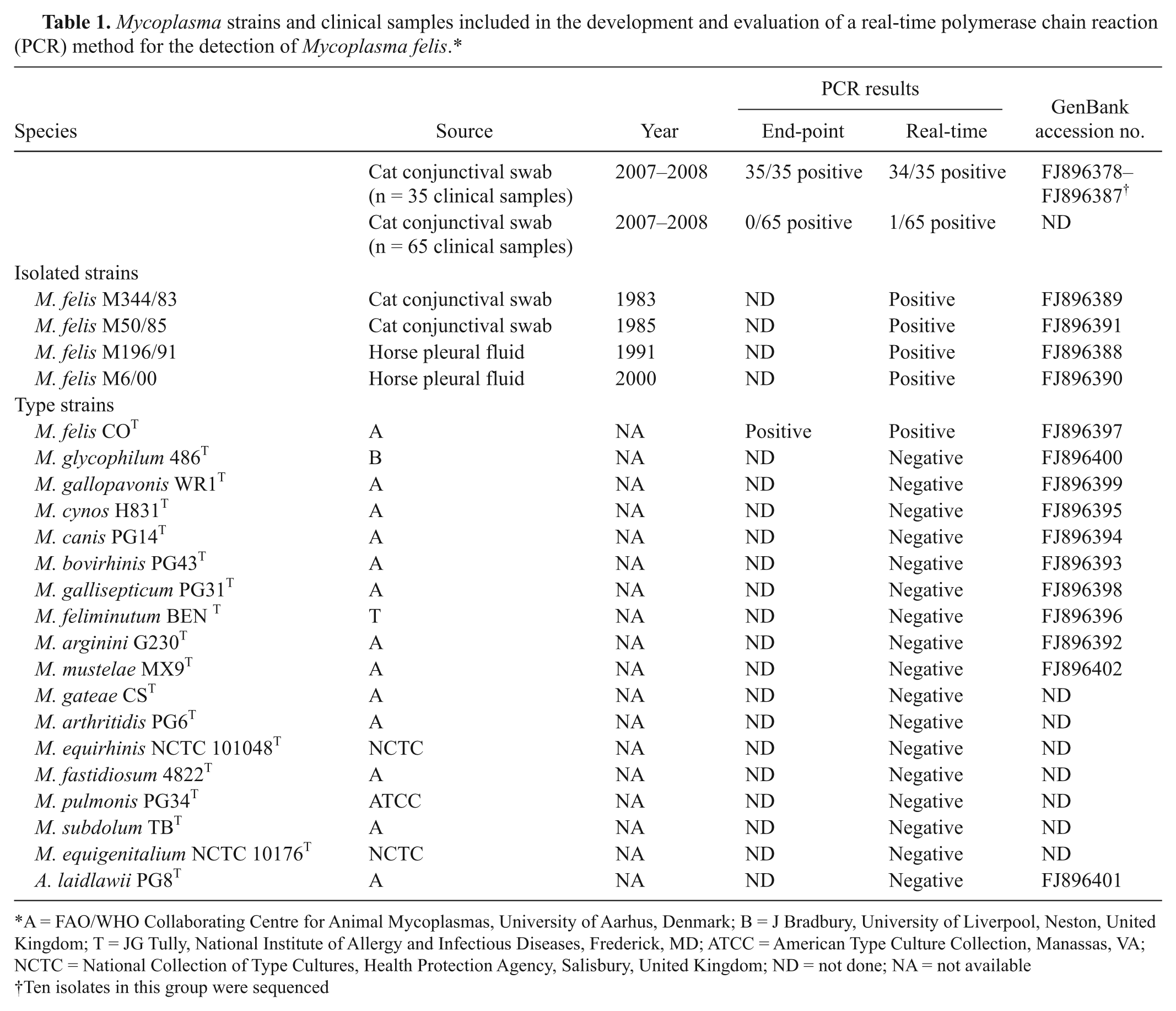
Infection by Mycoplasma felis is associated with ocular and respiratory disease in cats and respiratory disease in horses. A correct diagnosis is beneficial since the use of specific antimycoplasmal treatment can lead to resolution. The objective of the present study was to develop a real-time polymerase chain reaction (PCR) method based on dual-labeled fluorogenic probe technology, targeting the gene encoding elongation factor Tu (tuf ), for the fast and specific detection of M. felis. Specificity was achieved by basing the assay design on partial sequencing of the tuf gene in strains and clinical isolates of M. felis as well as other mycoplasma species. The detection limit of the developed assay was in the order of 10 copies of target DNA, and no cross-reaction was observed with a panel of several mycoplasma species. Compared to a previously published conventional PCR protocol, the novel assay had equal or slightly improved performance in terms of sensitivity and specificity when analyzing 100 conjunctival swab samples from cats with clinical signs of infection.
Several mycoplasma species are part of the normal bacterial flora of animals. Many of these mycoplasmas are regarded as harmless; however, Mycoplasma felis can be pathogenic for both cats and horses. Mycoplasma felis is rarely found in ocular swabs from healthy cats, but it is prevalent in swabs from cats with conjunctivitis,
6
which has been experimentally induced in kittens following inoculation.
