Abstract
In 2011, a 2-year-old horse in northern Queensland, Australia, was reported to have developed mild neurologic signs, and a blood sample was submitted for laboratory investigation. Virus isolation was performed using the blood sample, and an orbivirus was isolated. This was confirmed to be a strain of Elsey virus (ELSV) after transmission electron microscopy and nucleotide sequencing. The nucleotide sequence was compared with those in GenBank, and had 100% identity with ELSV previously reported from the Northern Territory, Australia. ELSV is taxonomically closely related to Peruvian horse sickness virus.
Keywords
The genus Orbivirus is the largest within the family Reoviridae, consisting of 22 known virus species. Reports of novel orbiviruses in new animal species are expanding the list of known orbiviruses.2,4 Orbiviruses are known to cause severe diseases of livestock, with arguably the greatest of economic impacts caused by Bluetongue virus (BTV) in cattle and sheep, Epizootic hemorrhagic disease virus (EHDV) in wild ungulates or cattle, as well as African horse sickness virus, Equine encephalosis virus, and Peruvian horse sickness virus (PHSV) in equids. 10 At least 7 serogroups of the known orbiviruses are endemic to Australia, infecting a wide range of host species, and known to cause mild to severe diseases in these host animals. 3 However, most of these infections remain asymptomatic.
The most extensively studied orbivirus to date is BTV, and other orbiviruses share morphologic and genomic similarities with BTV. 13 Structurally, the outer capsid proteins (VP2 and VP5) are more variable than components of the virus core, or the nonstructural proteins. The outermost protein, VP2, is encoded by segment-2 of the double-stranded (ds)RNA genome, and is an appropriate target for the development of type-specific, nucleic acid–based assays. 7 The pan-clade orbivirus PCR, known to detect the majority of the orbiviruses, is based on VP2 sequence present in GenBank (http://www.ncbi.nlm.nih.gov/; Oakey, unpublished). The genome segments coding for the virus core and/or the nonstructural proteins can be used as targets for serogroup (virus species)-specific, reverse transcription (RT)-PCR–based assays.2,13
In 2011, a 2-year-old horse in northern Queensland, Australia, was showing very mild neurologic signs. The ethylenediamine tetra-acetic acid (EDTA) blood sample collected was submitted to Biosecurity Queensland for a disease investigation. Simultaneously, southern Australian states had reported cases of arboviral infections in horses with a focus on flaviviruses and alphaviruses.5,11,12 Considering the nature of these outbreaks in equids, the blood sample was tested with a panel of arboviral PCRs that included flavivirus, 9 alphavirus, 14 and orbivirus (Oakey, unpublished). The sample tested negative for alphaviruses and orbiviruses. The sample tested for flavivirus was inconclusive, with respect to the positive control used in the assay, and, on that basis, was further subjected to virus isolation. With the history of circulating Murray Valley encephalitis in Queensland horses, 6 virus isolation was attempted using 2 different cell lines. Buffy coat from the blood sample was inoculated into Aedes aegypti (AA) C6/36 cells and incubated at 28°C for 2 weeks. The inoculum from passage-1 AA cells was further subcultured into BSR cells, a clone of baby hamster kidney (BHK)-21 cells, for an additional week at 34°C with supplementation of 5% CO2. The AA and BSR cell lines were maintained in a minimum essential medium supplemented with 10% fetal bovine serum a and the following antimicrobials: penicillin (100 IU/mL), streptomycin (100 μg/mL), and amphotericin B (2 ng/mL). a At the end of passage 2 in BSR cells, cells showed cytopathic effect (CPE) characterized by severe cytoplasmic vacuolation, stringing, rounding-up, and detachment. Supernatants from cell culture samples showing CPE were tested for flavivirus by PCR, which produced negative results. The samples were examined with negative staining transmission electron microscopy (TEM). The TEM revealed the presence of icosahedral particles ~60–70 nm (Fig. 1), suggesting the presence of an orbivirus in the cell culture supernatant.
