Abstract
Brachyspina syndrome (BS) is a rare monogenic autosomal recessive hereditary disorder identified in the Holstein breed caused by a 3.3-kb deletion in the bovine Fanconi anemia complementation group I (FANCI) gene on chromosome 21. In previous reports, the BS mutant allele was identified in North American and European Holstein populations. Because frozen semen and embryos have been imported into China from such regions during the past few years, BS has potentially spread into the dairy cattle population in China. In the present study, 206 Holstein bulls and 136 Holstein cows were tested for BS, with 10 BS carrier bulls and 3 carrier cows identified. Pedigree analysis showed that all 10 BS carrier bulls could be traced back to a common ancestor, the U.S. Holstein sire Sweet Haven Tradition. It is recommended that effective selection and mating strategies should be carried out to gradually eliminate this recessive gene from the Chinese Holstein population.
Brachyspina syndrome (BS), a genetic abnormality in the Holstein cattle breed that causes either early-term abortion (most common) or stillborn calves (rare) when an individual is homozygous recessive for the lethal gene, was first observed in Denmark in 2006. 2 Later, additional cases were reported in the Netherlands, 3 Italy, 6 Germany, 4 and Canada. 1 The cases were characterized morphologically by severely reduced body weight despite a normal or slightly prolonged gestation period, obvious shortening of the spine, long and slender limbs, inferior brachygnathism, and internal organ malformation, such as renal and gonadal dysplasia. At the same time, the defect might be responsible for some fertility problems, such as a high proportion of abortions and long calving interval. 5 A large proportion of cases probably die during embryonic or early fetal development (similar to CVM 7 ), thus causing reduced fertility and unpredictable economic losses to the breeders.
Previous studies mapped the gene and mutation causing BS to base positions 20156961 to 22499122 of bovine chromosome 21 (Btau_4.0) and defined a diagnostic single nucleotide polymorphism (SNP) haplotype to detect BS carriers (Georges M, et al., inventors; Université de Liège, et al., assignees: 2010 Feb 4, A genetic marker test for brachyspina and fertility in cattle. International patent WO/2010/012690). In 2012, the causal mutation for BS was successfully identified by next-generation sequencing as a 3.3-kb deletion in the bovine Fanconi anemia complementation-group 1 (FANCI) gene. 5 The deletion removed exons 25–27 of the 37 exons composing FANCI, and led to a frame-shift substituting the 451 carboxy-terminal amino acids with a 26 residue long illegitimate peptide. 5
In the European cases,2-4,6 the mutation was traced back to the elite U.S. Holstein sire, Sweet Haven Tradition (HOUSAM 1682485). However, in the Canadian case, 1 a more remote ancestor was suggested. Because descendants of this sire family have been commonly used in the dairy cattle breeding programs of many countries, it is presumed that the BS gene has been spread worldwide. The purpose of the present study was to detect and determine the frequency of BS in the Chinese Holstein population.
Frozen semen samples of 206 Holstein bulls were collected from the Beijing Dairy Cattle Center (Beijing, China), which is the predominant artificial insemination center in China. In addition, blood samples of 136 Chinese Holstein cows were randomly collected from 4 dairy herds in the Beijing region by trained veterinarians following standard procedures.
Genomic DNA was extracted from blood and semen samples by a standard phenol-chloroform method, and was diluted to a final concentration of 200 ng/μl. The quality and quantity of DNA was estimated by using 1% agarose gel electrophoresis prior to polymerase chain reaction (PCR) amplification.
The primers spanning the 3.3-kb deletion described previously 5 were used to detect the defective allele for BS. The forward and reverse primer sequences were 5’-GCTCAAGT AGTTAGTTGCTCCACTG-3’ and 5’-ATAAATAAATAA AGCAGGATGCTGAAA-3’, respectively. Only one 3,738-bp DNA fragment was amplified for normal animals, while 2 fragments, 409 bp and 3,738 bp, were expected to be generated for BS carriers (Fig. 1). The PCR was set up in a final volume of 25 μl containing 100 ng of genomic DNA template, 1 μM each of the above primers, 2.5 μl 10× PCR buffer (Mg2+ Plus), 4 μl of 2.5 mM deoxyribonucleotide triphosphate mixture, and 1.5 U Taq polymerase. a The mixture was incubated in a calibrated thermal cycler. b Initial denaturation was achieved at 94°C for 5 min, followed by 35 cycles of 30 sec at 94°C, annealing for 1 min at 58°C, and extension for 2.5 min at 72°C, then the final extension for 10 min at 72°C. The PCR products were separated on 1.5% agarose gels and were visualized under an ultraviolet transilluminator.
