Abstract
The objective of this study was to investigate whether intramuscular vaccination of healthy adult horses with a killed or a modified live equine herpesvirus type 1 (EHV-1) vaccine could induce transient positive PCR results in either blood or secretions collected on a nasopharyngeal swab. Four horses in each group received either a single killed or a modified-live vaccine intramuscularly. Two local commingled and 2 distant nonvaccinated controls were included for each group. All horses were observed daily for evidence of clinical abnormalities throughout the study periods. Blood and nasopharyngeal swabs were collected twice before vaccination and once weekly for 4 weeks after vaccination and submitted for PCR testing for EHV-1 by 2 independent laboratories using different real-time PCR methodologies. Serum samples collected from all horses on the vaccination day and 21 days later were tested for antibodies against EHV-1 using a serum neutralization test. Whereas the 2 vaccine strains tested positive in both EHV-1 PCR assays, nasopharyngeal swabs and whole blood collected from vaccinated and control horses had negative PCR test results for EHV-1 during the entire study period. Serum neutralization testing revealed a 2- to 4-fold increase in titers for all vaccinated horses, whereas titers in control horses were largely unchanged. The use of seropositive horses before immunization and the sampling frequency of 7 days may have prevented the occasional molecular detection of the vaccine virus in whole blood and nasopharyngeal secretions. However, the study results demonstrate that detection of EHV-1 DNA by PCR in vaccinated and unvaccinated healthy horses is not a common event.
Equine herpesvirus-1 (EHV-1) infection is widespread in horse populations throughout the world and produces well documented syndromes of respiratory disease, abortion, neonatal foal death, and myeloencephalopathy. 22 Although true seroprevalence against EHV-1 is difficult to determine because of widespread use of EHV vaccines in horses, recent evidence supports that exposure to EHV-1 is common during the first year of life, implicating the dam as the primary source for EHV-1 exposure in suckling foals. 4 , 5 As with other α-herpesviruses, primary exposure is often followed by life-long latent infection in recovered horses. Periodic reactivation of latent infections can lead to viral shedding and horizontal transmission. 2 Thus, early recognition of disease as well as viral shedding is crucial for implementation of management practices that decrease the risk of exposure of susceptible horses. Traditionally, virus isolation from a nasopharyngeal swab or blood has been the gold standard for diagnosis of viral shedding or viremia, respectively. Recently, PCR assays, which are more sensitive and rapid, have largely replaced the time-consuming procedure of virus isolation, as exemplified in recent outbreaks of EHV-1 myeloencephalopathy at riding schools, racetracks, and veterinary hospitals throughout North America. 6 , 8 , 21 Unfortunately, PCR assays only detect viral genomic DNA and are, therefore, unable to distinguish between lytic, dead, or latent virus. Further, vaccination with killed or modified-live EHV-1 vaccines may potentially confound interpretation of molecular diagnosis by producing false positive results (i.e., detection of vaccine virus or vaccine-associated reactivation of latent virus). This scenario was recently experienced by 2 of the authors (K.P.C. and H.C.S.) when nasopharyngeal swabs collected from clinically healthy horses 1–3 weeks after vaccination with a killed EHV-1 vaccine yielded positive PCR assay results. The possibility of having detected a field strain (primary exposure or reactivation of a latent stage) was also considered. This finding prompted the current study to investigate whether intramuscular vaccination of horses with either killed or modified-live EHV-1 vaccines would result in transient positive PCR results in nasopharyngeal secretions, whole blood, or both.
The study involved 14 adult horses, 8–17 years of age (mean 11.2 years) from Michigan State University research and teaching herds. There were 7 mares and 7 geldings of various breeds (6 Arabians, 4 Standardbreds, 2 Thoroughbreds, 1 Quarter Horse, and 1 Trahkener). The horses were randomly divided into 2 groups of 6 horses (4 vaccinated horses and 2 local, commingled controls). All horses had received bivalent EHV-1/4 vaccines in the past but none of the horses had received a booster within 6 months before the study period. Group 1 horses were studied in the fall (August to October 2005) and group 2 horses were studied in the winter (January to March 2006). Four horses in each group received either a single killed a (group 1) or a modified-live b (group 2) vaccine intramuscularly. Two additional nonvaccinated horses were used as distant controls for each group and were housed more than 200 m from the experimental group. These same distant control horses were used during the fall (group 1) and winter (group 2) studies. All horses were observed daily for evidence of clinical abnormalities (lethargy, decreased feed intake, elevated respiratory rate, nasal discharge, and coughing) throughout the study periods. Nasopharyngeal swabs c and whole blood (ethylenediaminetetraacetic acid [EDTA]-anticoagulated) samples for PCR analysis were collected in duplicate 7 days before vaccination, on the day of vaccination, and weekly thereafter for 4 weeks. Serum for serologic testing was also collected on the day of vaccination and 21 days later from all horses. All procedures performed on the horses were reviewed and approved by the Institutional Animal Care and Use Committee of the Michigan State University.
