Abstract
Equine coronavirus (EqCoV) infection has been documented in most reports through quantitative qPCR analysis of feces and viral genome sequencing. Although qPCR is used to detect antigen during the acute disease phase, there is no equine-specific antibody test available to study EqCoV seroprevalence in various horse populations. We developed an enzyme-linked immunosorbent assay (ELISA) targeting antibodies to the spike (S) protein of EqCoV and validated its use, using acute and convalescent sera from 83 adult horses involved in 6 outbreaks. The EqCoV S protein–based ELISA was able to reliably detect antibodies to EqCoV in naturally infected horses. The greatest seroconversion rate was observed in horses with clinical signs compatible with EqCoV infection and EqCoV qPCR detection in feces. The EqCoV S protein–based ELISA could be used effectively for seroepidemiologic studies in order to better characterize the overall infection rate of EqCoV in various horse populations.
Coronaviruses are part of the Coronaviridae family, which are positive-strand RNA viruses. The Coronavirinae subfamily is divided into 4 genera (Alphacoronavirus, Betacoronavirus, Deltacoronavirus, and Gammacoronavirus), based on serologic cross-reactivity and genetic differences.8,20 Equine coronavirus (EqCoV) is in the Betacoronavirus genus, as are Human coronavirus HKU1, Murine coronavirus (murine hepatitis virus; rat coronavirus), and Betacoronavirus 1 (EqCoV; bovine coronavirus; porcine hemagglutinating encephalomyelitis virus; and canine respiratory coronavirus). 21 EqCoV has been considered a pathogen affecting foals, but since 2010 there have been several reports from Japan, Europe, and the United States on outbreaks in adult horses.9,13,14,17 The most common clinical signs observed in adult horses infected with EqCoV are anorexia, lethargy, and fever, and less frequently diarrhea, colic, and neurologic deficits. The morbidity rate in outbreak situations has been 20–57%, and ~11% of clinically infected horses died or were euthanized because of endotoxemic shock, septicemia, or hyperammonemia-associated encephalopathy.4,5,10,13,17 Historically, the detection of EqCoV in fecal samples has been based on electron microscopy, culture, or antigen enzyme-linked immunosorbent assay (ELISA).3,6 Quantitative PCR has proven to be a sensitive and specific detection tool to determine fecal shedding during the acute phase of the infection. The use of EqCoV-specific serologic assays has, to our knowledge, not been investigated. Such assays would have the advantage of determining overall infection rates in specific horse populations and assessing individual infection using acute and convalescence serum samples. Therefore, the objectives of our study were to develop an ELISA targeting antibodies to the spike (S) protein of EqCoV and to validate the ELISA using serum samples from naturally infected adult horses involved in contemporary outbreaks.
The S2 region of the S protein of EqCoV was selected to establish the ELISA. For this purpose, amino acid sequences of the S protein of known EqCoV strains were lined up and reviewed (NC 99, GenBank accession AY316300; Tokachi 09, GenBank AB555560). Computer modeling predicted 3 immunodominant areas of the S protein.
a
A 250–amino acid (AA) fragment of the S protein with an additional 24 amino acids for expression and His-tag ((MASMTGGQQMGRGS) 610-SGATCSTDLQ RSNTEIVVGV CVKYDLYGIT GQGIFIEVNA
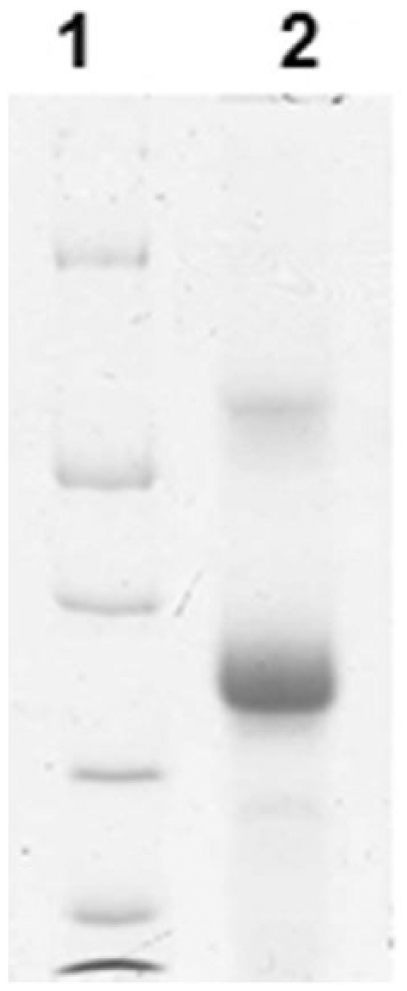
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis demonstrating the recombinant equine coronavirus S protein of ~30 kDa molecular weight (protein markers from top to bottom are 66, 45, 35, 25, and 18 kDa).
