Abstract
Toxoplasma gondii is among the most widespread parasites worldwide. Wildlife is recognized as an important reservoir and source of infection of T. gondii. The goal of the present work was to assess the performance of loop-mediated isothermal amplification (LAMP) as a diagnostic tool for T. gondii infection in the skeletal muscle and central nervous system (CNS) of free-ranging ungulates and carnivores. Fifty-seven wild animals were tested for the presence of T. gondii DNA by polymerase chain reaction (PCR) and LAMP. The use of LAMP amplification improved sensitivity in T. gondii molecular detection compared with conventional PCR on skeletal muscle (χ2 = 5.8, P < 0.05), having a lower minimum detection limit (0.1 tachyzoite) than PCR (1 tachyzoite). No significant differences existed between the detection capacities of both assays when performed on CNS. LAMP is a valid tool to improve the diagnosis of T. gondii infection in wild game meat. The technique provides a sensitive yet specific method that can be applicable to both field surveys and large-scale testing of wildlife samples.
Toxoplasma gondii is an apicomplexan parasite that is among the most common foodborne pathogens and the most prevalent infection in humans. 5 Wildlife species are often highly parasitized, 4 and the handling and consumption of raw and undercooked game meat have been identified as an infection source for human beings. 8 Infection by T. gondii can be detected through a number of different methods, from indirect identification of stage-specific immunoglobulins to DNA-based approaches such as polymerase chain reaction (PCR), quantitative real-time PCR (qPCR), and loop-mediated isothermal amplification (LAMP). Quantitative real-time PCR provides among the most sensitive diagnosis of toxoplasmosis as it is able to detect 1 fg of T. gondii DNA 11 while LAMP, targeting the same 529–base pair (bp) repeat fragment, has a minimum detection limit of 0.6 fg. 9 The minimum detection limit of a LAMP assay targeting the surface antigen 2 (SAG2) gene (0.1 T. gondii tachyzoite) is slightly higher, 10 but, due to extreme simplicity and speed, LAMP provides a valuable alternative for fast and effective diagnosis in contexts where economic resources are limited and no valid alternative are available (e.g., game meat screening in rural areas). The goal of the current work was to assess the diagnostic performance of LAMP in detecting T. gondii DNA in tissues of naturally infected free-ranging wildlife in comparison with conventional PCR.
From an existing tissue bank, samples from 30 wild animals known to be positive for T. gondii by PCR (results partially published 4 ) and 27 animals of the same species—wild boar (Sus scrofa), red fox (Vulpes vulpes), roe deer (Capreolus capreolus), chamois (Rupicapra rupicapra), and red deer (Cervus elaphus)—known to be negative by PCR were chosen. All 57 sampled animals were free-ranging wild ungulates or carnivores (n = 26 wild boar, n = 20 red fox, n = 5 roe deer, n = 3 chamois, n = 3 red deer) that had been hunted during regular hunting activities or found dead in Piedmont, Northwestern Italy. The samples were collected opportunistically and were originally submitted for passive wildlife disease surveillance. No ethical approval was necessary, and steps to ameliorate suffering were not applicable in this study. The skeletal muscle of all 57 animals was tested by both LAMP and PCR. Central nervous system (CNS; encephalon homogenate) was also collected and tested from 43 of these animals. Genomic (g)DNA was extracted from 25 mg of tissue, using a commercial kit a according to manufacturer’s instructions.
The PCR protocol targeted a highly reiterated 529-bp fragment of T. gondii DNA. 6 The PCR reaction was carried out as specified elsewhere. 4
The LAMP assay used 2 primer pairs targeting the SAG2 gene. 10 The LAMP amplification was optimized in a final volume of 25 µL, with 2 µL of the extracted DNA, 40 pmol of the FIP and BIP primers, 5 pmol of the B3 and F3 primers, 8 U of Bst polymerase b in 2.5 µL of buffer (20 mM Tris–HCl [pH 8.8], 10 mM KCl, 8 mM MgSO4, 10 mM (NH4)2SO4, 0.1% Tween 20), 0.8 M betaine, c and 1.4 mM deoxynucleotide mix. d The LAMP reaction was incubated for 60 min at 65°C and then inactivated at 80°C for 10 min. The resulting amplicons were visualized in 1.5% agar gel using a commercial nucleic acid stain. e The LAMP amplicons were also visualized directly in the reaction tube by adding the same diluted fluorescent detection reagent e to 2 µL of LAMP product.
