed by a board-certified veterinary pathologist (R.R.D.). DNA extraction was performed on paraffin-embedded, formalin-fixed equine ocular sections. The sample of Leptospira-infected porcine kidney was also extracted and served as a positive control for DNA extraction. Two individual 5-micron sections from each tissue specimen were deparaffinized in xylene, rehydrated in descending concentrations of ethanol, rinsed with 100% ethanol, centrifuged at 12,000x g, and air-dried. Tissue was digested overnight with proteinase K a at 56°C. Extraction of DNA was performed by using the MagneSil genomic fixed tissue system a according to the manufacturer's instructions. Samples of DNA with a final volume of 25 μL were stored at −20°C until analyzed. To ensure an adequate total DNA concentration in the samples for further experiments, a DNA quantification was performed on all equine ocular samples. The primers b F (forward) 5′-GGA GCA GAC ATC AAA TAA GTA GG-3′ and R (reverse) 5′-CCA TAA CCA TCA CCA TGA TAG G-3′ were used to target the ubiquitous equine membrane associated transporter gene. 10 A product size of 100 bp was targeted and SYBR green c was used to monitor and quantify double-stranded DNA levels. Real-time quantitative PCR was performed on a Stratagene Mx4000 d by using SYBR green reagents. c The PCR reaction contained 5 μl of template in a final volume of 20 μl. Each primer was used at a final concentration of 200 nM. The amplification protocol consisted of 15 min at 95°C followed by 40 cycles of amplification (denaturation at 94°C for 15 sec, annealing at 60°C for 1 min, and extension at 76°C for 30 sec), after which the reaction was stopped (95°C for 2 min) and melted (55–96°C), with plate readings every 1°C. All samples were subjected to real-time PCR for Leptospira. The primers b and probes b for this PCR were previously described 13 and were obtained from alignments of available Leptospira 16S rDNA partial sequences obtained from the GenBank nucleotide sequence database. The PCR primers F 5′-CCC GCG TCC GAT TAG-3′ and R 5′-TCC ATT GTG GCC GRA GAC AC-3′ were located between the positions 191 and 278 of the Leptospira 16S ribosomal subunit gene (GenBank accession no. DQ991474), with an expected product size of 87 base pairs (bp). The probe, 5′- (FAM)CTC ACC AAG GCG ACG ATC GGT AGC-3′(3IABlkFQ), had the fluorescent reporter dye, 6-carboxy-fluorescein (FAM) located at its 5′ end and the quencher Iowa Black FQ located at the 3′ end and corresponds to positions 225–248 of GenBank accession no. DQ991474. The PCR was performed on a Stratagene Mx4000 d and was carried out by using 5 μL of sample DNA, extraction control (porcine kidney), or known positive and negative DNA controls added to the 45 μL mastermix, e providing final concentrations of 200 nM of each primer and 200 nM of the FAM-IABlkFQ labeled probe, as previously described. 13 Immunohistochemistry for L. interrogans was also performed on all samples. Paraffin-embedded, formalin-fixed equine ocular sections and the section of Leptospira-infected porcine kidney were positioned on slides, hydrated, and placed in Tris buffer for at least 5 min before staining. Slides were treated with 3% H2O2 for 15 min, washed, treated with proteinase K for 5 min, washed, subjected to Protein Block f for 5 min, and drained. Deparaffinized sections were incubated in rabbit anti-Leptospira cocktail antibody g (a mixture of antibodies against L. interrogans serovars pomona, hardjo, icterohaemorrhagiae, canicola, grippotyphosa, and bratislava) at a 1:1,600 concentration for 30 min. Negative controls were treated with rabbit immunoglobulin G antibody h at a 1:1,000 dilution for 30 min. Subsequently, sections were incubated with a biotinylated secondary antibody i for 20 min and a streptavidin-biotin complex reagent i for 20 min, both at room temperature, followed by a Tris buffer rinse. Development proceeded for 5 min with NOVA Red. j NOVA Red chromogen is readily discernible from black-brown melanin pigment present in equine globes. Slides were counter-stained in Mayer hematoxylin and azure blue, then dehydrated, and cover-slipped. Additional controls were also included via omission of the primary antibody or both primary and secondary antibodies. The staining intensity and distribution of Leptospira antigen was evaluated by a masked board-certified veterinary pathologist (J.R.T.). Immunostaining from an average of 3 high-power (200x) microscopic fields was assessed by using the following semiquantitative grading scale: grade 0, none or occasional positive cells only; grade 1, sparse positive cells; grade 2, moderate positive cells; and grade 3, intense positive-staining cells.