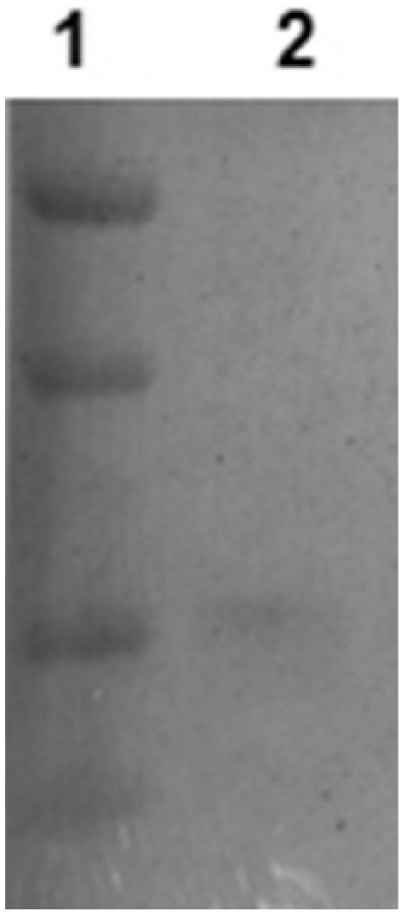
Western blot detecting the recombinant equine coronavirus (EqCoV) S protein of ~30 kDa molecular weight using a serum pool from 5 horses with confirmed EqCoV infection (protein markers from top to bottom are 56, 42, 28, and 21 kDa).
Microtiter plates were coated with 100 µL of recombinant S protein diluted in coating buffer c at a concentration of 0.156 µg/mL. Plates were then covered and stored at 4°C overnight. Serum samples from the study horses previously stored at −20°C were thawed overnight at 4°C. On the day of the analysis, the coated plates were washed 4 times with 200 µL of wash buffer c per well and gently tapped until dry. Then each well received 99 µL of sample dilution buffer c and 1 µL of serum; each sample was run in singlet. Optimal S protein and serum dilutions were determined prior to assay validation using standard checkboard titration procedures. Negative controls consisted of serum samples collected from 3 healthy adult horses from the university herd with no history of systemic disease and qPCR-negative feces for EqCoV. Positive controls consisted of serum samples collected from 3 horses 30 days after these horses developed clinical signs compatible with EqCoV infection and tested qPCR positive for EqCoV in feces. After the serum samples were loaded into the wells, the plates were covered and wrapped in aluminum foil and incubated for 2 h at room temperature on a titer plate shaker. Thereafter, the plates were washed 4 times, and 100 µL of diluted anti-horse IgG horseradish peroxidase conjugate d (dilution of 1:15,000 in 2% milk) was added. This step was followed by 1 h incubation as mentioned above. After washing the plate 4 times, 100 µL of enzyme substrate c was added to each well. The plate was then incubated at room temperature for 10 min. As a final step, 50 µL of stop solution (4.89 mL of 98% sulfuric acid diluted with 495 mL of distilled water) was added to each well. The optical density (OD) was measured at 450 nm in a microplate photometer. e The OD was measured within 15 min of adding the stop solution.
An OD cutoff titer was determined testing known positive (15 sick and EqCoV qPCR–positive horses) and known negative (47 healthy and EqCoV qPCR–negative horses) samples and performing receiver operator characteristic analysis. A pool of 3 positive and a pool of 3 negative control samples were used on each plate to determine quality control of the assay. Intra-assay coefficients of variation (CVs) were determined using 3 replicates of a total of 4 serum samples run on the same plate. Interassay CVs were determined using the same serum samples run in triplicate on 2 different plates on 2 different days. Analytical specificity of the S protein EqCoV ELISA was determined using pre- and post-vaccine serum samples from horses vaccinated with commercial equine vaccines containing the following antigens: Equid herpesvirus 1, Equid herpesvirus 4, equine Influenza A virus, Equine arteritis virus, Eastern equine encephalitis virus, Western equine encephalitis virus, and West Nile virus.