The sensitivity of both the PCR and LAMP assays was determined on 10-fold serial dilutions of gDNA of T. gondii tachyzoites (10 ng to 1 fg of total gDNA). The specificity of both PCR and LAMP assays was tested on heterogeneous DNA samples of Neospora caninum, Babesia spp., Theileria spp., and Leishmania infantum. These nontarget samples were obtained by direct DNA extraction from parasite cultures and identified by species-specific PCR and sequencing. The GenBank accession numbers are as follows: EF202082 N. caninum; KF773737 Babesia spp.; KF773741 Theileria spp.; and HF937257 L. infantum. Sterile water was used as negative control, and T. gondii gDNA was used as positive control. The LAMP and PCR assays were carried out under sterile conditions, in order to avoid possible contamination. Positive LAMP products were sequenced at a commercial facility, f and the resulting sequences were compared with those available in GenBank, to confirm assay specificity.
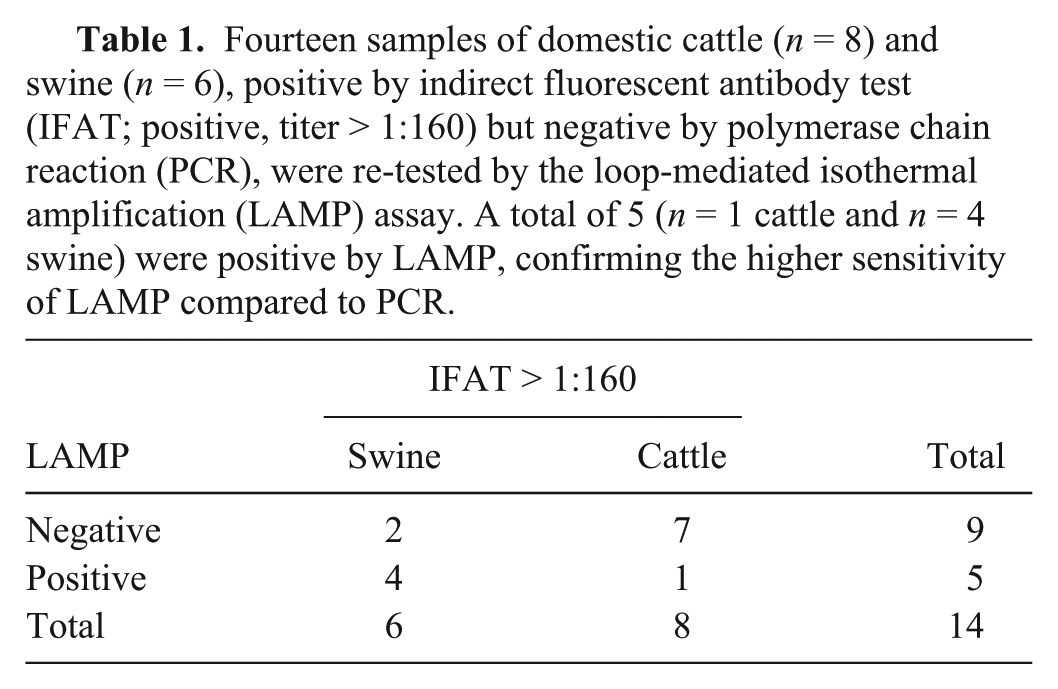
An indirect assessment of the specificity of LAMP compared with PCR was done using 14 samples (skeletal muscle) of domestic cattle (n = 8) and swine (n = 6) with discordant serological and biomolecular results. All samples were seropositive to T. gondii by indirect fluorescent antibody test (IFAT) but negative by conventional PCR (Table 1).
Fourteen samples of domestic cattle (n = 8) and swine (n = 6), positive by indirect fluorescent antibody test (IFAT; positive, titer > 1:160) but negative by polymerase chain reaction (PCR), were re-tested by the loop-mediated isothermal amplification (LAMP) assay. A total of 5 (n = 1 cattle and n = 4 swine) were positive by LAMP, confirming the higher sensitivity of LAMP compared to PCR.

McNemar test for paired data was used to compare the PCR and LAMP assays. The performance of both tests was compared separately for skeletal muscle and CNS. Statistical analysis was conducted using R software, version 3.0.1. 12
The LAMP assay was able to detect a minimum amount of T. gondii DNA equal to 10 fg (Fig. 1). This corresponds to ~0.1 tachyzoite, as the 80–megabase pair genome of T. gondii is equal to ~80 fg. 13 No cross-reaction was observed with conventional PCR or LAMP with any of the heterogeneous DNA samples used.
