Abstract
A potential mechanism by which highly pathogenic avian Influenza A virus subtype H5N1 could more readily infect human beings is through the infection of and adaptation in pigs. To detect the occurrence of such infection, monitoring of pig populations through serological screening would be highly desirable. In the current study, hemagglutination inhibition assays were able to detect antibodies against H5N1 developed in pigs, but because of antigenic variation between clades, the use of multiple virus strains were required. Whole recombinant virus and recombinant hemagglutinin antigen enzyme-linked immunosorbent assays (ELISAs) were generated that could detect antibody against multiple H5N1 strains, but which also detected antibody against endemic swine influenza viruses. A recombinant hemagglutinin antigen-based ELISA was as effective as the whole virus antigen ELISAs in detecting antibody against the H5N1 virus strains used and eliminated nearly all of the cross-reactivity with non-H5N1 virus antibody. The current study also highlighted the difficulty in establishing a decision (cutoff) value that would effectively counterbalance nonspecific reactivity against sensitivity. The results provide important information and considerations for the development of serological screening assays for highly pathogenic avian H5N1 viruses.
In 1997, a severe influenza-like illness was described in human patients in several countries in Southeast Asia. Diagnostic testing determined that they had been infected with a highly pathogenic avian Influenza A virus (HPAI) from the H5N1 subtype.2,9,11 Infection of human beings by avian influenza virus is uncommon, at least in part because avian influenza viruses prefer α-2,3 sialic acid glycoprotein receptors. The relatively few α-2,3 viral receptors in the upper respiratory tract of human beings may reduce susceptibility to infection by airborne avian virus and decrease the likelihood of transmission to a human host. 10 Nonetheless, direct interspecies transmission of avian influenza to human beings is possible and is postulated to have been responsible for the 1918 influenza pandemic. 10
Beyond the high mortality induced in poultry, the greatest concern with HPAI is the possibility of the virus adapting and establishing itself within the human population either through regular interspecies infection and horizontal transmission within the human population or through adaptation in an intermediate host, such as the pig. Evidence suggests the pig can serve as a “mixing vessel” supporting both human and avian influenza virus replication, attributed to the presence of both α-2,6 (human-like) and α-2,3 (avian-like) sialic acid receptors in the upper respiratory tract. 3 In contrast to human beings who in severe cases may require medical attention after infection with HPAI H5N1, pigs are only mildly affected by experimental or natural infection, suggesting the clinical impact of the virus on swine is minimal.1,6,7 The lack of clinical disease could result in underreporting of HPAI H5N1 infection in pigs, allowing the virus to persist in a small number of pigs within a population and thus potentially facilitating adaptation. An HPAI H5N1 virus has been isolated from swine in Indonesia that expressed a preference for the human-like α-2,6 sialic acid receptor suggesting that viral adaptation had occurred. 7 Because of the potential risk that HPAI H5N1 infection of pigs could pose to human health, as well as the potential negative economic impact on the swine industry itself, development of screening tests that can identify H5N1 HPAI infection in pigs would be valuable tools in preventing establishment of this virus in swine populations. 8 A serologic assay would be a good adjunct to polymerase chain reaction (PCR)–based tests for virus as antibody will circulate in serum for a longer duration than virus will be shed in nasal secretions.
To be available for use under biosafety level (BSL) 2 conditions at which veterinary diagnostic laboratories operate, H5N1 viruses in the assay would need to be attenuated. Three recombinant H5N1 (rH5N1) viruses (A/Vietnam/1203/04 [rVN04], A/Whooper Swan/Mongolia/244/05 [rWS05], and A/Japanese White Eye/Hong Kong/1038/06 [rJWE06] f ) were created using reverse genetics as previously described, 3 to attenuate the viruses for use in a BSL2+ environment. a Briefly, the viruses encode the hemagglutinin (HA) and neuraminidase genes from 1 of the H5N1 isolates as well as the remaining 6 genes from A/Puerto Rico/8/34 (PR8). The viruses were grown in either embryonated chicken eggs or Madin–Darby canine kidney cells. b The HA units (HAU)/ml were determined for each virus using 2-fold serial dilutions of the virus preparations in phosphate buffered saline (PBS) and incubating with 1% turkey red blood cells (RBCs) in PBS. The virus solutions were placed in 100-mm dishes and then ultraviolet (UV)-inactivated for 3 min using a UV crosslinker. c Ultraviolet inactivation of the virus was confirmed to be replication deficient using an immunocytochemistry TCID50 assay, as previously described. 4 Virus suspensions were diluted with PBS to a concentration of 128 HAU/ml, with the exception of the initial preparation of rWS05 that had a concentration of 96 HAU/ml.