Six independent outbreaks of EqCoV were investigated. The outbreaks occurred between March and September 2014 in the states of New Hampshire, South Carolina, Oregon, and California. In each of these outbreaks, EqCoV was found as the sole etiologic agent associated with clinical disease as reported previously.4,13,14,17 The population of horses per equine premises was 6–24 horses, and morbidity was 14–83% (Table 1). For each of the 83 study horses, clinical parameters, feces for EqCoV qPCR analysis, and acute and convalescent serum samples for EqCoV serology were available. The convalescent serum samples were collected 11–35 days following the first serum samples. The clinical signs observed in diseased horses included anorexia (96%), lethargy (92%), fever (84%), diarrhea (40%), and colic (28%). Based on the presence or absence of clinical signs and the qPCR results, the 83 horses were divided into 4 groups: diseased and EqCoV qPCR–positive (n = 15); diseased and EqCoV qPCR–negative (n = 10); asymptomatic and EqCoV qPCR–positive (n = 11); and asymptomatic and EqCoV qPCR–negative (n = 47; Table 1).
Demographic, clinical, and molecular results from 83 horses involved in 6 separate equine coronavirus outbreaks in 2014, by U.S. state and month.*
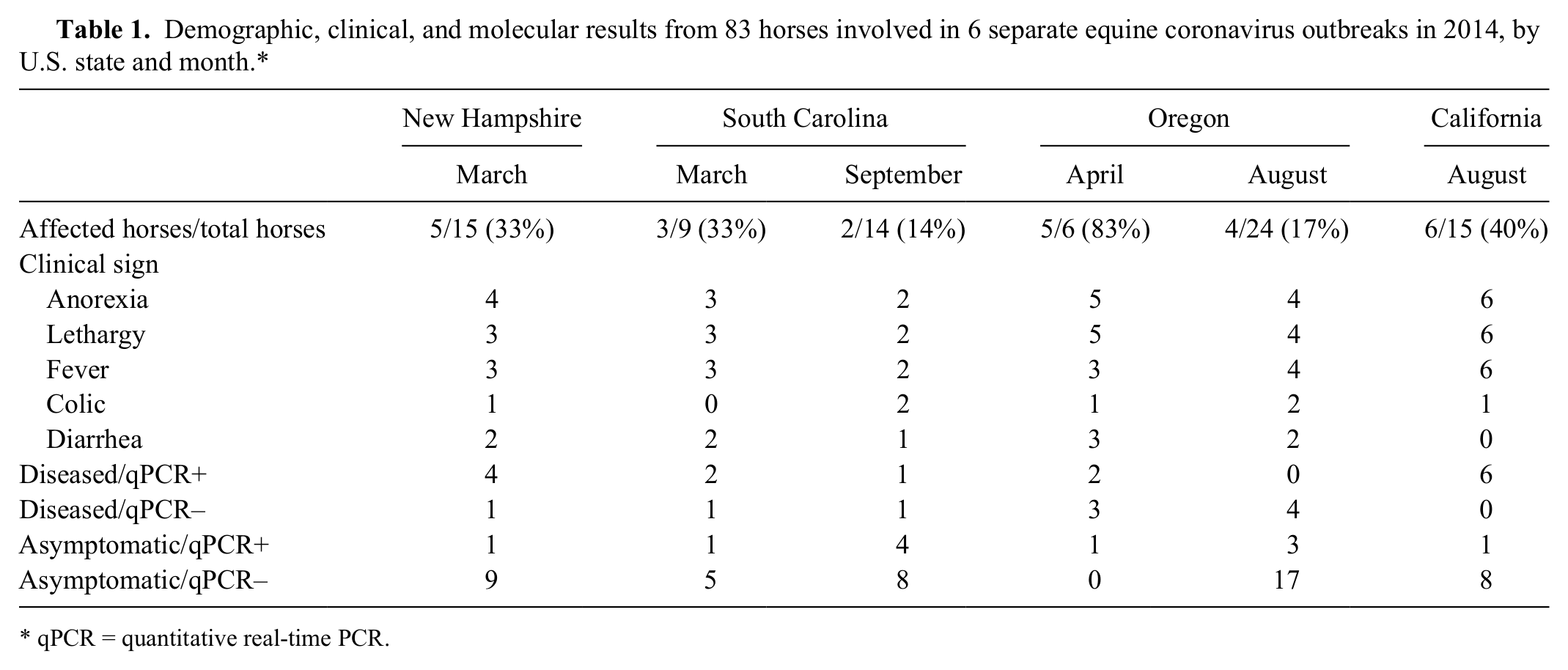
qPCR = quantitative real-time PCR.