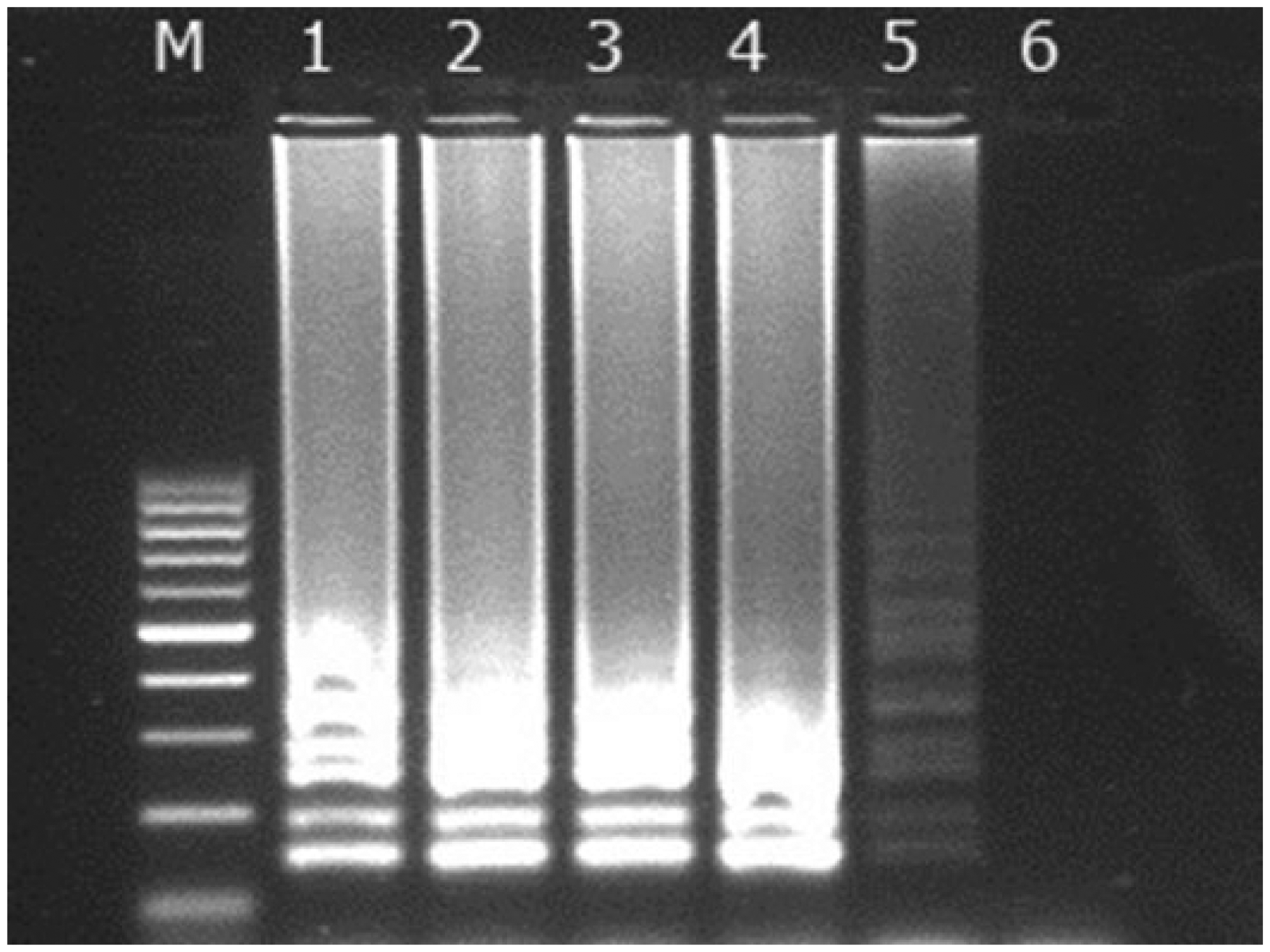
Lanes 1–6: result of loop-mediated isothermal amplification from 10 ng, 100 pg, 1 pg, 100 fg, 10 fg, and 1 fg of Toxoplasma gondii DNA; lane M: 100–1,000 base pair DNA ladder.