To obtain H5N1-specific antiserum for use in the development of immunological assays for screening of H5N1 antibodies, 5-week-old pigs were vaccinated with each of the 3 recombinant viruses. The vaccine preparations also contained a final concentration of 20% v/v adjuvant. d Pigs were vaccinated and booster doses administered at 14 and 33 days post initial vaccination (DPV). Serum samples were collected prior to vaccination and at 8, 14, 22, 33, and 40 DPV. All animal work was performed in compliance with the National Animal Disease Center Institutional Animal Care and Use Committee guidelines.
Antibody titers for the serum samples were determined using hemagglutination inhibition (HI) assay. Serum samples were treated with receptor destroying enzyme (RDE e ; 100 µl of serum to 300 µl of RDE solution) and incubated overnight at 37°C. After RDE treatment, serum was brought to a final 1:10 dilution by the addition of physiological saline (0.85% w/v NaCl solution) and serially diluted into 96-well U-bottom plates using 2-fold serial dilutions ranging from 1:10 to 1:2,560. An equivalent volume of virus at 8 HAU/ml concentration was added to each well and incubated at room temperature for 1 hr. Another volume of 1% turkey RBCs in PBS was added, incubated for another hour, and HI titers were determined as the reciprocal of the highest dilution completely inhibiting agglutination. Reported HI titers are the geometric mean of 3 individual HI assay replicates. At 8 DPV, rVN04- and rJWE06-vaccinated pigs had HI titers of 80, while rWS05 pigs had HI titers of 160. By 40 DPV, geometric mean HI titers increased to 508 for both rVN04-vaccinated pigs, 1208 and 806 for rWS05-vaccinated pigs, and 640 for rJWE06-vaccinated pigs.
Additionally, HI assays were performed using the serum samples collected at 40 DPV to determine the relative HI antibody cross-reactivity between the rH5N1 viruses (Table 1). The heterospecific HI titers differed substantially depending on the viral strain used for antibody detection. The HI assays using rVN04 or rWS05 viruses against heterologous serum samples differed by 4- to 16-fold from HI titers measured against homologous virus. However, when the rJWE06 virus was used for detection of antibody in serum from pigs vaccinated with rVN04 and rWS05, the HI titers of the heterologous serum samples were similar to those measured in homologous serum (within a 2-fold difference). These results indicate that, similar to HI tests against endemic swine viruses, the ability to detect antibody against H5N1 in swine serum using HI assays is dependent on the viral strains used for detection and would likely require use of multiple strains in order to assure detection of all virus clades. 5
Heterospecific geometric mean hemagglutination inhibition assay titers comparing antibody titers against the recombinant (r)H5N1 viruses at 40 days postvaccination.*

Boldface indicates homologous hemagglutination inhibition assay titers.