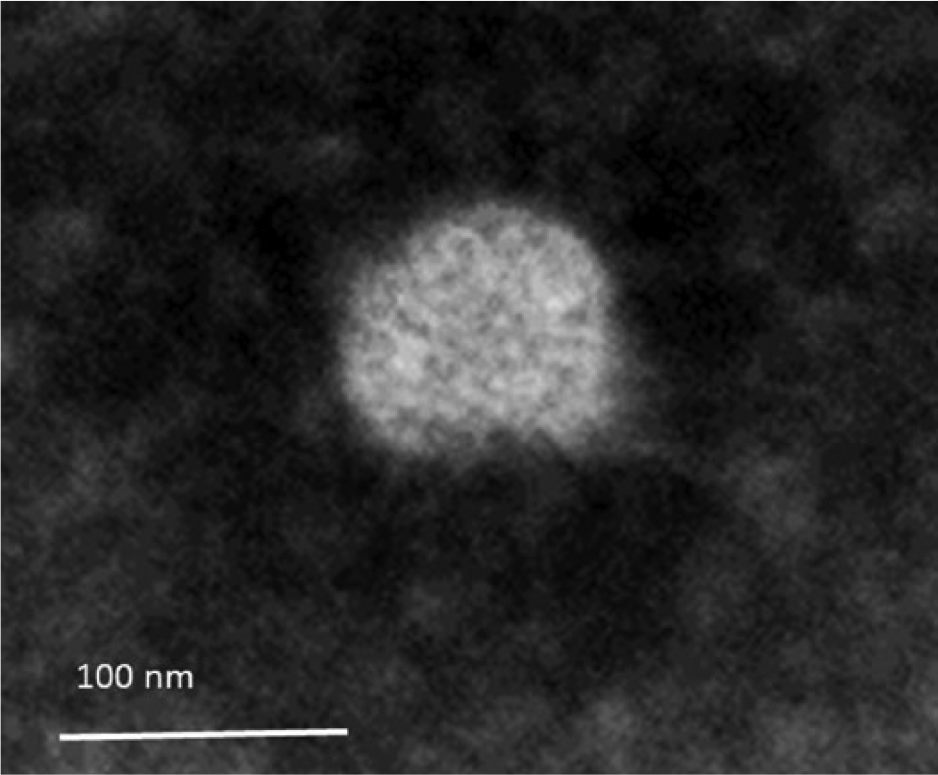
Transmission electron micrograph by negative staining showing presence of equine orbivirus, later confirmed as an Elsey virus. Bar = 100 nm. 100,000×.
Numerous orbiviruses are known to be endemic to Australia, including viruses from the Wallal and Warrego groups and Corriparta virus. Members of these groups have been involved with diseases in several species, and some of these viruses are known to cause neurologic signs. 3 In Queensland, ~2,573 samples from horses with reported neurologic signs were submitted to Biosecurity Queensland in the period 2010–2014, and <0.5% of these animals had a definitive diagnosis in terms of identifying and/or confirming the disease agent. This could indicate that arboviruses with the capacity to cause neurologic diseases are not being detected. Moreover, very little information is available for these viruses from horses in Australia.
During 1999, 2 horses from different locations in the Northern Territory (NT) of Australia (Katherine and Elsey Stations, Mataranka), showing signs of neurologic disease, were investigated. In both cases, an orbivirus, which was consequently named Elsey virus (ELSV), was isolated. 1 Subsequent nucleic acid sequencing showed that it was very similar to PHSV from South America, a mosquito-transmitted orbivirus that causes encephalitis among horses. The 97–100% similarity for individual genes indicated that they both belong to PHSV serotype 1. 1 The epidemiology of the infections caused by PHSV and Elsey is poorly characterized.