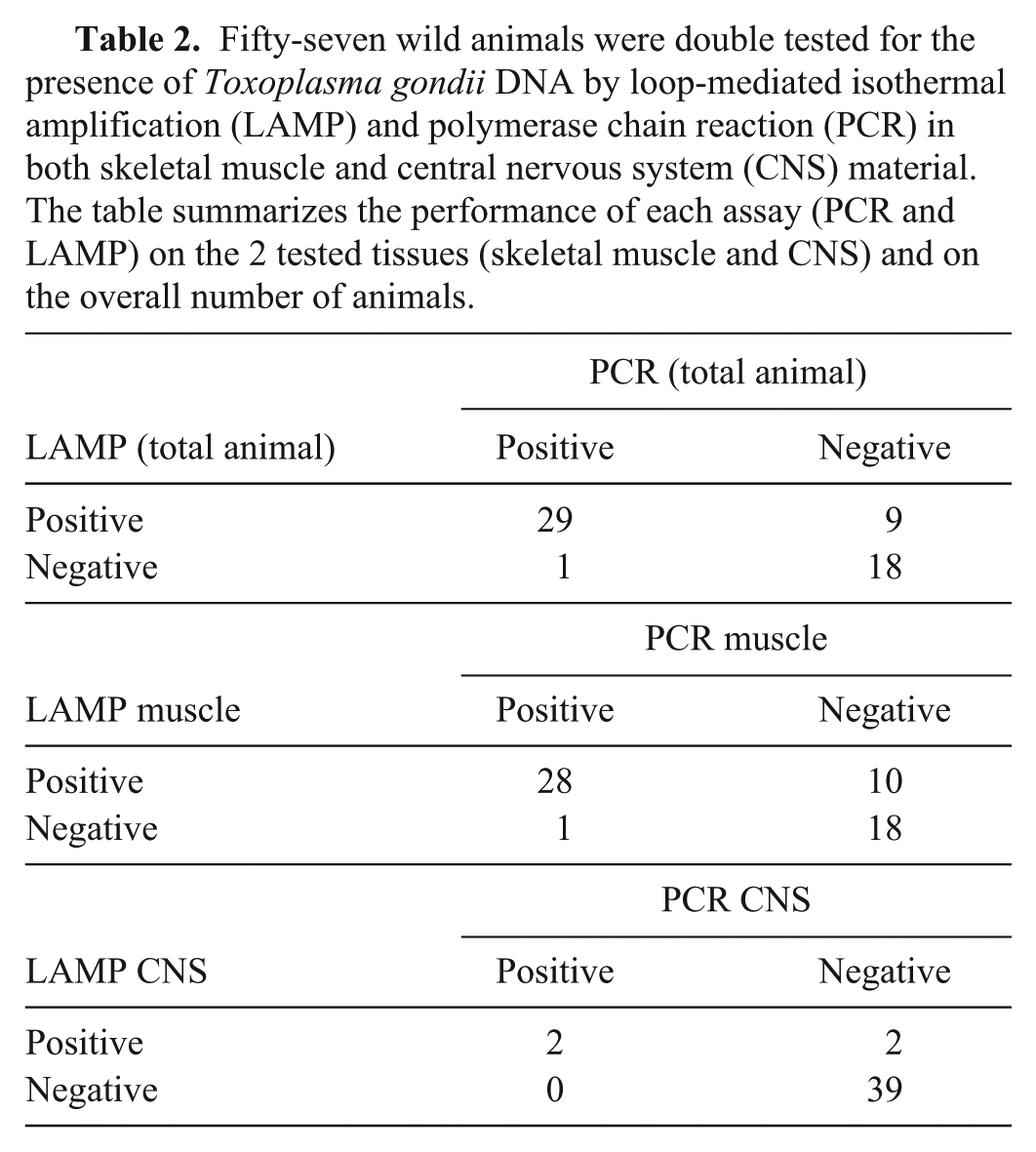
The LAMP and PCR assays gave significantly different results when used on wild animal samples. The LAMP assay proved to be more sensitive than the PCR assay (χ2 = 6.4, P < 0.05) as 9 samples negative by PCR tested positive by LAMP (Table 2). The diagnostic performances of the 2 assays were also separately assessed on skeletal muscle and CNS homogenate. The LAMP assay was significantly more sensitive compared to the PCR assay when performed on skeletal muscle (χ2 = 5.8, P < 0.05), while there were no significant differences when both assays were performed on CNS (P > 0.05; Table 2). Domestic cattle and swine with discordant serological (IFAT positive) and conventional PCR (negative) results as retested by LAMP confirmed the higher sensitivity of LAMP: of the 14 livestock samples, all negative by conventional PCR and positive by serology, 5 were positive in the LAMP assay. Fifteen randomly chosen LAMP-positive samples (n = 4 swine; n = 1 cattle; n = 10 wild animals) were sequenced to confirm the actual presence of T. gondii DNA. Resulting sequences were compared to those available in GenBank, and all showed a 100% homology to T. gondii SAG2 gene (accession AF357582).