Nasopharyngeal swabs placed in viral transport medium (minimal essential medium with 0.125% gentamycin and 0.1% amphotericin B) and whole blood samples were processed and analyzed for the presence of EHV-1 DNA at 2 different laboratories (Diagnostic Center for Population and Animal Health, Michigan State University, Lansing, Michigan (MSU) and the Lucy Whittier Molecular and Diagnostic Core Facility, Department of Medicine and Epidemiology, School of Veterinary Medicine, University of California, Davis, California (UCD). After collection, samples were placed in a refrigerator (MSU) or sent via overnight mail on ice packs (UCD). Both laboratories used a commercial DNA kit d to extract DNA from nasopharyngeal swabs and whole blood. Each swab was vortexed for 10 seconds, inverted, and spun down at 16,000 × g for 5 minutes to retrieve a cell pellet. After removing the swab and supernatant, each pellet was resuspended into 400 μl of phosphate-buffered saline solution, and the sample was divided into 2 equal aliquots, 1 for extraction and 1 as a backup sample. A total of 180 μl of whole blood was extracted from each blood sample. After extraction, the 2 sample types (nasal secretions and blood) were eluted in 100 μl water. e Samples were tested for the presence of EHV-1 by a SYBR Green-based real-time PCR assay developed and implemented for routine diagnostics at MSU. Forward and reverse primers, 5′-TTA GTG GTG TTT GCC GGG AC-3′ and 5′-TTC AGC TAA ATC GAC AAG GAG-3′, respectively, target a 168-bp fragment of the UL24 gene homologue of the EHV-1 genome (GenBank Accession no. X13209). Amplification was carried out in a 50-μl reaction volume containing 25 μl of the QuantiTect SYBR Green PCR mastermix, f 500 nM of each primer, and 5 μl of template DNA. Real-time PCR was performed in an iCycler iQ Real-Time Detection System with Optical System software version 3.1. g Cycling conditions were as follows: 95°C for 15 minutes and 40 cycles of 94°C for 30 seconds, 55°C for 30 seconds, and 72°C for 30 seconds. To determine the specificity of the amplicon, a postamplification melt curve analysis was incorporated in the run, consisting of raising the temperature in 0.5°C increments from 55°C to 95°C every 10 seconds. An EHV-1 positive sample exhibited a peak melting temperature of 87°C. The detection limit of the assay is 0.02 tissue culture infective dose (TCID50) per 5 μl (corresponding to 2 to 20 copies of EHV-1 DNA). Realtime TaqMan PCR assays h targeting the glycoprotein B (gB) gene of EHV-1 and the housekeeping gene glyceraldehyde-3-phosphate dehydrogenase (GAPDH) as quality control were used at UCD as previously reported. 17 For both EHV-1 PCR assays, the killed and modified-live vaccine strains and water were used as positive and negative amplification controls, respectively. Positive PCR results at the DNA level, if obtained, were to be further investigated by using absolute quantification, transcriptional activity at the messenger RNA level for the gB gene, and latency-associated transcripts as previously reported 16 to determine the viral state (dead, lytic, latent) present in either nasopharyngeal secretions or whole blood. Serum samples collected from all horses on the vaccination day and 21 days later were tested for antibodies against EHV-1 using a serum neutralization test 3 at MSU.
