Abstract
Molecular methods for the detection of Schmallenberg virus (SBV) RNA were rapidly developed after the emergence of this novel orthobunyavirus in Europe. The SBV epizootic wave has declined, but infectious SBV in SBV RNA–positive semen remains a possible risk for the distribution of SBV. However, the abilities of SBV molecular detection methods used at European laboratories have not yet been assessed, to our knowledge. The performances of extraction and real-time reverse transcription polymerase chain reaction (RT-qPCR) methods used at 27 German and 17 other European laboratories for SBV RNA detection in the matrices of whole blood, serum, tissue homogenate, RNA eluates, and bovine semen were evaluated in 2 interlaboratory trials with special emphasis on semen extraction methods. For reliable detection of viral genome in bovine semen samples, highly effective extraction methods are essential to cope with the potential inhibitory effects of semen components on PCR results. All methods used by the 44 laboratories were sufficiently robust to detect SBV RNA with high diagnostic sensitivity (100%) and specificity (95.8%) in all matrices, except semen. The trials demonstrated that the published recommended semen extraction methods (Hoffmann et al. 2013) and a combination of TRIzol LS with an alternative extraction kit have a considerably higher diagnostic sensitivity to detect SBV RNA in semen up to a detection limit of Cq ≤35 compared to other extraction methods used. A thorough validation of extraction methods with standardized semen batches is essential before their use for SBV RNA detection in bovine semen.
Introduction
The unexpected emergence of the orthobunyavirus Schmallenberg virus (SBV; Simbu serogroup) in Europe in 2011 confronted veterinary laboratorians with the challenge of swiftly developing molecular assays for robust and sensitive detection and monitoring of SBV infection in ruminant livestock.1,4–6,10,14,16,23,30 SBV has infected ruminants at an impressive pace, resulting in a seroprevalence of up to 100% in some SBV-affected regions and extensive spread throughout Europe as far as Turkey in Asia.2,8,17,22,28,29 Biting midges of the Culicoides genus have been determined as the major source of SBV infection.6,7,19 Adult ruminants infected with SBV show no or mild clinical signs. In contrast, abortion and malformation may occur in the offspring of dams infected for the first time with SBV during the first third of gestation. 22 The detection of infectious SBV in semen from naturally or experimentally SBV-infected fertile bulls suggested that natural and artificial insemination of dams might be an additional route of infection.16,22,24 The SBV epizootic wave has declined (World Trade Organization [WTO], 2012, http://ec.europa.eu/food/animal/diseases/schmallenberg_virus/docs/gen_1161_sbv_en.pdf), 12 but insemination of SBV-naive dams with cryopreserved semen contaminated with SBV is assumed to be a possible risk for distribution of SBV to SBV-free regions.21,22,24 The WTO removed restrictions on trade of live ruminants and their products from SBV-affected regions in July 2012 (WTO, 2012, http://ec.europa.eu/food/animal/diseases/schmallenberg_virus/docs/gen_1161_sbv_en.pdf). Nevertheless, several SBV-free countries implemented or sustained restrictions on the import of ruminant semen 16 (ProMED-mail, 2013, http://www.promedmail.org/direct.php?id=20130131.1524123). To ensure that traded semen is free of SBV, sensitive real-time reverse transcription polymerase chain reaction (RT-qPCR) testing of the semen matrix from SBV-infected bulls is required. However, reliable detection of SBV genome in bovine semen demands highly effective extraction methods to cope with the potential inhibitory effects of semen components on the PCR results.15,16 The high variability of excretion patterns also complicates the safe detection of SBV RNA excretion in semen from an SBV-infected bull.16,22,24
A limited serological ring trial indicated that the enzyme-linked immunosorbent assays and virus neutralization tests used in European laboratories are suitable for reliable detection of SBV antibodies in ruminant blood. 24 However, the performance characteristics of molecular diagnostic methods for the detection of SBV RNA in European laboratories have not yet been assessed, to our knowledge.
Schmallenberg virus contains a tripartite single-stranded, negative-sense genome structure characteristic of viruses belonging to the Orthobunyavirus genus. The 3 segments are named according to their size.9,10 RT-qPCR assays were developed for the detection of any of the small (S), medium (M), and large (L) segments.a,b,4,6,10
The Friedrich-Loeffler-Institut (FLI; Insel Riems, Germany) organized and arranged shipments for 2 interlaboratory method trials to German National and European laboratories in 2013. The objectives of the trials included the evaluation of the efficacy of 1) routinely used in-house extraction and RT-qPCR methods for SBV RNA detection in various matrices and of 2) previously published15,16 versus alternative methods used for extraction of SBV RNA from bovine semen with respect to the reproducibility of the RT-qPCR results.
Material and methods
Study design
The efficacy of methods used for SBV RNA extraction and detection in the matrices of whole blood, RNA eluate, serum, tissue homogenate, and bovine semen used at 44 laboratories from 13 European countries (Austria, Belgium, Croatia, Denmark, France, Germany, Ireland, Italy, The Netherlands, Spain, Sweden, Switzerland, and the United Kingdom) was evaluated in 2 method trials. A total of 27 German National (D) and 16 other European (EU) laboratories participated in the first trial. In the second trial, a larger number of samples for each matrix was provided to 5 European laboratories of a specialist SBV consortium (C); 4 laboratories (C1–3, and C5) out of the 5 participated in both trials.