The Queensland horse isolate obtained in 2011 was tested by PHSV PCR using primers Phsv-1101F (TTTGCATTATGTTATCCAAATCA) and Phsv-1510R (TCAGAATCATAACCTAAATAACA) designed to amplify a 410-bp fragment of segment 2 of the dsRNA fragment of PHSV. Briefly, viral RNA was isolated from cell culture supernatants showing CPE, using a robotic viral nucleic acid extraction kit b following the manufacturer’s instructions. PCR was performed using a 1-step RT-PCR kit, c with final concentrations of each primer at 400 nM, and using 5 µL of heat-denatured (95°C for 5 min) RNA extract as template. Reverse transcription was performed at 50°C for 30 min, hot-start polymerase activation at 94°C for 2 min, amplification by 35 cycles of 94°C for 30 s, 53°C for 30 s, 68°C for 45 s, and a final extension at 68°C for 5 min. Unused reaction components were removed using commercial enzyme d following the manufacturer’s instructions, and nucleotide sequencing was performed using a dideoxynucleotide sequencing kit. e The products were purified f and resolved using a genetic analyzer. g
Nucleotide sequences were aligned using base-calling and alignment software, h and the sequence was compared to those in GenBank. Distance tree of BLAST results of the Queensland virus sequence was generated using the neighbor-joining method (Fig. 2). The sequence clustered with previously published sequences representing PHSV (DQ248058) and ELSV (FJ225396). All 3 sequences were aligned using ClustalW software (alignment not shown), and only 1 nucleotide variation was observed between the Queensland isolate and the previously described isolate from NT (position G66A) and the Peruvian isolate (position A46G). On translation, the partial VP2 sequence from the Queensland isolate showed 100% identity with the ELSV sequence from NT (FJ225396). The 100% identity for partial VP2 outer coat protein between Australian isolates indicates that the Queensland virus belongs to serotype 1. Interestingly, there was no noted identity with VP2 of other reported orbiviruses, including BTV or EHDV, that are well described and represented in GenBank. The pan-clade PCR for orbiviruses is based on VP2 sequences present in the GenBank database. However, with noted differences in VP2 of these orbiviruses compared to ELSV sequences, the pan-orbivirus PCR failed to detect ELSV.
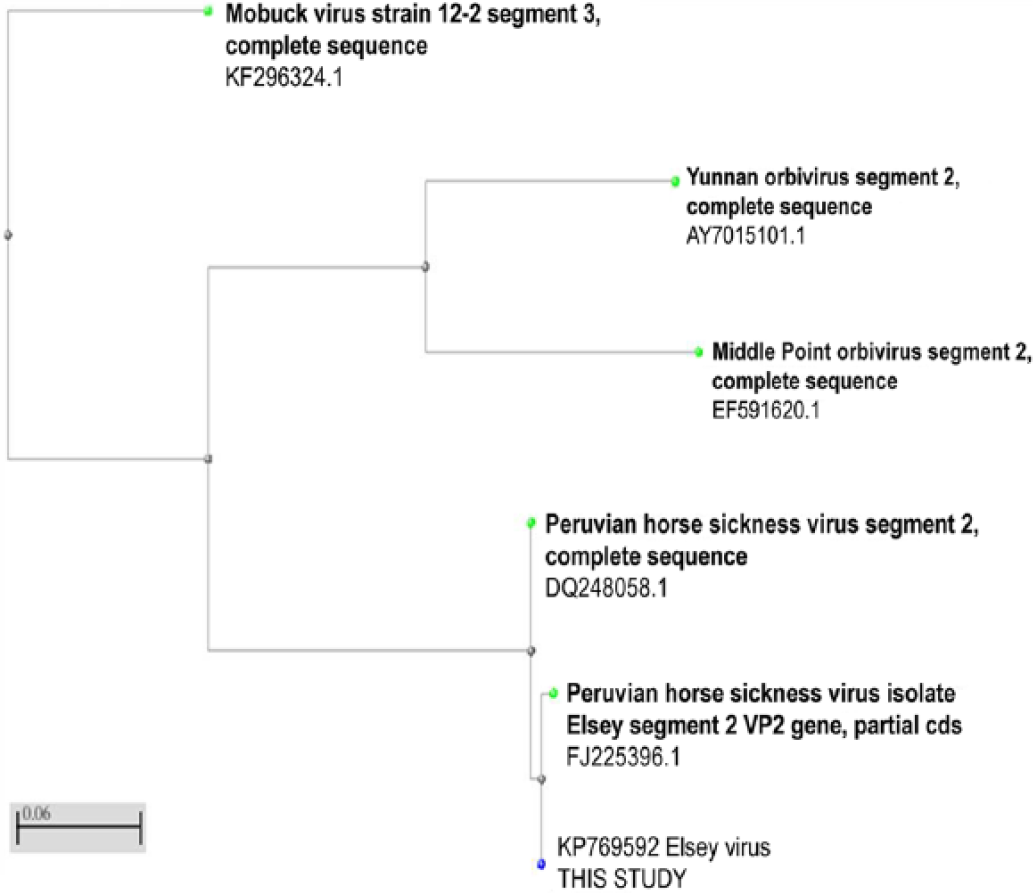
Distance tree of BLAST results using neighbor-joining method, and created within BLAST of our nucleotide sequence (http://www.ncbi.nlm.nih.gov/).
