Abstract
An autosomal recessive genetic defect termed JH1 has been associated with early embryonic loss in the Jersey cattle breed. The genetic basis has been identified as a cytosine to thymine mutation in the
Keywords
Jersey cattle are a small dairy animal popular for their high milk fat percentage, low maintenance costs, and tolerance to heat stress. 4 However, intense selection has led to rapid inbreeding accumulation. A rate of inbreeding of 0.14% per year from 2000 to 2007 and a small effective population size of ~68 was reported in Canadian Jerseys. 3 The average inbreeding level has increased to 6.00% and 7.09% for cows born in 2013 in Canada (Canadian Dairy Network, 2014, Inbreeding update—August 2014. Available at: http://www.cdn.ca/articles.php) and the United States (Council on Dairy Cattle Breeding, 2015, Trend in inbreeding coefficients of cows for Jersey. Available at: https://www.cdcb.us/eval/summary/inbrd.cfm?R_Menu=JE#StartBody), respectively.
Inbreeding can result in negative effects such as inbreeding depression and increased prevalence of rare deleterious recessive alleles.
1
A new autosomal lethal recessive defect (denoted as JH1) in Jersey cattle was reported in 2011 by identifying haplotypes with high population frequencies but lacking homozygotes.
5
The JH1 defect was found to have an extremely high carrier frequency in the U.S. Jersey population, and this defect was associated with early embryonic loss.
5
Most recently, the causative mutation of JH1 was identified as a transition from cytosine (C) to thymine (T) in the
Blood samples were collected from 449 Jersey cows that were imported from Australia in 2012. Genomic DNA was extracted using a commercial kit.
a
Using the DNA sequence of
The PCR product was 1,246 base pairs (bp) in length. There are 2
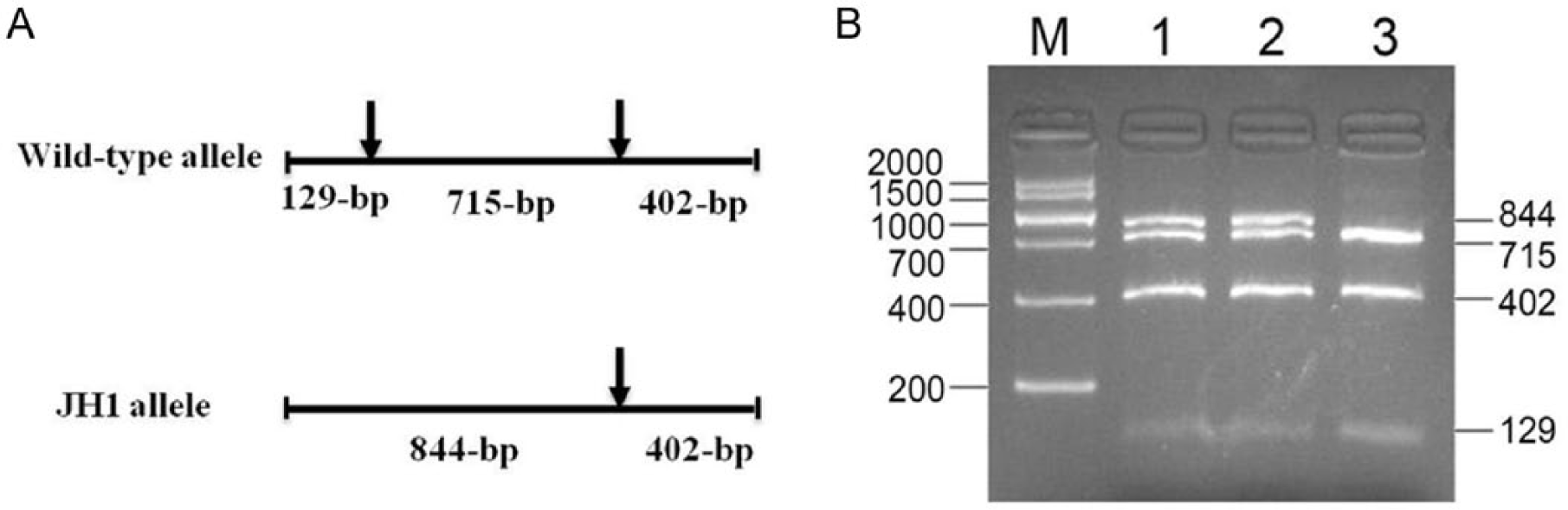
The restriction fragment patterns for different alleles (
The previous strategy for JH1 carrier testing is a haplotype-based method that relies on haplotype inference of 17 single nucleotide polymorphism (SNP) markers spanning ~5 Mb on BTA15. 5 However, for a small population without large SNP data for accurate haplotype inference, the haplotype-based method is inapplicable. The testing method reported herein is simple and cost-efficient because it directly examines the JH1 mutation by PCR-RFLP. Moreover, the reliability of the method was enhanced by utilizing another identical restriction site at 715 nucleotides downstream of the JH1 mutation, which helps prevent false-positive test results due to incomplete digestion in the RFLP analysis.