In each of the trials, a set of defined samples was provided by the organizer of the trial (FLI). The samples originated from domestic ruminants that were infected during the SBV epizootic in Germany between 2011 and 2013. The number of specimens by matrix, their corresponding reference quantification cycle (Cq) values, and sample labels provided in the first method trial are given in Table 1. For the second trial, 16 RNA eluates (Cq 19.2–35.7, no Cq), 16 serum (Cq 22.6–32.5, no Cq), 12 tissue homogenates (Cq 17.7–34.8), and 6 semen batches (Table 1) were provided.
Sample labels, matrix, origin, and reference quantification cycle (Cq) values as determined with Schmallenberg virus (SBV)-S3 real-time reverse transcription polymerase chain reaction* by the organizer of the method trial. All samples were collected during the SBV epizootic in Germany, 2011–2013.
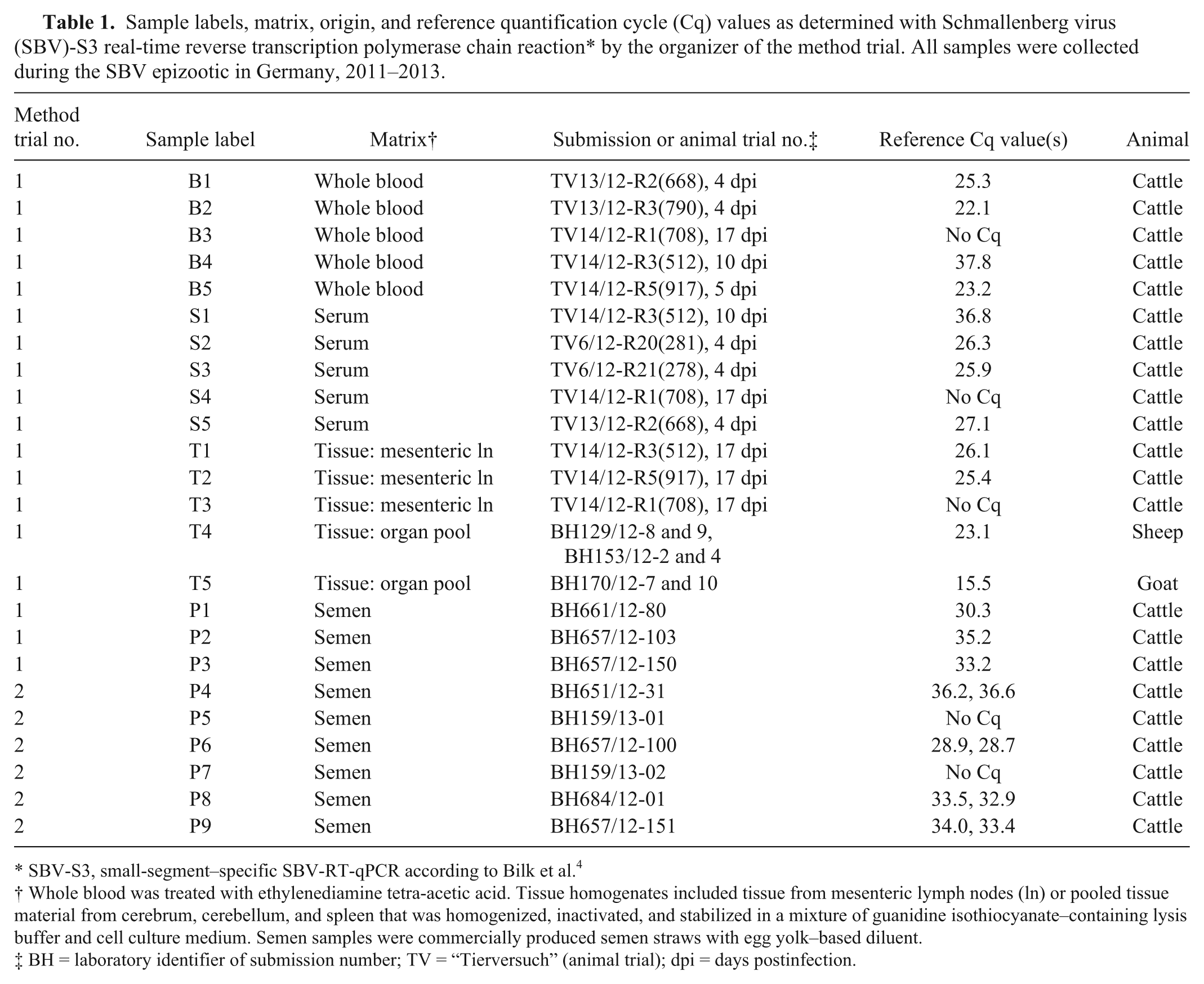
SBV-S3, small-segment–specific SBV-RT-qPCR according to Bilk et al. 4
Whole blood was treated with ethylenediamine tetra-acetic acid. Tissue homogenates included tissue from mesenteric lymph nodes (ln) or pooled tissue material from cerebrum, cerebellum, and spleen that was homogenized, inactivated, and stabilized in a mixture of guanidine isothiocyanate–containing lysis buffer and cell culture medium. Semen samples were commercially produced semen straws with egg yolk–based diluent.
BH = laboratory identifier of submission number; TV = “Tierversuch” (animal trial); dpi = days postinfection.
To determine reference Cq values for whole blood, serum, and tissue homogenates, the specimens were extracted c according to the manufacturer’s recommendations. Semen samples were extracted with a method that showed the highest diagnostic sensitivity (DSe) for SBV RNA detection in bovine semen in a previous study of extraction methods.15,16 Briefly, semen was lysed, d and viral RNA was subsequently purified with a magnetic bead–based commercial extraction kit without lysis buffer e using automated extraction. f For all samples, SBV RNA was amplified using a commercial PCR kit g and S segment–specifica,4 RT-qPCR with a real-time thermocycler h (Supplementary Tables 1, 2).4,15,16 To confirm the reference Cq values, each specimen was tested in replicate. For semen samples, at least 2 biological and 2 technical replicates (repeated RT-qPCR analysis of each biological replicate) were tested.