23
In the oropharynx, M. felis is considered part of the normal flora of approximately a third of the feline population,
16
but in shelter cats, isolation of M. felis from oropharyngeal swabs has been associated with upper respiratory tract infection.
1
While Mycoplasma felis is generally not present in bronchoalveolar lavages of clinically healthy cats,
16
it has been associated with lower respiratory tract disease,
25
as well as with cases of ulcerative keratitis,
5
chronic rhinosinusitis,
8
and arthritis.
10
Diagnosis of M. felis infection in such cases is beneficial, as appropriate therapies can effectively treat the disease.5,7,25 Other mycoplasma species found in the upper and lower respiratory tract of cats include M. arginini, M. pulmonis, M. arthritides, Acholeplasma laidlawii, and rarely M. feliminutum.17,24 Although Mycoplasma gateae can cause arthritis,
12
it is also the most common mycoplasma found in clinically healthy cats.
24
Thus, diagnostic methods targeting M. felis should not produce false positives when other mycoplasma species are present in a sample.
In horses, tracheal isolation of M. felis has been associated with sudden onset of coughing, nasal discharge, or pyrexia,
14
and with outbreaks of lower respiratory tract disease.
26
Mycoplasma felis has also been detected in pleural fluid of horses with pleuritis.
15
As in the cat, a fast and specific etiological diagnosis is beneficial, as appropriate treatment cures the disease.
15
Other known respiratory mycoplasmas of horses include M. equirhinis, M. arginini, M. subdolum, and M. fastidiosum.
9
The traditional diagnostic technique used for M. felis is microbiological culture, including identification of M. felis with specific antisera, which is done only in specialized laboratories and can take weeks. More recently, a method using polymerase chain reaction (PCR) for detection of M. felis has been reported.
4
The PCR assay was sensitive and faster than conventional culture and identification techniques, but the assay was based on traditional end-point PCR technology. The same assay was later adapted for real-time PCR using SYBR Green intercalating dye technology.
19
In addition to the advantage of reduced analysis time, real-time PCR analysis can be performed with closed PCR tubes, which makes it easier to handle a large number of samples and minimizes the risk of cross-contamination. The targeting of a third sequence element by the use of a dual-labeled fluorogenic probe enables increased specificity compared to SYBR Green-based assays and reduces the time and effort of each analysis by eliminating the need for melting curve analysis. The objective of the current work was the development of an improved M. felis assay based on real-time PCR using dual labeled fluorogenic probe technology. The gene encoding elongation factor Tu (tuf ) was chosen as the PCR target since it has been shown to be sufficiently conserved between mycoplasma species to allow sequencing while containing regions variable enough for species discrimination.11,20
Mycoplasma strains (see Table 1) were grown in F-medium as previously described
2
for 1–3 days. Ten milliliters of medium were pelleted for 15 min at 25,000 × g and then washed with 10 ml of 0.8% sodium chloride. The washed pellet was incubated with 20 µl of proteinase K solution (>600 mAU/ml) in 200 µl of buffer
a
for 10 min at 56°C with agitation, and DNA was extracted from the lysate with an automated workstation.
b
One hundred conjunctival swabs from cats with suspected M. felis infection were submitted at ambient temperature to the National Veterinary Institute (Uppsala, Sweden) during 2007 and 2008. The swabs were shaken in 1.5 ml of 0.8% sodium chloride for 15 min at room temperature, and then the liquid was centrifuged at 12,000 × g for 5 min. The collected cells were resuspended in 50 µl of Tris–ethylenediamine tetra-acetic acid buffer and incubated at 95°C for 20 min with agitation. After a final centrifugation at 12,000 × g for 1 min, the supernatant was frozen and stored at −20°C until needed.
For partial sequencing of the tuf gene, an initial amplicon was generated using the primers Msp27f and Msp28r.
11
Sequencing reactions and reads were performed using these primers for all clinical samples and strains sequenced. Additionally, M. felis samples and strains were read from the same initial amplicon using the internal primers Mftuf1R (5’-CAACGATACGAGGAACACCA-3’) and Mftuf2F (5’-AGAAGCACTTGCGGGAGATA-3’). The partial tuf gene sequences were submitted to GenBank with accession numbers as presented in Table 1.
Based on partial tuf gene sequences from mycoplasma strains, primers and probes were designed for real-time PCR using Primer3 v.0.4.0.