The JH1 defect is probably the most frequent inherited defect in Jersey cattle. From the common ancestor, an elite U.S. Jersey sire (Observer Chocolate Soldier) born in 1962, the JH1 mutation has been widely spread, leading to a carrier frequency of 20–25% in Jerseys in the United States. 5 The 10 most contributing ancestors accounted for ~60% of the gene pool of Canadian Jerseys in 2000–2007. 3 Notably, 2 of the bulls (Observer Chocolate Soldier and Soldierboy Boomer Sooner of CJ) are JH1 carriers (http://www.cdn.ca/query/individual.php).
In the current study, 31 JH1 carriers were identified among 449 cows, corresponding to a carrier frequency of 6.9%, which is substantially lower than that in the U.S. population (20–25%). 5 Pedigree analyses were performed to trace the ancestries of the carriers using the pedigree databases of the Canadian Dairy Network (https://www.cdn.ca/) and the Australian Dairy Herd Improvement Scheme (http://www.adhis.com.au/). Twenty of the 31 JH1 carriers showed paternal grandsires from U.S. bulls, indicating the high impact of U.S. ancestry on Australian Jerseys. Seventeen carriers were traced back to the common ancestor, Observer Chocolate Soldier. His grandson, Soldierboy Boomer Sooner of CJ, however, is the bull most responsible for the spread of the JH1 lethal allele (Fig. 2).
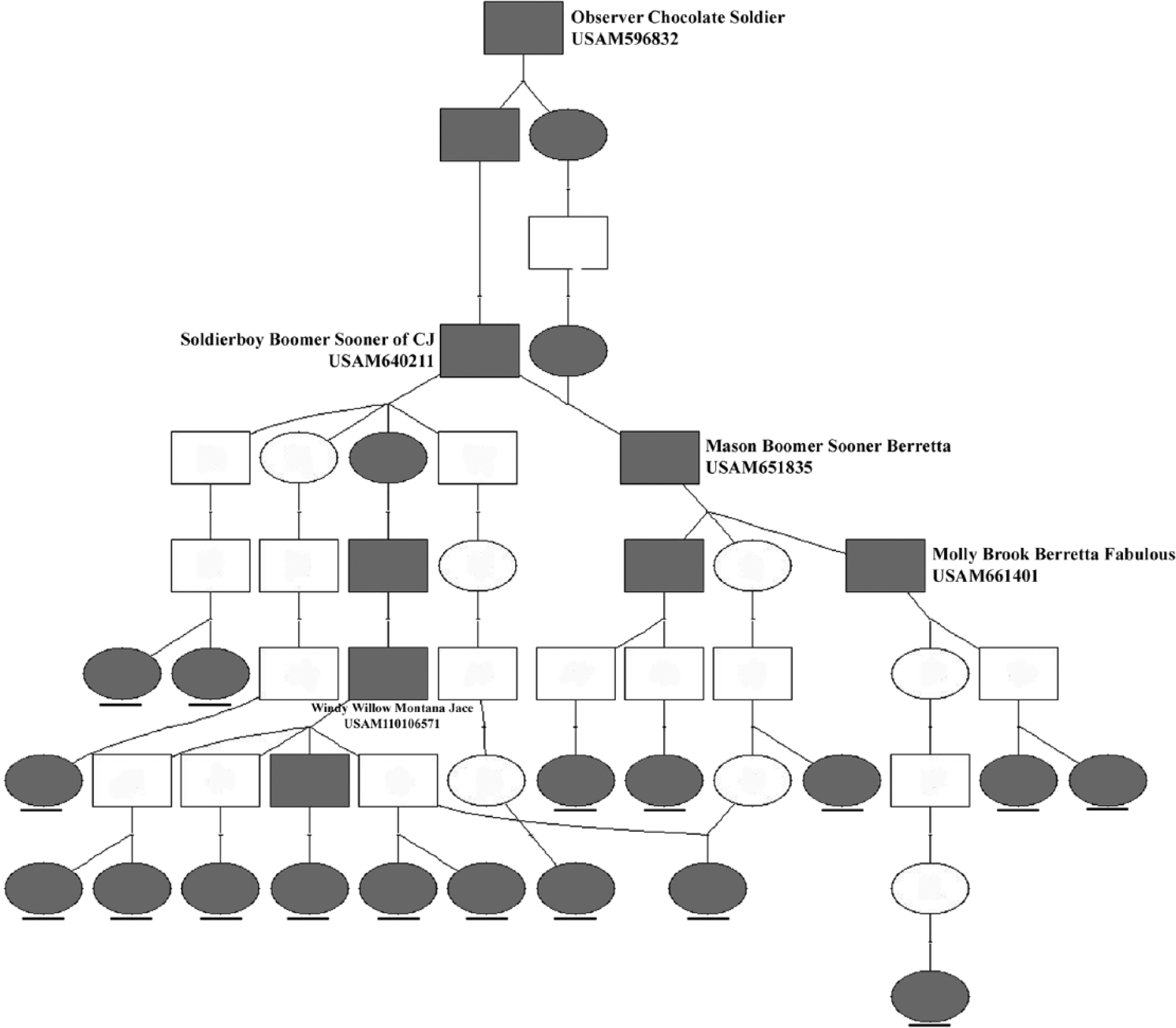
Pedigree network of the JH1 carriers constructed using the Pedigraph software (http://animalgene.umn.edu/pedigraph/). Open square = male without genotype; open circle = female without genotype; gray square = male carrier; gray circle = female carrier. The carriers that were identified in the current study are underlined.
The PCR-RFLP assay was validated in 3 ways. First, direct sequencing of PCR products was performed on 8 identified positives (carriers) and 13 negatives (noncarriers). As expected, the positives showed heterozygous peaks (C/T) at the causative mutation while the negatives were homozygous wild allele (C). Second, 9 U.S. Jersey bulls with known JH1 genotypes (5 carriers, 4 noncarriers) were tested using our new method, and all were confirmed. Third, 3 negatives and 3 positives shown by the PCR-RFLP assay were genotyped by a commercial SNP chip-based assay. d Results showed no discrepancies between the 2 assay strategies. These results demonstrated that the PCR-RFLP is a reliable assay for genotyping the JH1 defect.