In the first trial, extraction methods and PCR assays recommended by the trial organizer included in-house RNA extraction and PCR protocols for all matrices, except semen. For RNA extraction from semen, a highly sensitive method described previously15,16 (lysed d with or without purification e without lysis buffer) used in an automated f or manual magnetic bead separator was recommended. The laboratories were asked to report 1 result for each of the provided specimens. In the second trial, RNA extraction from serum and tissue homogenates with a commercial RNA kit, c RNA extraction from semen with an in-house method, and PCR testing of all specimens with RT-qPCR a were recommended as alternative methods in addition to the recommended methods given in the first trial. The participants were asked to provide 2 PCR results (technical duplicates) for all matrices, except semen. For semen samples, PCR results of 2 biological and 2 technical replicates were required. All participants received straws of additional semen batches as well as the corresponding reference Cq values in the first (Cq 28.1–31.3) and second method trial (Cq 30.2–36.4) to allow validation of the recommended and/or in-house RNA extraction methods commonly used at their laboratory before the analysis of the specimens provided for the trial.
If more than 2 Cq values for each specimen were provided, means of Cq values determined by the same extraction method were calculated and processed for graphical and statistical analysis. The efficacy of different methods was comparatively analyzed with regard to PCR results. The DSe and diagnostic specificity (DSp) were determined according to a previous study 20 and the OIE.31,32 Of the 6 specimens provided in the first (S1, B4, P2) and second (R7, T5, P1) method trial (Table 1), 2 (R7 and T5) contained SBV RNA loads at the detection limit (Cq ≥ 35). For specimens confirmed as SBV RNA–positive by the trial organizer, doubtful results were considered “positive” in our study, because doubtful results lead to repeated testing of a sample in the primary or another laboratory, or in virological and serological analyses of follow-up sample(s) from the donor animal.
Statistical analysis
Statistical analysis was conducted for PCR results determined for semen batches in the first trial. Wilcoxon rank sum tests were used to determine whether the mean ranks of Cq values of all respectively single semen batches obtained by extraction with the recommended methods (lysed d with or without purification e ) differ significantly from those obtained by other extraction methods. The alpha level was adjusted by continuity correction.
For all statistical analyses, a Cq value of 45 was given to samples identified as negative, and an alpha level of P < 0.05 was considered significant. Data processing and statistical analyses were conducted with R for statistical computing (http://www.r-project.org/; http://stat.ethz.ch/R-manual/R-patched/library/stats/html/wilcox.test.html; http://www.rstudio.org).
Results
RT-qPCR results for whole blood, serum, RNA eluates, and tissue homogenates
For the matrices of whole blood, serum, RNA eluate, and tissue homogenate, results were provided by all participants of the first (n = 43) and second (n = 5) trial, respectively. A considerable number of different extraction methods by matrix (n = 14–16) were used for SBV RNA extraction from whole blood, serum, and tissue homogenate, but the highest proportion of these specimens was extracted with a commercial kit c (40.82%, 60/147; Supplementary Table 3). All extraction and RT-qPCR methods (100%) were sufficiently robust to detect SBV RNA in each of the specimens.
In the first trial, SBV RNA was detected in at least 1 of 2 or in both borderline samples (S1 and B4) by 72% (31/43) and 49% (21/43) of the laboratories, respectively (Fig. 1). In the second trial, SBV RNA was detected in the 2 borderline samples T5 and R7 (Cq 34.8, respectively 35.7) by all 5 participants; although not every RT-qPCR method yielded a positive result (data not shown). Of the SBV RNA–negative samples, 95.8% (181/189 analyses) were correctly determined as negative for SBV RNA by 40 of 44 laboratories (Fig. 1). PCR protocols (Supplementary Tables 1, 2; data only shown for methods used for SBV RNA detection in semen) and temperature profiles (data not shown) corresponded with those used for SBV RNA detection in bovine semen.
