Abstract
An immunochromatographic strip was developed for the serological detection of pseudorabies virus (PRV) in swine. In the strip, the expressed protein of gB, one of the glycoproteins of PRV, labeled with colloidal gold, was used as the detector; staphylococcal protein A and swine anti–pseudorabies virus antibody were blotted on nitrocellulose membrane for the test and control lines, respectively. The specificity of the strip was 98.1%, and the sensitivity of the strip with reference anti-PRV serum was 96.0%. Swine serum samples (296) were collected to evaluate the characteristics of the strip in comparison with an existing commercial kit. The agreement was 93.6%. Furthermore, the dipstick assay based on the strip is rapid (5 min) and easy to perform with no requirement of professional skills, reagents, or equipment. This suggests that the immunochromatographic strip is an acceptable alternative for use in clinical laboratories lacking specialized equipment and for field diagnosis.
Pseudorabies virus (PRV) is the causative agent of Aujeszky’s disease. This pathogen has major economic consequences in pig husbandry,10,11 as it can infect livestock and wild animals, resulting in increased morbidity and mortality.2,3 In general, PRV mainly infects pigs at various production phases, causing high mortality in newborn piglets and abortion in pregnant sows, and results in significant economic losses for the pork industry. 3 The infected animals displayed multiple clinical signs, including high fever (usually 40–42°C), depression, anorexia, cough, shivering, diarrhea, and systemic neurological symptoms. Infected pigs were emaciated and weak in appearance, and ultimately died.8,12 Unexpectedly, growing and finishing pigs also died during this epidemic period, with a reported mortality rate of 10–30%.
Vaccination is a key strategy for the prevention and control of PRV in developing countries. To validate the efficacy of the vaccine, a rapid and simple test is needed in routine field practice to monitor antibody titers against PRV induced by vaccines. In our study, a simple and rapid assay (immunochromatographic strip test) was developed for the detection of anti-PRV antibody.
Pseudorabies virus gB belongs to the family of highly conserved herpesvirus glycoproteins that are implicated in the processes of virus entry and cell-to-cell spread.5,9 In herpesviruses, the N terminus and the regions upstream of the transmembrane-spanning domain of gB were shown to harbor epitopes associated with neutralization. 7 Previously, it was also shown that the majority of continuous and discontinuous epitopes of PRV gB identified were associated with complement-dependent virus neutralization and induced a strong antibody response during natural infection of swine. 15 Although no quantitative correlation has been reported between antibody level and clinical outcome, it seems likely that antibodies against immunodominant regions could mediate the clearance of virus and virus-infected cells and contribute to protective immunity against PRV.
In our study, the main antigen coding region of the gB (1,044 bp) gene was cloned from the vaccine strain PRV China/HB/98. The polymerase chain reaction products were inserted into a pET28a plasmid.