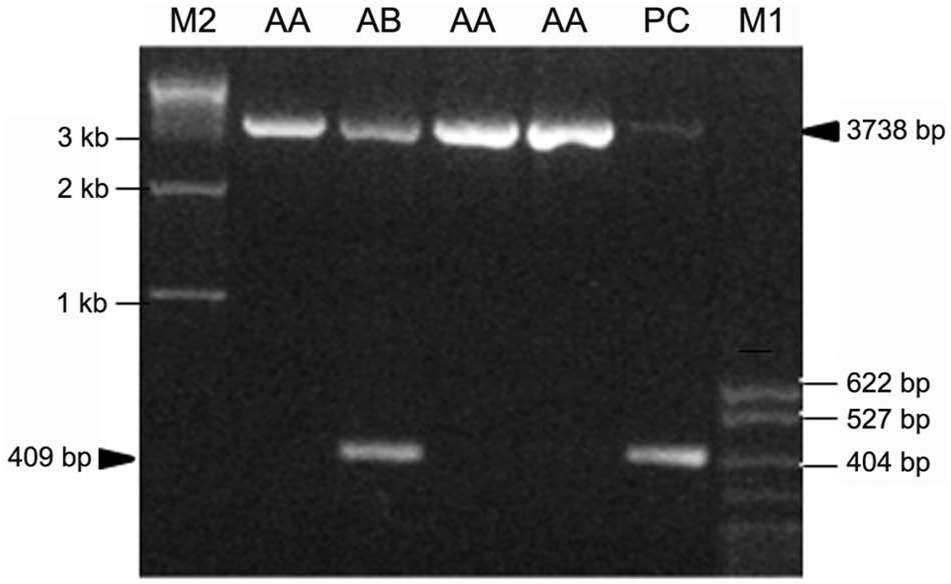
Polymerase chain reaction products of FANCI gene of Chinese Holstein bulls and cows. Lane M2: 1-kb DNA ladder; lane AA: normal animals; lane AB: brachyspina syndrome carrier; lane PC: positive control; lane M1: pBR322/Mspl marker DNA.
Using the above PCR-based genotyping method and a known BS carrier, Gillette Jordan (CANM7588022), as positive control, a total of 206 Holstein bulls and 136 Chinese Holstein cows were checked for BS. Of these, 10 bulls and 3 cows were identified as BS carriers. Thus, the frequency of the heterozygote carriers was 4.9% in bulls and 2.2% in cows. As expected, no mutant homozygote was found.
Pedigree analysis was implemented to trace the ancestries of these carriers through the pedigree databases of the Dairy Association of China and Holstein Association USA. All of the 10 BS carrier bulls were traced back on either the sire or dam side to the common ancestor, Sweet Haven Tradition, in generation VII (Fig. 2), and no more remote ancestors were found. The analysis showed that this genetic defect spread into China mainly through the progeny of Sweet Haven Tradition, especially Bis-May Tradition Cleitus (HOUSAM 1879085) and Rothrock Tradition Leadman (HOUSAM 1983348). However, the ancestry of the 3 BS carrier cows was not clear, so analysis of their pedigrees could not be conducted.