To determine if antigenic variation among HPAI H5N1 viruses would similarly interfere with enzyme-linked immunosorbent assay (ELISA) format serologic testing, single antigen indirect ELISAs were generated using each rH5N1 virus. Additionally, to determine if potential cross-reactivity issues were characteristic of the hemagglutinins (as opposed to other virus proteins), a commercially available recombinant HA protein (rHA) from A/Vietnam/1203/04 k also was used as an antigen. Components for the ELISAs were optimized using a checkerboard format to compare concentration of antigens (2–256 HAU/ml of each of the rH5N1 viruses or 0.03–4 µg/ml recombinant HA protein from A/Vietnam/1203/05 f ) in different combinations with homologous serum sample dilutions (1:10–1:20,480). Additional assays were performed to optimize the conjugate antibody dilution (1:500–1:64,000). Using the optimized conditions (Table 2), plates g were coated with 100 µl of diluted rH5N1 virus or rHA resuspended in PBS and incubated overnight at 37°C. Plates were washed 3 times using PBS with 0.05% Tween 20 (PBST) and tapped onto paper towels between washes to remove residual liquid. Serum samples were then diluted with ELISA diluent (PBS with 0.5% gelatin, 0.15% Tween 20, and 4% goat serum), and 100 µl of a diluted sample was added to a well and incubated at room temp for 1.5 hr. After the plates were washed 3 times with PBST, 100 µl of anti-swine HRP h diluted 1:500 in ELISA diluent was added to each well and incubated for 1 hr at room temperature. The plates were washed 3 times with PBST prior to the addition of 100 µl of substrate (10 mg of o-phenylenediamine in 20 ml of 0.05 M phosphate citrate buffer, containing 0.03% sodium perborate). The plates were incubated until color development was complete (approximately 1 min) and stopped with 1 N sulfuric acid. Using a plate reader, g sample absorbance (optical density [OD]) was determined at a wavelength of 492 nm. The serum from the pigs vaccinated with the attenuated H5N1 viruses were tested by the above-described ELISAs.
Optimized enzyme-linked immunosorbent assay (ELISA) protocol conditions determined from checkerboard analysis for the whole virus and recombinant hemagglutinin ELISA assays.*
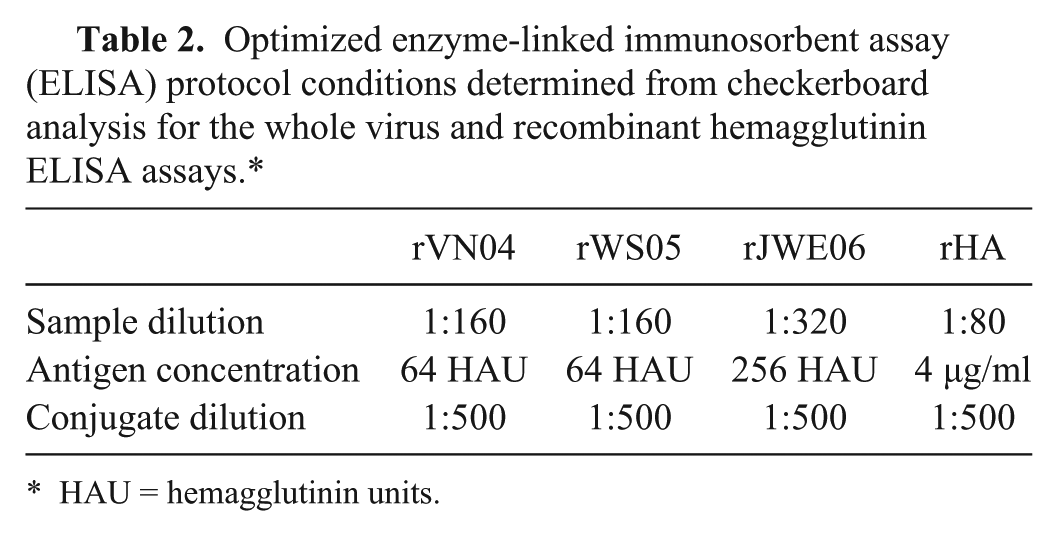
HAU = hemagglutinin units.
Because nearly all conventional swine have antibody to multiple endemic swine influenza viruses, cross-reactivity by these antibodies in the H5N1 ELISAs would preclude their use for surveillance. Serum from pigs infected in a previous study with influenza viruses (A/Swine/Wisconsin/R33f/01 [H1N2], A/Swine/Iowa/40776/93 [H1N1], A/Swine/Iowa/35233/99 [H1N1], A/Swine/Texas/4199-2/98 [H3N2], and A/Swine/Wisconsin/R7c/01 [H3N2]), 1 pig per virus, were also tested. 5 Homologous HI titers for the endemic swine serum samples ranged from 160 to 1,280.