a
The recombinant plasmids were then transformed into Escherichia coli BL21 (DE3)
a
and protein expression induced with 0.4 mM isopropyl-β-
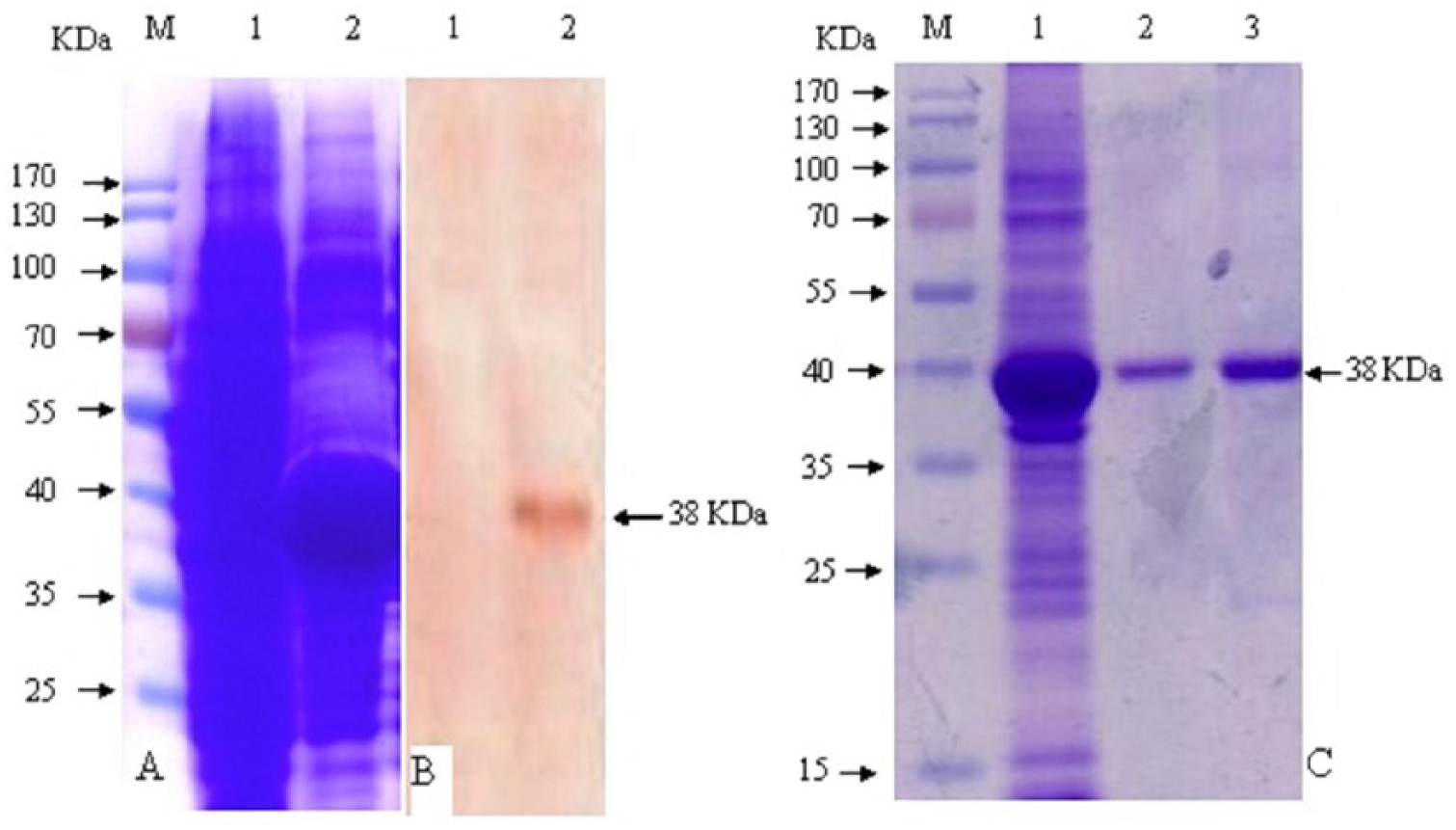
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and Western blot analysis of the expressed glycoprotein B (gB).
Approximately 200 μL of gB protein (1 mg/mL) solution was added to 10 mL of colloidal gold solution (pH 7.0) with constant stirring for 30 min, and then 1 mL 0.02 M sodium borate (pH 7.0) containing 10% bovine serum albumin (BSA) was added and then stirred for 10 min to stabilize the conjugate. The colloidal gold–labeled antigen was washed twice with wash buffer (0.02 M sodium borate, pH 7.0; 1% BSA; 0.1% sodium azide) by centrifugation at 20,000 × g and 4°C for 25 min. The precipitate was resuspended in 1 mL wash buffer. SPA b (1.0 mg/mL) and swine anti-PRV antibody IgG (1.0 mg/mL) were dispensed by biojets c attached to a commercial dispenser c on cellulose ester membrane (300 mm × 25 mm) d as the test and control lines, respectively. The test and control lines were situated 0.5 cm apart at the center of the membrane. The membrane has a nominal capillary flow time of 120 sec/4 cm and a nominal membrane thickness of 135 μm. These reagents were applied in the form of dots applied at 50 dots/mL/cm to the membrane, and then dried for 1 hr at 40°C. Sample and absorbent pads were made from nonwoven, 100% pure cellulose fiber d The sample pad was cut to 15 mm × 300 mm and saturated with a buffer (pH 8.0) containing 20 mM sodium borate, 2.0% (w/v) sucrose, 2.0% (w/v) BSA, and 0.1% (w/v) NaN3 and dried for 10 min at 72°C. The absorbent pad was cut to 20 mm × 300 mm. Then the sample pad, conjugate pad, blotted membrane, and absorbent pad were assembled on the plastic backing support board sequentially with a 1–2 mm overlap. The master card was cut to 3.0-mm wide strips using a guillotine cutter. c The strips were then sealed in a plastic bag with desiccant gel and stored at 4°C until used.