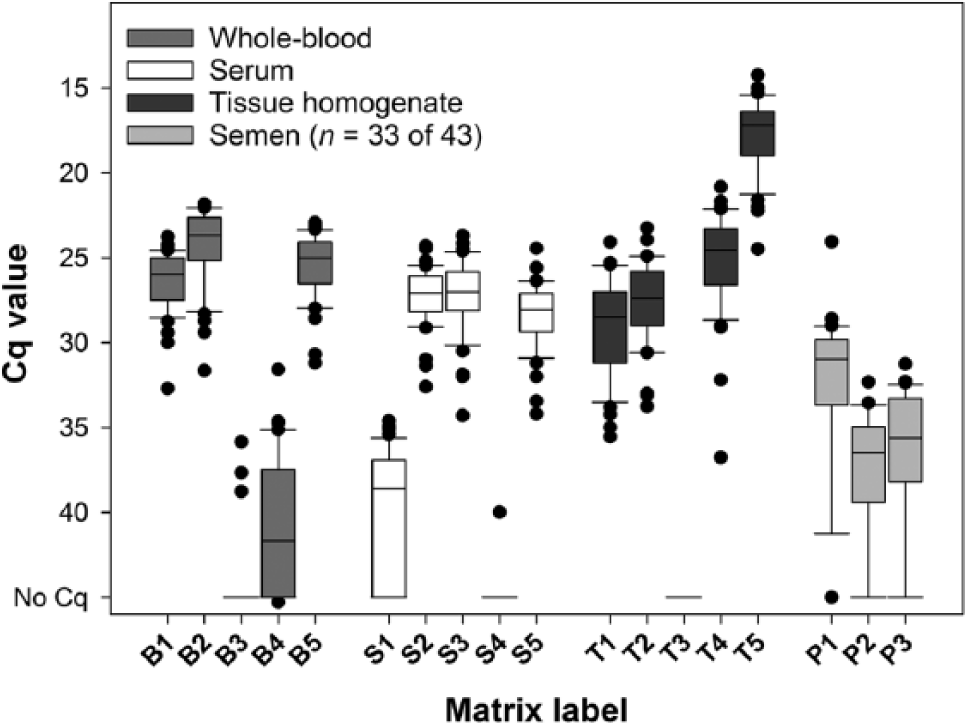
Polymerase chain reaction (PCR) results of 27 German and 16 other European laboratories that participated in the first method trial to evaluate the efficacy of methods for the detection of Schmallenberg virus (SBV) RNA by matrix. Semen batches (P1–P3) were tested by 33 of 43 laboratories. In the matrices of whole blood (B), serum (S), and tissue homogenate (T), SBV RNA was detected with all PCR assays used by the 43 laboratories with similar quantification cycle (Cq) values, except for the 2 samples with Cq values at the detection limit (Cq ≥36.8). All except 2 of the 33 laboratories that tested semen batches detected SBV RNA in bovine semen. The boxes indicate the lower and upper quartiles and the median. The ends of the whiskers mark the 10th and 90th percentiles. Individual outliers outside of that range were plotted as dots.
SBV RNA extraction from bovine semen
In the first trial, the 3 semen batches P1–P3 were tested for SBV RNA by 18 of the 27 German and 15 of the 16 European laboratories. SBV RNA was detected in bovine semen by all except 2 (EU6 and EU14) of the laboratories (93.9% of 33; Supplementary Tables 1, 4). Four additional laboratories had difficulty detecting SBV RNA in 1 or both semen batches with reference Cq values ≥33 (P2 and P3; Fig. 1; Supplementary Tables 2, 4).
A total of 20 different combinations of methods were used for SBV RNA extraction from semen. One-third of the laboratories extracted SBV RNA from semen with the recommended methods (TRIzol LS d with or without purification with MagAttract Virus kit e without lysis buffer; 10/33), with a combination of TRIzol LS d and an extraction kit other than the MagAttract Virus kit e (12/33) or with other TRIzol preparations and extraction kits (12/33), respectively (Table 2). One European laboratory (EU14) analyzed the 3 semen batches twice using 2 different extraction methods (Supplementary Table 1). The highest DSe (100%) for SBV RNA detection in bovine semen was found for the recommended protocol compared to other extraction methods. A combination of TRIzol LS d with any other extraction kit also revealed a high DSe (97%, 35/36), whereas the use of other TRIzol preparations with/or other kits revealed a considerably lower proportion of positive results (Table 2). Extraction with the recommended protocol compared with other methods revealed statistically significant differences between means of Cq values analyzing the 3 batches together (P1–P3) and separately (P1 and P2), respectively, except for P3 (Fig. 2).