Fifty-seven wild animals were double tested for the presence of Toxoplasma gondii DNA by loop-mediated isothermal amplification (LAMP) and polymerase chain reaction (PCR) in both skeletal muscle and central nervous system (CNS) material. The table summarizes the performance of each assay (PCR and LAMP) on the 2 tested tissues (skeletal muscle and CNS) and on the overall number of animals.
Consumption and trade of wild and farmed game meat increased across Europe together with the number of free-living wild ungulates, especially roe deer and wild boar. 1 Epidemiological studies on several outbreaks have identified the handling and consumption of raw or undercooked game as a source of toxoplasmosis. 2 Serological and molecular studies showed a level of infection of hunted wildlife ranging from 2% to 68%,7,14 and the European Food Safety Authority estimated that approximately half of the game meat produced in Europe is seropositive for T. gondii. 3 The need for a rapid and effective method for a sensitive detection of T. gondii in human-consumed meat has long been recognized. Some methods, such as qPCR are among the most accurate, but require highly sophisticated equipment and resources. The LAMP assay has been proposed as an alternative method to improve diagnostic sensitivity (minimum detection limit ranging from 0.6 to 10 fg).9,11 An added advantage of LAMP is that DNA amplification can be easily detected by visual inspection of the test tubes, alleviating the need for gel electrophoresis, and thus, making the method suitable for field tests. 11 Despite the high sensitivity showed by the LAMP assay, specificity is assured by the simultaneous use of 4 specific primers and was also demonstrated by the increased concordance between IFAT and LAMP as reported by our data on domestic livestock and by sequencing results.
The current study uses LAMP to assess T. gondii infection in wild game tissues. As already reported for PCR,4,9 LAMP assay carried out on the skeletal muscle of both wild and domestic animals is more sensitive than the same assay performed on CNS. An effective diagnosis of T. gondii infection in wild animals is needed to improve the hygienic standards of hunted and farmed game meat. The data presented in this article confirms that the use of LAMP assay can improve the diagnosis of T. gondii infection in wild herbivores and carnivores.
Footnotes
Acknowledgements
We acknowledge the valued help of Drs. Isabella Nicola and Fabio Bosio who processed some of the samples used in the study, as well as Drs. Davide Grande and Simona Bruno who performed postmortem examination of the studied wildlife.
Authors’ contributions
S Zanet contributed to conception of the study, contributed to analysis and interpretation of data, and drafted the manuscript. E Ferroglio contributed to conception and design of the study. A Trisciuoglio contributed to acquisition and analysis of data. F Chiesa and DM Nucera contributed to analysis of data. G Marello contributed to analysis and interpretation of data. M Bergallo and MS Gennero contributed to interpretation of data. All authors critically revised the manuscript; gave final approval; and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
GenElute mammalian genomic DNA miniprep kit, Sigma-Aldrich, St. Louis, MO.
b.
Large fragment DNA polymerase, New England Biolabs, Ipswich, MA.
c.
Betaine, Sigma-Aldrich, St. Louis, MO.
d.
Deoxynucleotide mix 10 mM, Sigma-Aldrich, St. Louis, MO.
e.
GelRed nucleic acid gel stain, VWR, Milan, Italy.
f.
Macrogen Inc., Amsterdam, The Netherlands.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was partially supported by funds from the Piedmont Regional Administration (“Convenzione tra la Regione Piemonte e il Dip. di Scienze Veterinarie dell’Università degli Studi di Torino per la razionalizzazione ed integrazione delle attività di raccolta e smaltimento degli animali selvatici morti o oggetto di interventi di contenimento”) and APHAEA EU project (“Harmonised approaches in monitoring wildlife population health, and ecology and abundance”).