Reports on retrospective serologic surveillance on sick and healthy horse samples collected since the 1999 incident in NT showed an ~11% seroprevalence for antibodies to ELSV by immunoglobulin G enzyme-linked immunosorbent assay. 1 Although the possibility that ELSV causes neurologic syndromes in horses may be reasonably high, these viruses have also been isolated from healthy horses. Significant differences between Australian ELSV and PHSV from South America were noted. PHSV has been reported to cause no CPE in mammalian cells; however, extensive CPE is observed in BSR cells infected with Australian ELSV, although this was after enrichment in AA cells. Clinically, these 2 viruses differ, as PHSV in South America causes high mortality in infected horses whereas the Australian horse infected with ELSV in 2011 recovered after a few weeks. During the 1999 incident in NT, both sick horses were euthanized after the onset of illness, therefore the severity of the disease caused by ELSV was not fully assessed. However, in 2011, in the absence of a definitive diagnosis, the Queensland horse was not euthanized and later was reported to have recovered from the infection.
Australian horses are known to become infected with viruses such as Murray Valley encephalitis virus, Equid herpesvirus, Hendra virus, and Kunjin virus, which is a subtype of West Nile virus. 6 Although several of these infections are fatal, some are associated with neurologic syndromes. As new viruses are being reported over time 8 and more confirmed cases of infections become available, it is necessary to consider the role of orbiviruses, especially in neurologic cases, to help provide definitive diagnoses. Our study describes ELSV infection of horses in Queensland, although detected 15 years after the incident in Northern Territory. There is a probability that the virus remained unnoticed either because of its transient nature in infected animals or because disease is an uncommon event. It is also possible that most of the infected horses remain normal and that the virus is detected infrequently because virus isolation is not routinely undertaken on these animals. However, more evidence is required to confirm either of these theories. Overall, more data is required to determine the underlying rate of orbiviral infections in horses in Queensland. The accession number for the sequence submitted to GenBank is KP769592
Footnotes
Acknowledgements
We thank the laboratory staff of Tropical and Aquatic Animal Health Laboratory, Townsville, and Biosecurity Sciences Laboratory, Brisbane, Queensland for their work in processing of samples for this case and reviewing the manuscript. Electron microscopy was performed at the University of Queensland; we thank Howard Prior and Rick Webb for electron microscopy work.
Authors’ contributions
K Agnihotri contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; critically revised the manuscript; and gave final approval. B Pease contributed to analysis and interpretation of data. J Oakey contributed to analysis and interpretation of data, and critically revised the manuscript. G Campbell contributed to conception of the study, and contributed to acquisition of data. K Agnihotri, B Pease, and J Oakey agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Sigma-Aldrich, St. Louis, MO.
b.
EZ-1 viral mini kit V2.0, Qiagen Pty Ltd, Doncaster, Victoria, Australia.
c.
Superscript III one step kit (endpoint), Life Technologies Australia Pty Ltd, Mulgrave, Victoria, Australia.
d.
USB Affymetrix ExoSAP-IT, In Vitro Technologies Pty Ltd, Cleveland, Queensland, Australia.
e.
Big Dye Terminator V3.1 kit, Life Technologies Australia Pty Ltd, Mulgrave, Victoria, Australia.
f.
DyeEX II kit, Qiagen Pty Ltd, Doncaster, Victoria, Australia.
g.
ABI 7500 XL, Applied Biosystems, Foster City, CA.
h.
Sequencher v4.8 software, Gene Codes Corp., Ann Arbor, MI.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Departmental financial support.