The ELISA was able to reliably and repeatedly classify negative and positive control sera using an OD cutoff of 1.958. At that cutoff titer, the EqCoV S protein ELISA had a sensitivity of 100% and a specificity of 90.48%. The assay was found to be linear over an OD range of 0.557–3.487. Intra-assay CVs were 0.8–6.4%, with an average of 3.1%. The interassay CVs were 1.9–4.2%, with an average of 3.0%. None of the antisera against non-EqCoV vaccine–derived viruses yielded a positive OD on either the pre- or postvaccine sera.
Seroprevalence during the acute onset of the 6 outbreaks was 26% for all 83 study horses, with a range of 9–70% depending on the clinical and qPCR status. Seroprevalence in convalescent serum samples increased to 100% for diseased EqCoV qPCR–positive horses and 70% for diseased EqCoV qPCR–negative group horses. For asymptomatic horses, the seroprevalence in the convalescent serum samples increased to 27% for EqCoV qPCR–positive and 17% for EqCoV qPCR–negative horses. Regardless of the qPCR results, 86% of the sick horses tested seropositive at the time the convalescent sample was taken. In the group of healthy horses, this percentage was 21%. The rate of seroconversion was 80% for diseased, EqCoV qPCR–positive horses; 10% for diseased, EqCoV qPCR–negative horses; 18% for asymptomatic, EqCoV qPCR–positive horses; and 6% for asymptomatic, EqCoV qPCR–negative horses.
Hence, the EqCoV S protein–based ELISA was able to reliably detect antibodies to EqCoV. The greatest seroconversion rate was observed in horses with clinical signs compatible with EqCoV and EqCoV qPCR detection in feces. We recognize the limitation of using study cases and group assignment based solely on clinical and qPCR status. Unfortunately, there is no established antemortem gold standard to confirm EqCoV infection.
The commercially available bovine coronavirus ELISA kit f uses whole virus as antigen, and the test has been shown to have a sensitivity of 84.6% and a specificity of 100%.1,12 In the present study, all 15 diseased and EqCoV qPCR–positive horses had evidence of specific antibodies to EqCoV in the convalescent serum samples, which reflects the high sensitivity of the S protein–based ELISA. The present ELISA was based on recombinant S protein of EqCoV because of the technical difficulty in growing large amounts of EqCoV. Amino acid sequences of the spike protein of EqCoV have been shown previously to be highly conserved among different strains of EqCoV. 13 The use of recombinant protein for the detection of antibodies to various pathogens using an ELISA is routinely accepted and has the advantage of producing a pure and high-quality antigen.11,18,19
Although natural outbreaks represent true dynamic situations, the timing of initial sample collection to determine serologic responses can vary depending on the identification of the initial index case(s). The present study showed that 26% of study horses had detectable EqCoV antibodies at the time the acute serum samples were collected, with group seroprevalences varying depending on the disease and qPCR status. The incubation period of EqCoV is short, and clinical disease develops 48–72 h after natural and experimental infection.5,10 Clinical signs persist for a few days to 1 week and generally resolve with minimal supportive care. In experimentally infected horses, the detection of EqCoV by qPCR in feces occurs 24 h after horses become febrile (day 2–4 postinfection), and fecal shedding persists for 10–12 days. 10 The viral kinetics of EqCoV and their association with clinical disease support the finding that most of the diseased and EqCoV qPCR–positive horses experienced seroconversion, whereas seroconversion in diseased or asymptomatic EqCoV qPCR–negative horses was low. The low seroconversion rate in diseased and asymptomatic qPCR-negative horses is consistent with previous studies on enteric equine pathogens. 15 Although the pathogenesis of EqCoV has only been described in a small number of clinically infected horses, the severity of clinical disease, magnitude and duration of viral fecal shedding, and serologic responses likely depend on the degree of enteritis induced by EqCoV.4,5 One needs to take into account that some horses remain asymptomatic after infection. The percentage of asymptomatic horses determined by qPCR has been observed to be 11–29%. 17 In the present study, 11 of 58 (19%) asymptomatic horses had evidence of recent infection with EqCoV based on detectable antibodies in convalescent serum samples.
Footnotes
Authors’ contributions
LJ Kooijman contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; and critically revised the manuscript. SM Mapes contributed to conception and design of the study; contributed to acquisition and analysis of data; and critically revised the manuscript. N Pusterla contributed to conception and design of the study; contributed to acquisition, analysis, and interpretation of data; drafted the manuscript; critically revised the manuscript; gave final approval; and agreed to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Notes
Declaration of conflicting interests
The author(s) declared no conflicts of interest with respect to the research, authorship, and publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The study was supported by the Center for Equine Health, School of Veterinary Medicine, University of California, Davis, with additional contributions from public and private donors.