To use the immunochromatographic test strip, a serum sample was 50-fold diluted with normal saline solution first, then 100 μL diluted sample was applied to the sample pad, whereby the solution migrates toward the absorbent pad. With a positive sample, the antibody binds to the antigen conjugate forming a gold antigen–antibody complex, which binds to SPA and gives a red-colored band at the test-line region. The absence of this band suggests a negative result. Samples with ambiguous results were retested; if the test provided another ambiguous result, the sample was considered as negative. To serve as a procedural control, a red band at the control-line region always appears regardless of the presence of anti-PRV antibodies.
The specificity of the strip was determined with 265 reference serum samples, including 160 PRV-positive sera and 50 negative sera, as well as 55 positive sera against a variety of other viruses (Classical swine fever virus [CSFV], Porcine reproductive and respiratory syndrome virus [PRRSV], Porcine circovirus-2 [PCV-2], Japanese encephalitis virus [JEV], Foot-and-mouth disease virus [FMDV], and porcine parvovirus [PPV]). As shown in Figure 2, a positive result appeared with 2 visible red bands at the test and control line positions, and a negative result appeared with only 1 red band at the control line position on the strip. The results from swine serum samples are summarized in Table 1. In 160 PRV-positive samples and 50 PRV-negative samples, the strip gave 3 false-negative and 2 false-positive results, and all of the 55 other pathogens (CSF, PRRS, PC, JE, FMD, and PP) positive serum samples gave negative results, which indicated that the specificity of the strip was 98.1% (260/265).
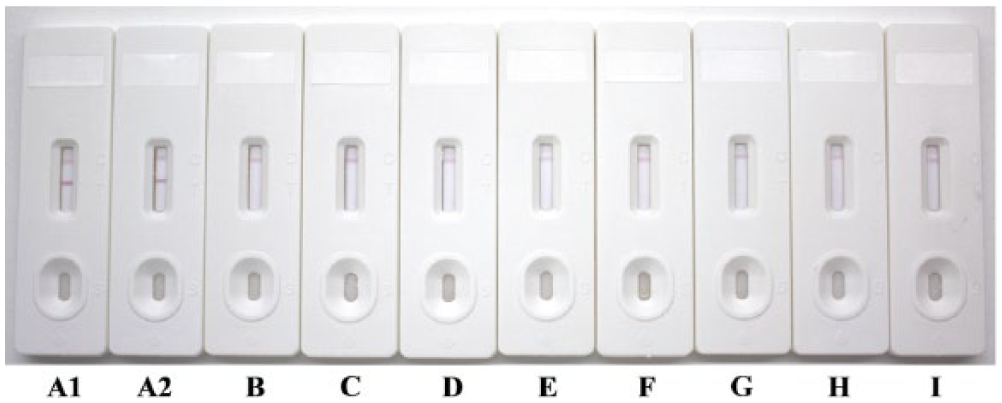

The specificity of pseudorabies virus (PRV) antibody detection by immunochromatographic strip test.