18
The primer–probe combinations were positioned to ensure that the mycoplasma species in Table 1 other than M. felis had substantially divergent sequence in the target regions, and that the sequence was conserved in the available tuf gene sequences of feline strains and clinical samples (Table 1). Several combinations of primers and probes were initially evaluated in terms of specificity and amplification efficiency. The primers Mftuf3F (5’-TAAATTAGCTCTTGATGGTGTTCCT-3’), Mftuf3R (5’-TTCAAAGTCTTTTTCTGGAGTTTCA-3’), and probe Mftuf3P (5’-[6-FAM]- TGAGAAGAAAAAGTTATGGAATTAATGGATGCA-[BHQ1a]-3’) were selected for further evaluation. The product size of the PCR was 100 base pairs (bp). Specificity of the primers and probe were verified in silico by using the Basic Local Alignment Search Tool (BLAST) algorithm (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi; last accessed on March 20, 2011). For assay evaluation, a 400-bp fragment of the tuf gene from the M. felis strain COT containing the assay target region was synthesized, cloned into a pCR 2.1 vector, and verified by sequencing using a service provider.
c
For the real-time PCR assay, a total reaction volume of 25 µl was used, with 0.5 U of DNA polymerase with buffer,
d
3 mM magnesium chloride, deoxyribonucleotide triphosphate mix at 0.1 mM of each nucleotide, each primer at 0.3 µM, probe at 0.1 µM, and 2 µl of whole-genome DNA template extracted as described above. A real-time PCR instrument
e
was used with the following settings: 94°C for 10 min, 45 cycles of 95°C for 15 sec, and 59°C for 60 sec. The threshold cycle (Ct) threshold was set to a normalized fluorescence of 0.02 in the instrument software, and any Ct <35 was considered a clear positive. Samples with Ct values ≥35 were re-analyzed for confirmation, with the sample declared positive if the fluorescence threshold was reached in this second run, regardless of Ct value. Each PCR run included a positive control reaction containing M. felis COT genomic DNA and a negative control with no template added. All conjunctival swab samples were analyzed in parallel for Chlamydiaceae using a PCR assay that includes an internal amplification control, as previously described.
21
When analyzing DNA extracts from strains of other mycoplasma species, PCR inhibition was ruled out by running a replicate of each sample spiked with 0.3 ng of M. felis COT DNA in parallel. Representative positive and negative real-time PCR reactions from the analysis of M. felis cat conjunctival swab samples were subjected to electrophoresis on 2% agarose gel to confirm the presence or absence of amplicon. Conventional PCR analysis was performed according to a published method.
4
For the novel real-time assay, a 10-fold dilution series starting from 108 plasmid template copies per reaction in triplicate produced a linear relationships between log template amount and Ct value over 8 orders of magnitude, with R
2
= 0.996. Based on this curve, the amplification efficiency (AE) was 1.1, using the formula AE = [10(−1/M)] − 1. All replicates containing approximately 10 target copies were positive. In agreement with this finding, a dilution experiment performed using genomic DNA extracted from M. felis COT culture found all replicates positive down to a dilution corresponding to 12 fg of genomic DNA per reaction. The M. felis chromosome size has been estimated at 1,055 kbp by pulsed field electrophoresis,
13
which corresponds to approximately 1.2 fg. PCR-positive reactions were observed at the single-copy dilution level at the expected Ct values, but with loss of signal in some replicates, likely due to complete absence of template material. Thus, the developed PCR appears to be able to detect very low copy numbers or single copies of target DNA. All 5 available M. felis strains tested positive using the presented assay, while no cross reaction was observed with any of the 17 other mycoplasma species assayed using 5 ng of DNA extracted from a culture of each species as template (Table 1).
As further evaluation, the assay was used to re-analyze 100 DNA samples extracted from conjunctival swabs from cats with clinical signs of ocular infection, in which there were a total of 35 positive and 65 negative when analyzed using a conventional PCR assay.
4
For 98 of the 100 clinical samples re-analyzed, the conventional PCR results were confirmed by the new real-time PCR assay. One sample previously negative when analyzed by conventional PCR tested positive by real-time PCR with the only Ct value >35 in the panel. One sample previously positive by conventional PCR tested negative by real-time PCR and was, by sequencing, found to have a tuf gene sequence identical to that of Mycoplasma cynos H831T. While the true M. felis infection status of the sampled cats is undetermined, the results suggest that the presented assay has equal or slightly improved sensitivity and specificity compared to the conventional PCR protocol. There was no PCR inhibition observed in any sample, and all positive and negative control reactions produced the expected results. Two stored strains isolated from horses with pleuritis in 1991 and 2000 tested positive with the developed assay, and sequencing revealed that in these isolates (GenBank accession nos. FJ896388 and FJ896390, respectively) an identical tuf variant had a single base mismatch to the reverse primer. A 10-fold dilution curve of genomic DNA carrying this mismatch from 3 ng to 30 fg yielded R2 = 0.997 and efficiency = 0.96. As described by others, single base mismatches in a primer can have very limited effect on amplification performance
22
although the impact likely depends on the primer sequence itself and the mismatch position.
3
Since the effects of the mismatch in the current case were benign and the assay target region highly favorable in terms of species specificity, the assay was not redesigned.
In conclusion, detection of the pathogen M. felis in samples from cats and horses is clinically relevant. In the current study, a novel real-time PCR assay was developed that could represent an attractive option for rapid, sensitive, and specific detection of M. felis for research and routine diagnostic applications.