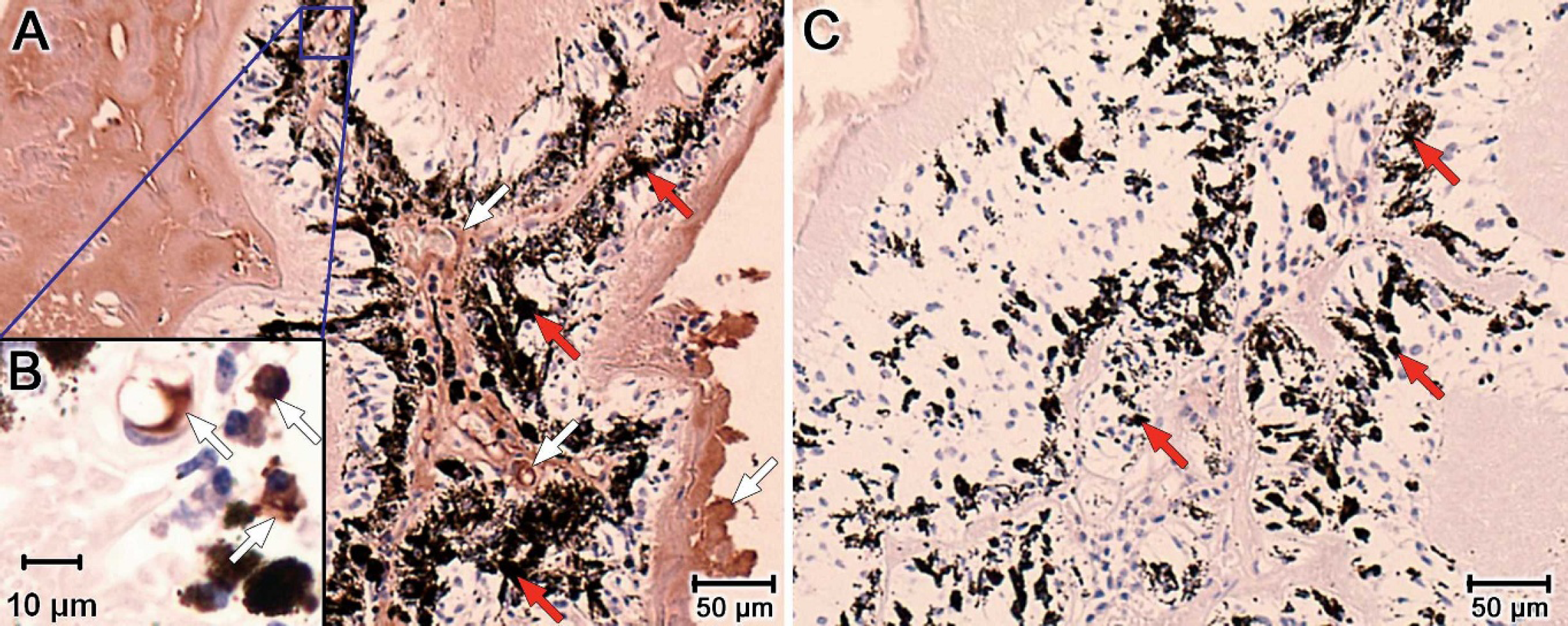
The presence of amplifiable DNA was confirmed on all equine ocular tissue samples after extraction by performing PCR for an equine genomic target. Amplifiable DNA was identified in 29 of 30 samples after initial extraction with 1 normal control sample being negative. This sample was extracted a second time, and DNA PCR was repeated, confirming that amplifiable DNA was present in the second extraction and in the final extraction of 30 of 30 samples. Real-time PCR evaluation for Leptospira DNA was negative for all equine ocular samples and for the negative DNA control. The extraction control (porcine kidney) and positive DNA control were positive, showing amplification of the 87-bp target Leptospira product. Immunohistochemistry indicated that 2 of 10 eyes from the ERU affected experimental group and the section of Leptospira-infected porcine kidney exhibited positive immunoreactivity. Positive results were repeatable on multiple sections. Of the positive horses in the ERU group, grade 2 focal immunoreactivity was noted in the corneal epithelium of 1 horse and in the uveal tract (inflammatory cells within the ciliary body) of another horse. A representative photomicrograph is shown in Figures 1A and 1B. The control sample of porcine kidney exhibited multifocal areas of mixed grade 2 and grade 3 immunoreactivity. Positive samples were confirmed by repeating immunohistochemistry on a minimum of 3 sections from the same sample. The 8 additional samples from the experimental (ERU) group were negative, as were all samples from control groups and the negative controls (Fig. 1C). Statistical evaluation with the Fisher exact test revealed no significant difference (P = 0.1) in the detection of Leptospira antigen between the experimental (ERU) group and the control groups.