Diagnostic sensitivity for Schmallenberg virus (SBV) RNA detection in 3 SBV RNA–positive bovine semen batches (reference quantification cycle values 30.3–35.2) by extraction procedures used by 18 German and 15 other European laboratories that participated in the first method trial. Supplementary Table 3 provides details on the used extraction kits and polymerase chain reaction protocols.*
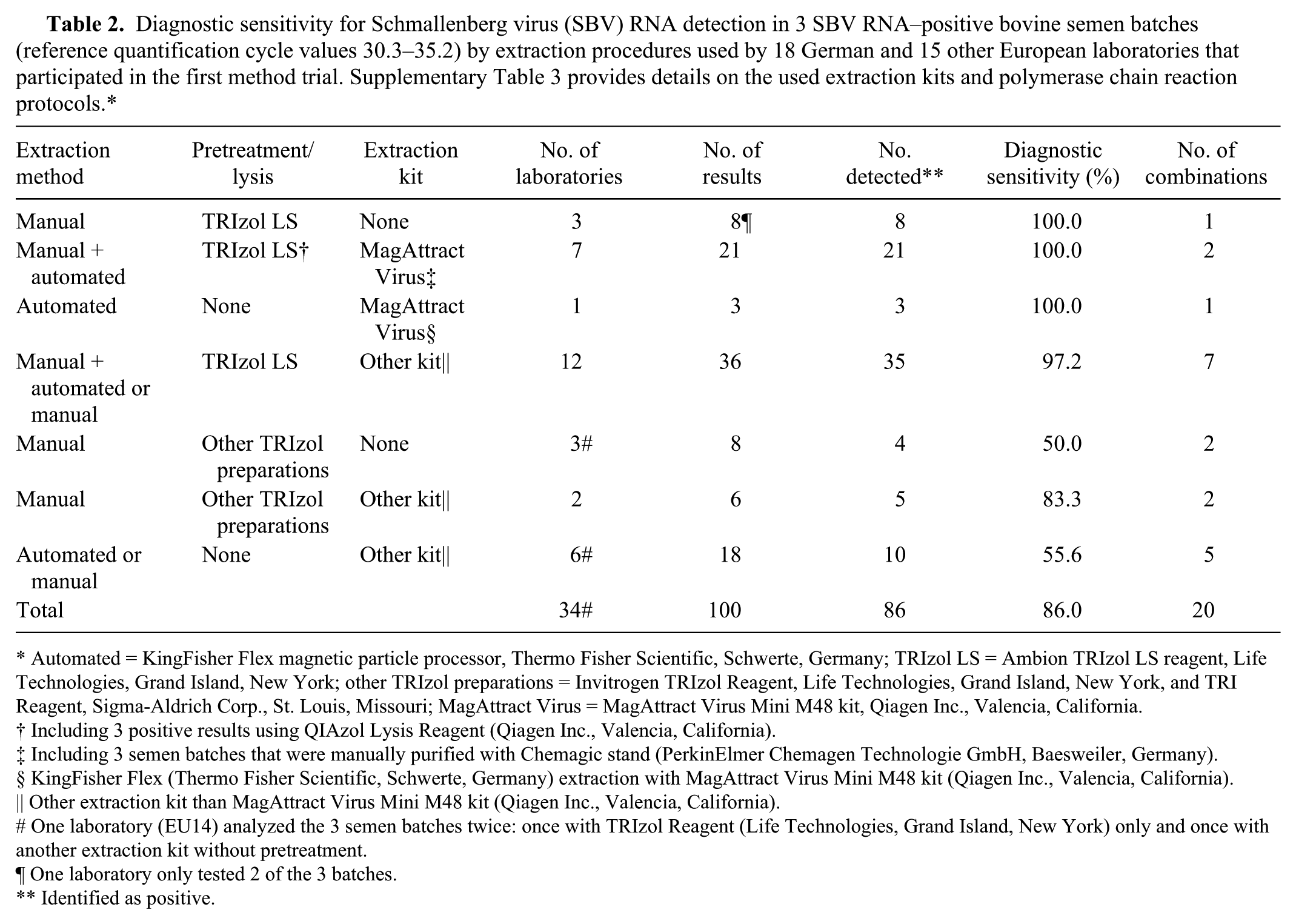
Automated = KingFisher Flex magnetic particle processor, Thermo Fisher Scientific, Schwerte, Germany; TRIzol LS = Ambion TRIzol LS reagent, Life Technologies, Grand Island, New York; other TRIzol preparations = Invitrogen TRIzol Reagent, Life Technologies, Grand Island, New York, and TRI Reagent, Sigma-Aldrich Corp., St. Louis, Missouri; MagAttract Virus = MagAttract Virus Mini M48 kit, Qiagen Inc., Valencia, California.
Including 3 positive results using QIAzol Lysis Reagent (Qiagen Inc., Valencia, California).
Including 3 semen batches that were manually purified with Chemagic stand (PerkinElmer Chemagen Technologie GmbH, Baesweiler, Germany).
KingFisher Flex (Thermo Fisher Scientific, Schwerte, Germany) extraction with MagAttract Virus Mini M48 kit (Qiagen Inc., Valencia, California).
Other extraction kit than MagAttract Virus Mini M48 kit (Qiagen Inc., Valencia, California).
One laboratory (EU14) analyzed the 3 semen batches twice: once with TRIzol Reagent (Life Technologies, Grand Island, New York) only and once with another extraction kit without pretreatment.
One laboratory only tested 2 of the 3 batches.
Identified as positive.
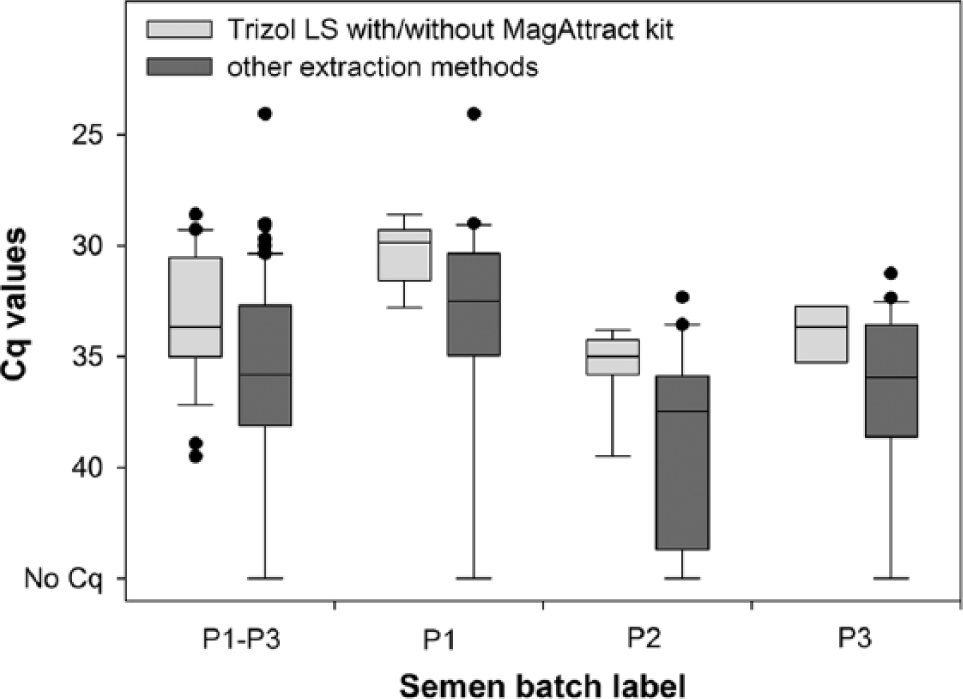
Polymerase chain reaction results of 18 German and 15 other European laboratories that participated in the first method trial to evaluate the efficacy of methods for the detection of Schmallenberg virus (SBV) RNA in bovine semen (reference quantification cycle [Cq] values of P1, P2, and P3: 30.3, 35.2, respectively, 33.2) by extraction method. The means of Cq values of all semen batches (P1–P3) and of P1 and P2 obtained by the recommended extraction methods (TRIzol LS Reagent [Life Technologies, Grand Island, NY] with/without MagAttract Virus Mini M48 kit [Qiagen Inc., Valencia, CA]; SBV RNA detected in 100.0% of 29 analyses; light gray) statistically significantly differed from the mean of Cq values obtained by other extraction methods (SBV RNA detected in 80.3% of 71 analyses; dark gray; see also Table 2). Explanation of the boxplot design is given in Figure 1.