The specificity of the immunochromatographic strip with reference swine serum samples.*
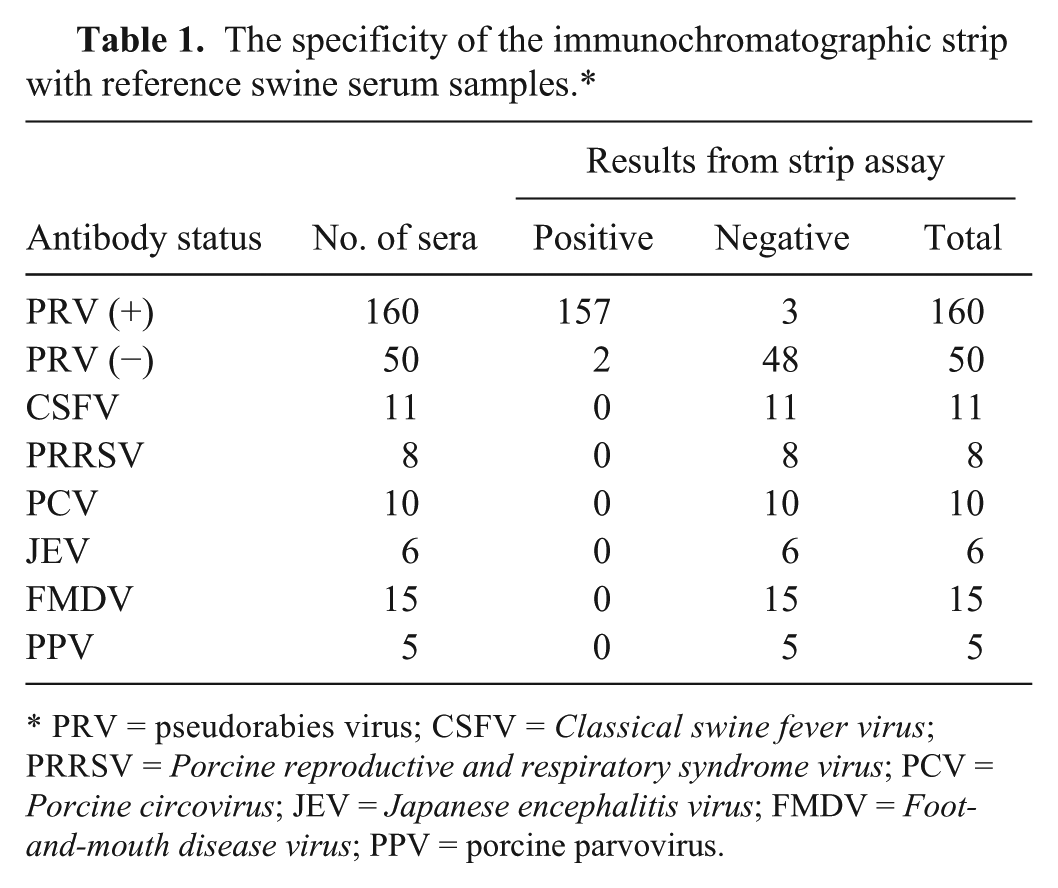
PRV = pseudorabies virus; CSFV = Classical swine fever virus; PRRSV = Porcine reproductive and respiratory syndrome virus; PCV = Porcine circovirus; JEV = Japanese encephalitis virus; FMDV = Foot-and-mouth disease virus; PPV = porcine parvovirus.
The sensitivity of the immunochromatographic strip was tested with 100 PRV gB antibody–positive pig serum samples (which were screened by a commercial PRV gB enzyme-linked immunosorbent assay [ELISA] e ). Ninety-six samples gave a positive result, which indicated that the sensitivity of this strip was 96% (96/100).
A total of 296 serum samples derived from vaccinated or infected swine and 50 sera derived from naive swine were tested with the immunochromatographic strip and commercial PRV gB ELISA, e simultaneously. The immunochromatographic test strip provided clear positive or negative results with the 296 field samples. The strip and the commercial ELISA provided similar results (Table 2). The agreement of the immunochromatographic test strip with the commercial PRV gB ELISA was 93.6% (277/296). This indicated that the 2 test methods show good correspondence.