Serum viral neutralization titers in healthy horses vaccinated with a killed or a modified-life EHV-1 vaccine.*
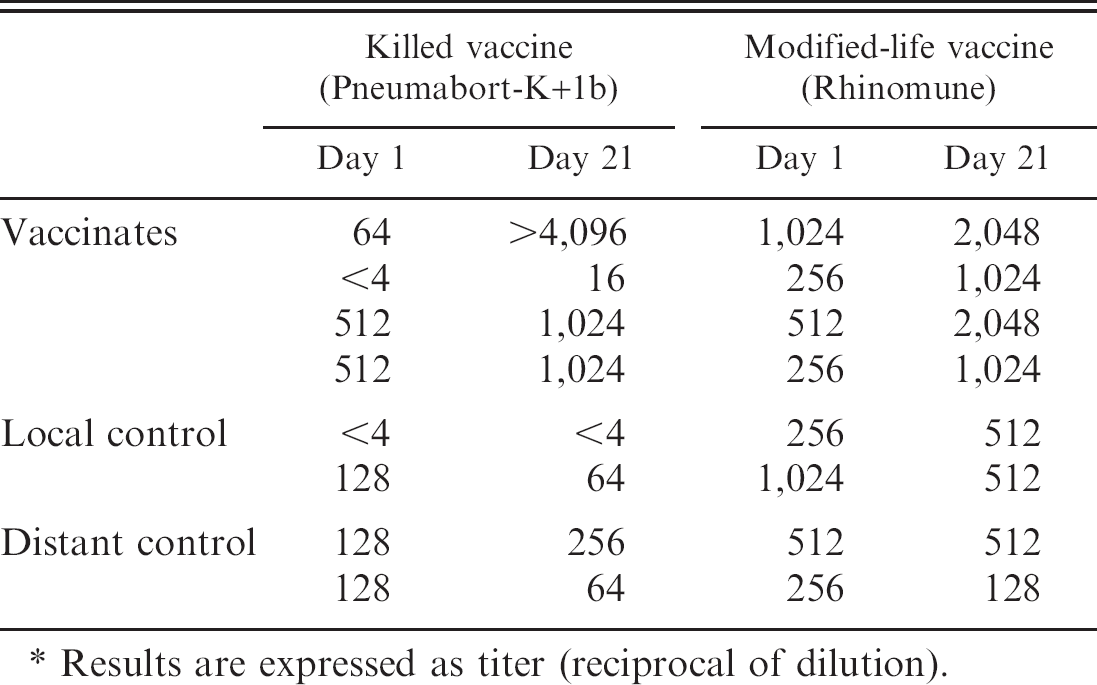
Results are expressed as titer (reciprocal of dilution).
Whereas the 2 vaccine strains tested positive in both EHV-1 PCR assays, nasopharyngeal swabs and whole blood collected from vaccinated and control horses had negative PCR test results for EHV-1 during the entire study period. Serum neutralization testing revealed a 2- to 4-fold increase in titers for all vaccinated horses, whereas titers in control horses were largely unchanged (Table 1).
In recent years, tissue distribution of novel vaccine constructs, such as plasmid vaccines, has been thoroughly investigated in mice, and it was found that the amount of plasmid DNA in tissues distant from the injection site decreases rapidly after vaccine administration. 11 However, biodistribution of viral DNA after intramuscular injection of killed or modified-live vaccines is poorly documented in the veterinary literature. We hypothesized that modifiedlive vaccine virus was more likely to spread from the intramuscular injection site via the systemic circulation to pathogen-specific predilection sites and more likely to be detected by PCR testing than killed vaccine virus. A variety of factors could contribute to the extent of tissue distribution of vaccine virus, including virulence of the attenuated pathogen, antigen mass, vaccine formulation, and host immune status at the time of vaccination. When the modified EHV-1 vaccine strain (RacH) was originally attenuated by serial passage in porcine kidney cells, isolation of the virus from blood and nasopharyngeal secretions of vaccinated horses was only successful when low-passage viral strains were administered. 12 Further, the RacH strain used in the commercial vaccine has never been detected from field cases of abortion or respiratory tract infections using Southern blot analysis 13 or PCR 14 for strain characterization. However, no study has specifically investigated the detection of EHV-1 vaccine strains after intramuscular administration using PCR. Detection of modified-live vaccine DNA by PCR has been reported from bursa of Fabricius tissue of specific pathogen-free (SPF) chicks vaccinated with infectious bursal disease virus, 1 tissues from sheep vaccinated with the Bartha vaccine strain of Aujeszky disease virus, 9 and peripheral blood mononuclear cells (PBMCs) of dogs and domestic ferrets vaccinated against canine distemper virus. 20 , 23 On the other hand, attempts to detect vaccine strains by PCR in feces of healthy dogs vaccinated with modified-live canine coronavirus strains were unsuccessful. 15 Kawashima et al. 10 were able to detect a vaccine strain of measles in peripheral blood mononuclear cells of 6 patients with autoimmune hepatitis but in only 1 of 12 healthy control patients, leading these authors to conclude that host immune status likely plays a role in the persistence of vaccine strains in peripheral blood leucocytes or organ tissues.