An association between the RNA purification matrices (magnetic bead or silica membrane) and the outcome of PCR results could not be reliably evaluated because for these combined extraction methods 75% (63/84 analyzed in 21/28 laboratories) of the semen samples were pretreated with a TRIzol-based reagent; a few laboratories did not pretreat the samples with any TRIzol-based reagent before the extraction with a membrane-based (2/28) or a magnetic bead–based (5/28) method. The exclusive membrane-based extraction revealed positive results for the 6 semen samples, while of the semen samples extracted with magnetic beads only, 47% (7/15) were found to be SBV RNA positive. For extraction methods that included a pretreatment with TRIzol LS, d all except 1 semen sample were identified as positive (Table 2).
In the second method trial, 6 different extraction methods were used. The highest proportion of false-negative results (7/10) was found for the semen batch (P4) with the highest Cq value (reference Cq 36.4). This batch also tested negative for SBV RNA by 2 laboratories that used the recommended extraction methods. The other 3 SBV RNA–positive semen batches (P6, P8, P9) were identified as positive by all laboratories with all except 1 of the 6 extraction methods (Supplementary Tables 2, 4). The 2 semen batches negative for SBV RNA (P5 and P7) were identified as SBV RNA–negative by all 5 SBV consortium laboratories (DSp 100%).
SBV RNA detection in semen
In the first trial, >75% (26/33) of the laboratories used the PCR kit g (15/33) and/or the RT-PCR kit i (12/33) for the detection of SBV RNA in semen. The remaining 7 laboratories used 6 other PCR kits. In the second trial, all 5 laboratories used either of the 2 PCR kits,g,i and 3 laboratories also used an additional kit.
Any of the S segment–specific RT-qPCR assaysa,i were used for SBV RNA quantification by all except 2 (EU12 and EU14) of the 44 laboratories (95%). The latter 2 laboratories used another commercial S segment b – and an L segment–specific RT-qPCR assay, respectively, after extraction with methods distinct from the recommended extraction protocol. Details of the PCR kits and assays used are given in Supplementary Tables 1 and 2. One or none of the 3 semen batches (P1–P3) were detected as SBV RNA–positive by the 2 laboratories, but SBV RNA was detected in all other matrices using the same RT-qPCR assays after extraction with the same or different methods (data not shown).
Discussion
Molecular methods for the detection of SBV RNA in blood, tissue, and semen were swiftly established after the emergence of SBV in Europe.1,4–6,10,14,16,23,30 In order to improve overall test performance and to establish confidence in the reliability of laboratory test procedures, interlaboratory testing is recommended.3,11 Therefore, 2 method trials were conducted to evaluate the efficacies of extraction and RT-qPCR methods used at 27 German and 17 other European laboratories in 13 different countries to reproduce PCR results predetermined by the trial organizer.
SBV RNA detection in blood of ruminant livestock allows monitoring of new acute cases in SBV-affected as well as in SBV-free regions. Adult ruminants have short-term RNAemia during the first week after SBV infection, followed by seroconversion between 4 days and 3 weeks after a positive PCR result.14,21,26,29 The presence of SBV RNA in spleen and mesenteric lymph nodes for up to several weeks is a common finding in most SBV-infected ruminants after RNAemia has ceased,24,27 whereas in fetuses the detection of SBV antibodies in precolostral blood or of SBV RNA in cerebrum, brainstem, amniotic fluid, and in spinal and umbilical cords was found suitable for the confirmation of a previous SBV infection 4 (World Organization for Animal Health (OIE), 2013, http://www.oie.int/fileadmin/Home/eng/Our_scientific_expertise/docs/pdf/A_Schmallenberg_virus.pdf). Therefore, whole blood, serum, and tissue homogenates from mesenteric lymph nodes, spleen, and different parts of the brain were chosen for the 2 independent trials (Table 1). The detection of infectious SBV in bovine semen by subcutaneous injection of cattle and mice with SBV RNA–positive semen and the high variability patterns of SBV RNA excretion in semen after SBV infection of bulls demands reliable methods for safe detection of SBV RNA in semen from SBV-infected bulls, although the possibility of venereal SBV transmission remains to be further investigated.16,22,24
All extraction and RT-qPCR methods used by the 44 participating laboratories (100%) were sufficiently robust to reliably detect SBV RNA in the matrices of whole blood, RNA eluate, serum, and tissue homogenate, although a marked variety of extraction and PCR methods had been used (Table 2). More than two-thirds of the 44 laboratories also detected SBV RNA in at least 1 of the samples with a Cq value at the detection limit (Cq ≥35), whereas the proportion of SBV RNA–negative samples erroneously determined as positive or doubtful was low (DSp 95.8%).