Comparison of the immunochromatographic strip and commercial pseudorabies virus (PRV) glycoprotein B (gB) enzyme-linked immunosorbent assay (ELISA).*
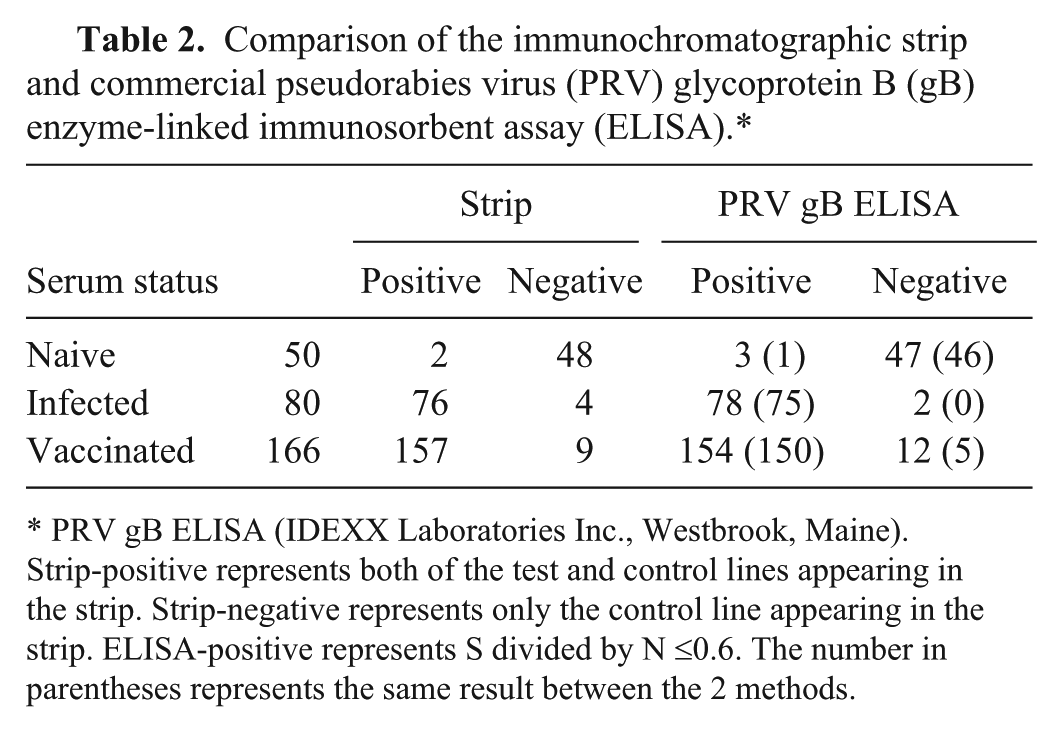
PRV gB ELISA (IDEXX Laboratories Inc., Westbrook, Maine). Strip-positive represents both of the test and control lines appearing in the strip. Strip-negative represents only the control line appearing in the strip. ELISA-positive represents S divided by N ≤0.6. The number in parentheses represents the same result between the 2 methods.
In China, the first report of a PRV outbreak occurred in the 1950s; inactivated vaccines and gE gene–deleted vaccines with virus strains Bartha-K61, HB-98, and Bucharest were imported from Hungary to China in the 1970s. From the 1990s until late 2011, >80% of pigs in China were vaccinated with the Bartha-K61 vaccine, and pseudorabies was well controlled. However, beginning in late 2011, pseudorabies has occurred on many large pig farms in animals that have been vaccinated. Newly emerging variants of PRV compromise the protection provided by current vaccines and cause death in all ages of vaccinated pigs. To circumvent such problems, both killed or modified-live PRV vaccines based on current PRV variants have been developed. An inactivated gE and gI-deleted vaccine based on PRV ZJ01 was developed, and the vaccine can provide protection to the newly emerging variant PRV challenge. A live gE-deleted PRV vaccine was also reported to provide complete protection to lethal challenge with PRV variant TJ strain. Glycoprotein B, as a main protective antigen, was necessary in the PRV vaccine.