The main concern about the detection of EHV-1 vaccine strains in nasal secretions and/or whole blood that prompted this study was not related to EHV-1 virulence but rather how positive PCR test results should be interpreted in recently vaccinated horses. The increasing application of sensitive molecular diagnostic techniques for detection of pathogens in practice settings has presented new dilemmas with regard to how test results are interpreted and used by both equine practitioners and regulatory veterinarians. This pilot investigation failed to detect DNA from EHV-1 vaccine strains by PCR testing of nasopharyngeal secretions or blood collected from healthy horses for 4 weeks postintramuscular vaccination. In addition to the small number of horses in each vaccination group, additional limitations of this study included the use of healthy adult horses with preexisting antibodies to EHV, as well as sampling horses on a weekly versus daily schedule. As demonstrated by others, healthy individuals are less likely to have modified-live vaccine strains persist in tissues. 10 Polymerase chain reaction testing of samples collected from recently vaccinated stressed horses (e.g., after prolonged transport or racing) or horses hospitalized for treatment of other disorders could yield different (positive) results and has not been critically evaluated. The presence of serum antibodies against EHV is common in adult horses because of the ubiquitous nature of EHV infection and the routine practice of vaccinating horses against EHV-1/4. The presence of serum antibodies against EHV in this study's experimental horses could have neutralized vaccine virus at the injection site and prevented the virus from reaching other organ systems. Obtaining horses that are naive to EHV for research purposes is challenging, and the horses used in this investigation are more representative of the general population that would be vaccinated. In agreement with previous studies, both vaccines induced an anamnestic response. 7
The pilot study results demonstrate that detection of EHV-1 DNA by PCR in both vaccinated and unvaccinated healthy horses is not a common event. The molecular detection of EHV-1 in this study may have been compromised by prior vaccination of the study horses and sampling frequency; however, the study results are in disagreement with a recent study finding a molecular prevalence of latency (positive nested PCR assay results for EHV-1 in peripheral blood) of 47% in healthy horses. 7 This discrepancy may be related, in part, to different sensitivities of the PCR assays employed and the higher risk of carryover contamination associated with open-tube assays. Further, it is important to emphasize that positive test results using PCR assays only indicate detection of viral DNA and are unable to differentiate between live or dead virus or detection of latent or vaccine virus. Practicing veterinarians and regulatory officials that receive positive PCR test results may be unaware of the complexities involved in test interpretation and could decide to quarantine equine facilities or cancel competitions by misinterpreting test results. These possibilities support the need for a consensus on use and interpretation of molecular diagnostic techniques in evaluation of suspected field EHV-1 disease and other infectious diseases. Use of novel realtime PCR assays that discriminate between EHV-1 viral states is now possible by 1) targeting several genes (e.g., glycoprotein, latency-associated transcripts), 2) detecting viral genomic DNA and transcriptional activity of the target genes at the messenger RNA level, and 3) using absolute quantification. Standardized use of these novel PCR assays by both commercial and research laboratories is necessary for more accurate investigation of disease outbreaks in the future. As examples, viral threshold loads are used in selected human infectious diseases (i.e., HIV, HCV) to determine disease stage and response to antiviral therapy 18 , 19 and have recently been used diagnostically in EHV-4 infected horses to discriminate between lytic and nonlytic infection. 16
Acknowledgement. This project was supported by funding from Pfizer Animal Health, New York, NY.
Footnotes
a.
Pneumabort-K+1b, Fort Dodge Animal Health, Fort Dodge, IA.
b.
Rhinomune, Pfizer Animal Health, New York, NY.
c.
Fox converting swabs, Green Bay, WI.
d.
DNEasy Blood Kit, Qiagen, Valencia, CA.
e.
DEPC-Treated water, Ambion, Austin, TX.
f.
Qiagen, Valencia, CA.
g.
Bio-Rad Laboratories, Hercules, CA.
h.
ABI PRISM 7700 Sequence Detection System, Applied Biosystems, Foster City, CA.