In contrast, considerable differences were found between SBV RNA RT-qPCR results obtained from semen samples (Fig. 1). The 2 trials confirmed that the previously published and recommended extraction methodsd–f,15,16 have a significantly higher DSe (100%; Table 2; Fig. 2) in detecting SBV RNA in bovine semen with Cq values below the detection limit of Cq ≤35 (DSe of 92.7% for all semen samples analyzed in the 2 trials) compared with other extraction methods—independent from the conducting laboratory. Interestingly, a similarly high proportion of positive results was obtained by a combination of TRIzol LS d with any other than the recommended extraction method.e,f In contrast, the use of other TRIzol preparationsj–l without or in combination with any other extraction method revealed a considerably lower proportion of true-positive results (Table 2). The use of TRIzol LS d only or in combination with any extraction kit considerably improved SBV RNA extraction from bovine semen and, consequently, the PCR results. TRIzol j was most frequently used as an alternative to TRIzol LS. d The 2 reagents have the same ingredients, but TRIzol j has a lower total concentration (Life Technologies, http://www.lifetechnologies.com/de/de/home/brands/product-brand/trizol.html). Accordingly, adopting the lysis protocol published for TRIzol LSd,16 to TRIzol, j SBV RNA extraction from semen may be less efficient using the latter reagent. The results suggest that the use of different extraction protocols was the major reason for the significantly different PCR results (Fig. 2). A higher sensitivity and consistency of PCR results using a TRIzol LS d –based extraction method compared with other extraction methods was reported in previous studies. TRIzol–chloroform-based extraction efficiently removes PCR inhibitors and cellular waste present in semen and prevents the clogging of membranes of subsequently used membrane-based extraction kits.16,24,25 Magnetic bead–based methods might therefore avoid the drawback of membrane-based extraction. Sensitive PCR testing after automated f extraction of SBV and Bluetongue virus RNA with a magnetic bead–based extraction kit without a TRIzol pretreatment was reported by other groups.13,22 In our study, SBV RNA was detected in semen samples extracted with the recommended automated magnetic bead–based extractione,f without pretreatment with TRIzol LS d (Table 2) and confirms the high sensitivity reported previously.15,16 On the other hand, the results of our study indicate that other factors, such as ingredients of individual kits, have a considerable influence on the PCR results with difficult sample matrices: only half of magnetic bead, but all of membrane-based extractions, conducted without pretreatment with any of the TRIzol preparationsd,j-l revealed true-positive PCR results in the first trial. However, further studies with a higher number of samples are required to confirm this observation.
Our study results were not markedly influenced by different RT-qPCR kits and assays (Supplementary Tables 1, 2), temperature profiles (data not shown), and laboratory procedures because of the high number of similar RT-qPCR protocols and the significant differences found for PCR results statistically analyzed with respect to the extraction methods (Table 2). The high proportion of false-negative PCR results reported by 2 laboratories (EU12 and EU14) that used an SBV S segment b – and L segment–specific RT-qPCR assay, respectively, other than the previously described assay a and the Qiagen virotype assay i was most likely because of the use of alternative extraction methods. Despite a slightly lower sensitivity reported for an SBV L segment–specific RT-qPCR compared with the S segment–specific assay,a,10 the amount of SBV RNA in the semen samples was sufficiently high for reliable SBV RNA detection. These 2 laboratories detected SBV RNA in all other matrices, except in samples with SBV RNA amounts at the detection limit. An efficient extraction method would result in high amounts of SBV RNA, which is a prerequisite for sensitive PCR testing. The consistently negative results determined for SBV RNA–negative semen batches indicate a high DSp (100%), but a larger number of samples is required to confirm this result.
To date, the dynamics, clinical importance, and duration of SBV RNA excretion in semen from bulls after SBV infection remain unknown. SBV RNA was not detected in male genitals (prostate, testicle, urethral bulb, epididymis) ~3 weeks after experimental SBV infection of bulls that excreted SBV RNA in their semen. 24 The high variability of excretion patterns further complicates the safe detection of SBV RNA excretion in semen from SBV-infected bulls.16,22,24 SBV RNA may be consecutively or intermittently excreted on a single day up to at least 3 months after SBV infection, despite seroconversion. Furthermore, semen with high and low viral loads from SBV-infected bulls may contain infectious SBV. However, the possibility of venereal transmission remains to be investigated.18,21,24
For reliable detection of SBV, it is important to test semen from SBV-infected bulls with highly sensitive methods.16,21 Although semen batches with predefined reference Cq values were provided for the validation of the PCR protocols used at the participating laboratories, not all SBV RNA–positive semen batches were detected as positive. Accordingly, a thorough validation of extraction methods with standardized reference semen batches is essential to establish extraction methods and PCR protocols for reliable detection of SBV RNA in bovine semen. Furthermore, testing of at least 2 biological and 2 technical replicates has been recommended for SBV RNA detection in semen. 16
In conclusion, the extraction and RT-qPCR protocols used by the participating German and other European laboratories for SBV RNA detection in all matrices, except semen, were found to be robust and produced positive results for all clearly SBV RNA–positive specimens and in most of the samples with SBV RNA loads at the detection limit. In contrast, the application of a highly effective extraction method that reduces or compensates for the inhibitory effects of semen components on PCR results was found critical for reliable detection of SBV RNA in bovine semen. The high sensitivity of the published and recommended extraction protocols for SBV RNA detection in bovine semen were confirmed by both trials. Although several other extraction procedures in combination with TRIzol LS d showed similar detection sensitivity, alternative methods should be thoroughly validated with standardized reference semen batches. Highly sensitive methods that allow the detection of a SBV genome load below the detection limit of Cq 35, and testing of biological and technical replicates are strongly recommended because SBV RNA–positive semen with a Cq value of 34.2 has been shown infectious for cattle injected subcutaneously. 21 Our investigation of viral genome detection in semen might be used as reference for future validations of extraction methods for other orthobunyaviruses and possibly RNA viruses to allow their sensitive detection in semen samples.