In summary, a PCR-RFLP genotyping test was developed for the JH1 defect, and the carrier frequency was examined in an imported Jersey population in China. The high frequency of the JH1 carriers suggests that evaluating the genetic defect by pedigree analysis and our new method is necessary for the imported Jersey cattle. In addition, all active sires should be screened so as to avoid the risk of carrier-to-carrier matings and to gradually eliminate the deleterious gene.
Footnotes
Acknowledgements
Yi Zhang and Gang Guo contributed equally to this work. We thank Prof. JSF Barker for his assistance in editing the English used in this article and Robert Watson for providing the U.S. Jersey samples for the validation work.
Authors’ contributions
Y Zhang contributed to conception and design of the study; contributed to analysis of data; drafted the manuscript; and critically revised the manuscript. G Guo contributed to design of the study; contributed to interpretation of data; and drafted the manuscript. H Huang and L Liu contributed to design of the study; contributed to acquisition of data; and drafted the manuscript. L Lu contributed to design of the study; contributed to interpretation of data; and drafted the manuscript. L Wang contributed to conception of the study; contributed to analysis of data; and drafted the manuscript. L Fang contributed to acquisition of data, and drafted the manuscript. Y Wang contributed to design of the study; contributed to acquisition of data; and critically revised the manuscript. S Zhang contributed to conception and design of the study; contributed to interpretation of data; drafted the manuscript; and critically revised the manuscript. All authors gave final approval and agree to be accountable for all aspects of the work in ensuring that questions relating to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
a.
Tiangen Biotechnology Co Ltd, Beijing, China.
b.
Takara Biotechnology Co Ltd, Dalian, China.
c.
New England Biolabs Inc., Ipswich, MA.
d.
GeneSeek, Lincoln, NE.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: National Key Technologies Research and Development Program (2011BAD28B02, 2012BAD12B01), Beijing Science and Technology Program (Z141100003714121), China Agricultural Research System (CARS-37), and Program for Changjiang Scholars and Innovative Research in University (IRT1191).