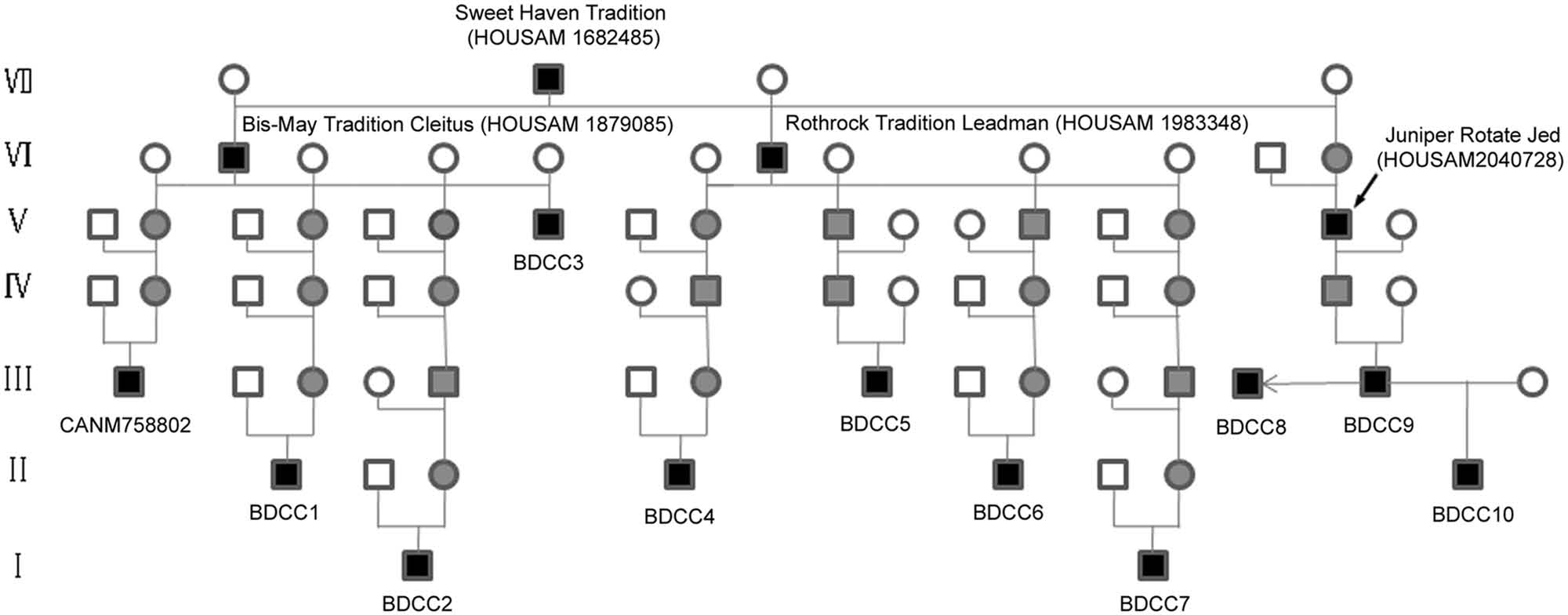
Genealogical diagram of 10 identified brachyspina syndrome (BS) carrier bulls (BDCC1–BDCC10) and Gillette Jordan (CANM7588022). BDCC8 is a clone of BDCC9; white circle = normal female; white square = normal male; black square = male BS carrier; gray square = presumed male BS carrier based on pedigree analysis; gray circle = presumed female BS carrier based on pedigree analysis.
In the current study, the BS allele was identified in Chinese Holsteins, which adds further information to its suspected worldwide dissemination. The frequency of BS carriers in the present study (4.9% in bulls and 2.2% in cows) is lower than that reported in the Netherlands (7.4%) and the United States (6%).5,8
Although the elimination of all BS carrier bulls would be the most efficient method to control this genetic disorder, many carrier bulls are still listed commercially for artificial insemination in China. Direct culling of all at-risk sires is not economically beneficial for the dairy industry because high-rank sires possess many other favorable genes. Selective mating (i.e., avoidance of carrier × carrier matings) would be an effective and less costly way to manage the defect. In addition, screening all young bulls for the BS gene and culling carriers prior to progeny test would be a desirable way to reduce the recessive allele frequency in the Chinese Holstein population.
Footnotes
Acknowledgements
The authors thank Prof. J. S. F. Barker for his English editing help on this article.
a.
LA PCR buffer, LA Taq polymerase; Takara Biotechnology Co Ltd, Dalian, China.
b.
Mastercycler, Eppendorf Inc., Germany.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by China Agricultural Research System (CARS-37), the National Key Technologies R & D Program (2011BAD28B02, 2012BAD12B01), the Program for Changjiang Scholars and Innovative Research in University (IRT1191), and Beijing Innovation Team of Technology System in the National Dairy Industry.