Footnotes
Acknowledgements
Patrick Zitzow, Susanne Zahnow, Karin Lissek, Christian Korthase, and Martina Abs are acknowledged for excellent technical assistance, and the German bull breeding centers for providing the semen samples. We thank Akbar Dastjerdi and Anna LaRocca for managing the testing at AHVLA.
Authors’ contributions
C Schulz and B Hoffmann contributed to conception and design of the study. M Beer contributed to conception of the study. C Schulz contributed to acquisition, analysis, and interpretation of data. WHM van der Poel, C Ponsart, AB Cay, F Steinbach, and S Zientara contributed to acquisition and analysis of data. M Beer and B Hoffmann contributed to analysis and interpretation of data. C Schulz drafted the manuscript. WHM van der Poel, AB Cay, F Steinbach, S Zientara, M Beer, and B Hoffmann critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
SBV-S3, small-segment–specific SBV-RT-qPCR according to Bilk et al. 4
b.
LSI VetMAX Schmallenberg virus real-time PCR kit (s gene), Life Technologies, Darmstadt, Germany.
c.
QIAamp viral RNA mini kit, Qiagen GmbH, Hilden, Ger-many.
d.
Ambion TRIzol LS reagent, Life Technologies, Grand Island, NY.
e.
MagAttract virus mini M48 kit, Qiagen Inc., Valencia, CA.
f.
KingFisher Flex, Thermo Fisher Scientific, Schwerte, Germany.
g.
Applied Biosystems AgPath-ID one-step RT-PCR kit, Life Technologies, Grand Island, NY.
h.
CFX96 real-time system, Bio-Rad Laboratories GmbH, Munich, Germany.
i.
virotype SBV RT-PCR kit, Qiagen Inc., Valencia, CA.
j.
Invitrogen TRIzol reagent, Life Technologies, Grand Island, NY.
k.
TRI reagent, Sigma-Aldrich Corp., St. Louis, MO.
l.
QIAzol lysis reagent, Qiagen Inc., Valencia, CA.
m.
KingFisher family, Thermo Fisher Scientific, and BioSprint, Qiagen and are the same instruments distributed by different manufacturers.
n.
Chemagic stand, PerkinElmer Chemagen Technologie GmbH, Baesweiler, Germany.
o.
Corbett Life Science X-Tractor gene, CAS-1820 nucleic acid extractor; Qiagen Inc., Valencia, CA.
p.
MagNA Pure LC 2.0, Roche Diagnostics GmbH, Mannheim, Germany.
q.
NorDiag Magnatrix 8000, DiaSorin AB, Solna, Sweden.
r.
NucleoMag VET kit, Macherey-Nagel GmbH & Co. KG, Düren, Germany.
s.
INSTANT virus RNA kit (now merchandised as “innuPrep virus RNA kit”), Analytik Jena AG, Jena, Germany.
t.
Applied Biosystems MagMax pathogen RNA/DNA kit, Life Technologies, Grand Island, NY.
u.
NucleoSpin RNA virus kit, Macherey-Nagel GmbH & Co. KG, Düren, Germany.
v.
InvitTrap spin tissue RNA mini kit, STRATEC Molecular GmbH, Berlin, Germany.
w.
MagNA Pure LC total nucleic acid isolation kit, Roche Diagnostics Corp., Indianapolis, IN.
x.
LSI MagVET universal isolation kit, Life Technologies, Grand Island, NY.
y.
BioSprint 96 one-for-all vet kit, Qiagen Inc., Valencia, CA.
z.
BioSprint 96 DNA blood kit, Qiagen Inc., Valencia, CA.
aa.
NorDiag Vet NA kit (now merchandised as “Bullet stool kit”), DiaSorin, Dublin, Ireland.
bb.
QIAamp cador pathogen mini kit, Qiagen Inc., Valencia, CA.
cc.
QuantiTect probe RT-PCR kit, Qiagen Inc., Valencia, CA.
dd.
Brilliant II QRTPCR master mix with ROX (1-step), Agilent Technologies Inc., La Jolla, CA.
ee.
QuantiTect virus kit, Qiagen Inc., Valencia, CA.
ff.
qScript one-step qRT-PCR kit, Quanta BioSciences Inc., Gaithersburg, MD.
gg.
QuantiFast probe RT-PCR, Qiagen Inc., Valencia, CA.
hh.
EZ1 Advanced XL, Qiagen Inc., Valencia, CA.
ii.
MagNa Pure 96 DNA and Viral NA small volume, Roche Diagnostics Corp., Indianapolis, IN.
jj.
EZ1 RNA universal tissue kit, Qiagen Inc., Valencia, CA.
kk.
M-MLV reverse transcriptase, Promega Corp., Madison, WI.
ll.
FastStart TaqMan probe master, Roche Diagnostics Corp., Indianapolis, IN.
mm.
QuantiFast pathogen RT-PCR +IC kit, Qiagen Inc., Valencia, CA.
nn.
RNeasy mini kit, Qiagen Inc., Valencia, CA.
oo.
High Pure viral nucleic acid preparation kit, Roche Diagnostics Corp., Indianapolis, IN.
pp.
ABI 7500 or 7300, Life Technologies, Grand Island, NY.
qq.
LightCycler 480, Roche Diagnostics Corp., Indianapolis, IN.
rr.
Mx3005P qPCR system, Agilent Technologies Inc., Santa Clara, CA.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study was supported by the German Federal Ministry of Food, Agriculture and consumer protection and the European Union as outlined in Council Decision 2012/349/EU concerning a financial contribution by the Union for studies on Schmallenberg virus.