Immunization efficiency evaluation mainly depends on serological tests. The serological tests used most commonly for PRV antibody determination are the virus neutralization test, 1 ELISA, 4 and latex agglutination test. 14 Although these assays provide accurate and sensitive detection of PRV antibodies, they require specialized equipment and technical expertise. As intact virus is involved in these tests, there is the risk of incomplete inactivation and escape of virus from laboratories. The immunochromatographic strip is a technique in which a cellulose membrane is used as the carrier and a colloidal gold–labeled antigen or antibody is used as the tracer. This technique has several advantages over traditional immunoassays, such as simplicity of the procedure, rapid operation, immediate results, low cost, and no requirements for skilled technicians or expensive equipment. Because of these characteristics, the immunochromatographic strip test is suitable for on-site detection of antibodies or antigens. 6
The gB protein overexpressed in E. coli, and, as is commonly the case, formed insoluble inclusion bodies. A large fraction of this protein could be solubilized in a denaturing buffer, thus allowing its purification by Ni-chelation chromatography. The problem of obtaining refolded protein was found to be best achieved by rapid dilution. Variations in the final yield are found depending on the combination of expression level and the efficiency of the refolding. The conditions of expression, purification, and refolding of gB protein were optimized, and the results were demonstrated by SDS-PAGE and Western blot.
The new immunochromatographic strip was optimized, and positive and negative sera were tested. The specificity and sensitivity of the strip was 98.1% and 96.0% respectively, with no cross-reaction with other pathogen antibody–positive sera such as CSF, PRRS, PC, JE, FMD, and PP. A total of 277 out of 296 samples showed the same results when compared with the commercial PRV gB ELISA kit; the rate of agreement was 93.6%. These results illustrated that the new immunochromatographic strip showed good performance and can be applied in-clinic to the evaluation of gB antibodies of PRV.
The strip test developed in the present study can be performed rapidly and easily, requiring no special equipment and skilled personnel. Thus, it has the potential for use in-field to monitor antibodies induced by PRV vaccines. The detection of PRV antibody is the basis for establishing a proper immune program and evaluating the effect of vaccination on swine. However, the strip cannot discern vaccine-induced antibodies and antibodies derived from natural exposure. Although there is a linear relationship between the extent of color of the test line and the titers of PRV antibodies, quantitative assay cannot be visualized clearly by the naked eye. Further investigations need to be focused on finding a portable strip reader to provide the end-user with options for on or off screening and quantitative results.
Footnotes
Acknowledgements
We wish to thank Drs. Weitao Xie, Yue Liang, and Guiyue Zhang for supplying serum samples.
Authors’ note
Xiujie Li and Yaning Sun contributed equally to this work.
Authors’ contributions
S Yang and G Zhang contributed to conception and design of the study, and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved. XJ Li, Y Sun, S Yang, Y Wang, J Yang, Y Liu, Q Jin, XW Li, and C Guo contributed to acquisition of data. S Yang contributed to analysis of data.
a.
E. coli BL21 (DE3), Merck KGaA, Darmstadt, Germany.
b.
SPA, Sigma-Aldrich, St. Louis, MO.
c.
Gene Genius Bio Imaging System, Quanti 3000 Biojets, XYZ Biostrip dispenser, CM-4000 cutter; Bio-Dot, Irvine, CA.
d.
HiFlow Plus cellulose ester membrane, Cellulose fiber; EMD Millipore Corp., Billerica, MA.
e.
IDEXX PRV gB ELISA kit, IDEXX Laboratories Inc., Westbrook, ME.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the grant from key program of the National Natural Science Foundation of China (c180501) and Special Fund for Agro-scientific Research in the Public Interest (201203039).