All 3 recombinant ELISAs were able to detect antibody, both homologous and heterologous, in nearly all samples collected after 22 DPV but only inconsistently in serum samples collected earlier. The VN04 ELISA was able to detect antibody in more of the earlier samples. The HI test was more sensitive for all 3 recombinant viruses, detecting geometric mean antibody titers of 80–160. However, all 3 ELISAs also detected antibody in 8 of 10 of the non-H5 serum samples (5 samples run in 2 replicates). The rHA ELISA detected antibody in nearly all samples from the recombinant H5–vaccinated pigs collected after 22 DPV. However, antibody was detected by this test in only 2 of the replicate samples of non-H5 serum, and the OD values were just above the cutoff value.
Data from 2 independent ELISA replicates were combined and analyzed using receiver operating characteristics (ROC) i for determination of the cutoff values and calculation of the sensitivity and specificity for each assay (Table 3). For ROC analysis of these assays, all non-H5 serum samples collected between 8 DPV and 40 DPV, regardless of the virus used for vaccination, were considered positive. To minimize the potential for false positive results (which could pose serious economic difficulties for a herd under test), cutoff values were selected for each assay that eliminated detection of non-H5N1 serum samples; at these cutoff values, all 4 ELISAs had specificities of 100% (Table 3). Based on these parameters, the rWS05 and rJWE06 ELISAs appeared to have similarly low sensitivities of 37% and 38%, respectively. The rVN04 ELISA out-performed both of the other whole virus assays with a sensitivity of 48%. Analyzed in this manner, the results suggest that the majority of positive samples from rH5N1-vaccinated pigs were undetected with this assay. This apparent lack of sensitivity may have been artificially enhanced as a result of the decision threshold imposed by the relatively high cross-reactivity with the non-H5 antibodies. This cross-reactivity could be expected since the recombinant viruses contain a PR8 influenza backbone ancestrally related to classical swine influenza.
Specificity and sensitivity for each enzyme-linked immunosorbent assay determined through receiver operating characteristic analysis and calculation of cutoff values.
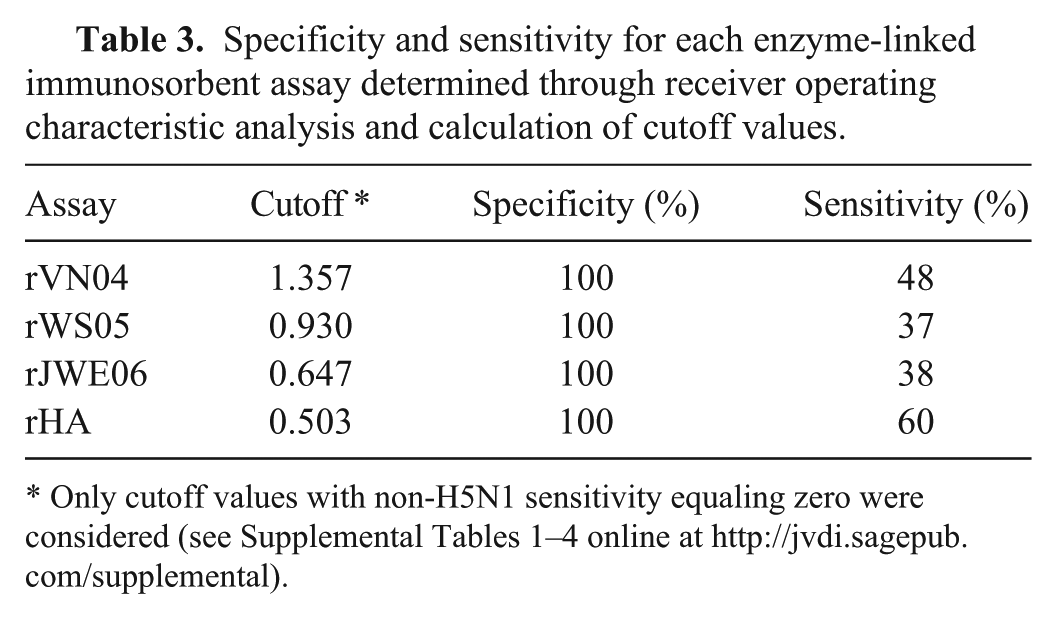
Only cutoff values with non-H5N1 sensitivity equaling zero were considered (see Supplemental Tables 1–4 online at http://jvdi.sagepub.com/supplemental).