The hypothesis that Leptospira DNA would be detectable in fixed archival equine ocular tissue affected with ERU was rejected. The hypothesis that Leptospira antigen would be detectable in fixed archival equine ocular tissue affected with ERU was accepted. However, Leptospira antigen had a low prevalence in ocular tissues from horses affected with ERU. Detection of Leptospira components (DNA, antigen) was lower in this study compared with previously reported evaluations of fresh ocular tissue from horses with ERU. There are several explanations for decreased leptospiral DNA and antigen detection in the fixed equine ocular samples evaluated in this study. Although amplifiable DNA was confirmed in all equine samples by SYBR green PCR and the presence of Leptospira DNA was detected in the extraction control (fixed porcine kidney), target DNA fragmentation may have occurred as a process of fixation and resulted in negative results by real-time PCR for Leptospira. Fixation may have damaged antigenic portions of Leptospira bacteria present in sections of equine ocular tissue. Although the control section of porcine kidney was processed in an identical manner and revealed strong positive staining for Leptospira sp. antigen, uncontrolled sample variation of formalin-fixation time and time spent in paraffin blocks also may have affected our results. The porcine kidney might have had a higher number of Leptospira organisms present than what was anticipated in the equine ocular tissue, possibly overestimating the value of the PCR assay. In previous studies, Leptospira DNA and antibodies toward Leptospira have been isolated from aqueous and vitreous humor samples. Although a small amount of aqueous and vitreous humor is preserved by fixation, Leptospira organisms could have been lost during the fixation process if they were present in small numbers and isolated solely in the aqueous/vitreous compartments. All eyes included in this study were in the chronic or end-stage categories of ERU, whereas the majority of ocular tissue used in previous studies 2,5,8,11,15 that detected Leptospira sp. was from horses affected with active, earlier stages of ERU. The immune-mediated etiology of ERU is well documented and widely accepted. Recent publications demonstrated immunogenic potential of small components of Leptospira sp., which suggested that the bacteria may play a role in the initiation of some cases ERU. In the ERU-affected equine tissue samples used in this study, Leptospira may have been present initially and subsequently cleared by the immune system, with no organisms present in ocular tissue with end-stage disease. Alternatively, Leptospira may have never been present in ERU-affected eyes evaluated in this study. The detection of leptospiral antigen in 2 ERU-affected eyes is a trend unique to this study but was not statistically significant when compared with control groups. This finding may be because of true bacterial presence or could be a false-positive result from antigenic similarity between equine ocular tissues and Leptospira bacteria, which was previously documented with immunoblotting techniques. In these studies, a protein epitope involved in an antigenic relationship with equine cornea and lens was located internal to the surface of Leptospira bacteria. 12 One positive case in this study revealed Leptospira immuno-reactivity of the cornea, which was consistent anatomically with the previous literature, although the other positive case showed increased staining of inflammatory cells within the ciliary body of the uveal tract. A discrepancy noted in this study was that Leptospira antigen was detected in 2 samples but Leptospira DNA was not detected by PCR (a more sensitive testing method). A possible explanation for this finding would be that different sections of the same eye were used for immuno-histochemistry and PCR with bacteria only being present in certain sections. Alternatively, antigenic similarity between equine ocular tissues and Leptospira bacteria could have caused false-positive reactivity with Leptospira antibodies. 12 Scientific impact of this study is multifactorial. This study introduces 2 techniques that could be tools for diagnostic pathologists or laboratory diagnosticians who might want to determine leptospiral involvement in cases of ERU when only fixed tissue is available. It also provides insight into the role of Leptospira in chronic, end-stage ERU and represents a new demographic of ERU horses (because 90% of ERU globes in this study were from midwestern and eastern American horses). Currently, there are no epidemiologic studies that linked Leptospira and ERU in the midwestern or eastern United States. To determine if geographic variation of Leptospira involvement in ERU is present, additional studies are needed to compare Leptospira incidence in fresh and fixed ocular tissues from horses in different geographic localities.
Acknowledgements. The authors thank Sunny J. Younger for her assistance with the PCR evaluations and Marilyn E. Beissenherz for her assistance with the immunohistochemical analysis.
Footnotes
a.
Promega Corporation, Madison, WI.
b.
Integrated DNA Technologies Inc., Coralville, IA.
c.
Qiagen QuantiTect SYBR Green PCR Kit, Valencia, CA.
d.
Stratagene Inc., La Jolla, CA.
e.
TaqMan Universal PCR Mastermix Mix, Applied Biosystems, Foster City, CA.
f.
DakoCytomation, Carpenteria, CA.
g.
National Veterinary Services Laboratories, Aimes, IA.
h.
Sigma, St. Louis, MO.
i.
LSAB2 System-HRP, DakoCytomation, Carpenteria, CA.
j.
Vector Laboratories, Burlingame, CA.