In contrast, the rHA antigen ELISA had a greater ability to differentiate between anti-H5 and anti–non-H5 antibody. By ROC analysis, at the cutoff value (the highest OD value at which no non-H5 antibody was detected but at which H5N1 specificity was 100%) the H5N1 sensitivity was 60%, which is substantially higher than that of the whole recombinant virus assays. These results suggest rHA ELISAs may be preferable to recombinant whole virus assays. Selecting a lower cutoff value would result in an increase in sensitivity, but would decrease the ability of the assay to differentiate between endemic swine influenza H1 and H3 subtypes and H5N1 serum samples.
Serological assays provide a number of benefits when screening large populations for the prevalence of H5N1 influenza in pigs, especially the ability to detect the occurrence of infection for an extended period of time after active viral replication has ceased. As stated earlier, pigs may present with subclinical symptoms and go unnoticed through the course of infection, making acquisition of viral RNA for use in real-time RT-PCR assays difficult. This study utilized 3 different serological assays and illustrated the hurdles encountered when analyzing serum samples from potentially H5N1 infected pigs.
Results obtained in the current study indicate whole recombinant H5N1 ELISAs prepared as described would not be useful because of cross-reactivity with antibody against endemic swine viruses, but that the rHA indirect ELISA format may be more promising. In its current state, the rHA ELISA would not be useful as a differential diagnostic tool, but could be useful as a surveillance tool. Decreasing the cross-reactivity of the assay to non-H5N1 serum samples may increase the sensitivity, by allowing lower cutoff values. Such assay conditions in which detection of non-H5N1 sera is minimized may have been achieved by including non-H5N1 sera during concentration optimization procedures during development. Future work is needed to further optimize the ELISA to increase sensitivity and minimize the effects that heterosubtypic antibodies have on specificity. Even if sensitivity of the rHA ELISA could be improved through methodology improvements, antigen variation among H5 hemagglutinins might require use of a multiplex of different rHAs. The differences in cross-reactivity between H5 antigens exhibited in both HI and the whole recombinant virus ELISAs in the present study would suggest that as another difficulty.
A number of the commercially available H5N1 ELISAs for avian species also use rHA antigen to coat the plates, such as developed by Gentaur j and Green Spring. i Comparison of the 4 ELISA assays described herein with these commercially available H5N1-specific ELISAs may have proven valuable in development efforts. The other H5N1 ELISAs utilizing a similar platform appear to be primarily developed for use with avian serum samples and would require both modification and optimization of the assay for use with swine serum samples. However, such modification of a commercially available avian H5N1 ELISA kit may be a more cost efficient approach. In conclusion, this work lays the groundwork for the development of a serological assay for detecting antibodies against H5N1 that could be standardized for diagnostic use in pigs.
Footnotes
Acknowledgements
The authors would like to thank Dr. Richard Webby for providing the recombinant H5N1 virus for this project, and Dr. Chong Wang for performing the ROC statistical analysis for the ELISA data.
Declaration of conflicting interests
The author(s) declared no potential conflict of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding was provided by the National Center for Infectious Disease, Centers for Disease Control and Prevention, and the U.S. Department for Health and Human Services (1 UO1 CI000357-01).
a.
Richard Webby, St. Jude Children’s Research Hospital, Memphis, TN.
b.
MDCK cell line, American Type Culture Collection, Manassas, VA.
c.
Stratalinker 2400 UV crosslinker, Stratagene, La Jolla, CA.
d.
Emulsigen-D adjuvant, MVP Technologies, Omaha, NE.
e.
Receptor destroying enzyme, Lonza, Allendale, NJ.
f.
recombinant A/Vietnam/1203/04 HA protein, Protein Science Corp., Meriden, CT.
g.
Immulon 2HB plate and Multiskan plate reader with Ascent software, Thermo Scientific, Waltham, MA.
h.
goat anti-swine HRP-conjugated antibody, KPL, Gaithersburg, MD.
i.
Chong Wang Department of Veterinary Diagnostic and Production Animal Medicine, College of Veterinary Medicine, Iowa State University, Ames, IA.
j.
Avian influenza (H5N1) ELISA kit, Gentaur, Kampenhout, Belgium.
k.
Avian Influenza (H5N1) Elisa Kit, Green Spring, Shenzhen, China.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
