Mycoplasmd bovis is potentially the most economically important Mycoplasma species in cattle, with cost estimates to the U.S. cattle industry reaching $108 million per year.
18
Clinical signs include mastitis, vaginitis, and abortions in adult cattle as well as pneumonia, septic polyarthritis, otitis media, and conjunctivitis in calves.
5,16,20,22
Prevalence of M. bovis is often underestimated in both beef and dairy herds because of the difficulties in diagnosis and the variety of clinical signs associated with this infection and disease caused by this pathogen.
15
Bacterial isolation, considered the gold standard test for mycoplasmal infections, is difficult and time consuming because of the fastidious nature of mycoplasmas, which require 1) specialized media and incubation conditions; 2) extended culture time as a result of the slow growth of the organism (up to 14 days for completion); and 3) additional testing for speciation.
10
Other respiratory pathogens are often isolated more easily and quickly, and attempts at detection and identification of M. bovis may not be pursued.
8,12
The microscopic features associated with this bacterium can resemble those caused by other respiratory agents, including Mannheimia haemolytica and Haemophilus somnus (now Histophilus somni).
9
Published polymerase chain reaction (PCR) techniques to detect M. bovis include gel-based and nested methods using a variety of targets.
2,7,9,13
A real-time PCR method based on 16S ribosomal (r)RNA has been demonstrated to be very effective for the detection M. bovis in milk and lung tissue samples, but the assay is designed specifically for use with one brand of thermocycler,
a
which limits its usefulness.
4
Constitutive or “housekeeping” genes that encode for products that are necessary bacterial replication and survival are generally highly conserved and consequently are good candidates for genetic detection of the agent.
19
The deoxyribodipyrimidine photolyase uvrC gene, which is part of the excision DNA-repair system Uvr ABC, has been shown to be highly conserved and therefore may prove useful for M. bovis identification.
3
The aims of the present study were 1) to evaluate a real-time PCR assay based on the uvrC gene target that could be completed without the requirement for a specific real-time PCR platform; and 2) to identify a more rapid and sensitive M. bovis diagnostic method than current culture techniques, which could be more universally applied in diagnostic settings.
Validation testing was performed on bovine clinical specimens submitted to the Iowa State University Veterinary Diagnostic Laboratory (ISU VDL) from 2005 to 2007. Three hundred eighty-five samples submitted for Mycoplasma culture during this time period were selected for comparison testing by real-time PCR, including 154 lung tissues, 122 milk samples, 70 joint fluids, 26 nasal swabs, 8 bronchoalveolar lavage (BAL) fluids, 1 tracheal wash, and 4 ear swabs. Fifty-one of the lung tissue samples were also culture positive for other common respiratory pathogens, including M. haemolytica, H. somni, Pasteurella multocida, and Arcanobacterium pyogenes.
Samples for bacterial culture were prepared as described elsewhere.
21
Briefly, 1–2 g of lung tissue was homogenized in 2.5 ml of sterile 0.9% saline using a stomacher,
b
with the resultant fluid collected. Joint fluid and BAL were diluted with equal volumes of saline. Joint swabs, ear swabs, and nasal swabs were rinsed in 2.5 ml of sterile saline. Milk samples were tested undiluted.
Culture was carried out directly from the sample and after enrichment and was performed according to the procedure described previously.
21
For direct sampling, 0.1-0.2 ml of the milk or saline-sample mixture was plated onto Mycoplasma agar.
c
Culture plates were housed in a moisture chamber placed in an incubator at 35°C providing 3–5% CO2. To enrich the samples, 4–6 drops of the salinesample mixture or undiluted milk were added to a 3-ml Mycoplasma broth tube,
c
with 0.25 mg/ml of cefoperazone.
d
Tubes were incubated at 35°C with 3–5% CO2 for 48-72 hr (based on turbidity) and plated on Mycoplasma agar for incubation as above. Culture plates were examined under a dissecting microscope for evidence of Mycoplasma colony morphology every 48 hr for 14 days. Suspect colonies were stained with fluorescent antibodies as previously de-scribed.
6
Briefly, an impression smear of colonies was made on a glass slide; fixed in acetone at room temperature for 7 min; incubated in a prewarmed moisture chamber at 37° C for 1 hr with primary antibody; rinsed 2 times for 5 min each with sterile phosphate buffered saline (PBS); rinsed with distilled water; allowed to dry; incubated with fluorescein isothiocyanate-conjugated secondary antibody in a prewarmed moisture chamber at 37°C for 1 hr; rinsed 2 times for 5 min each with sterile PBS; rinsed with distilled water; allowed to dry; and observed under a microscope for the presence of fluorescence.
Immunohistochemistry (IHC) for M. bovis antigens in tissue samples was performed as described.
1
Briefly, tissue sections on slides were deparaffinized and hydrated with distilled water; placed in staining racks; quenched in 3.0% hydrogen peroxide; rinsed in distilled water; digested in 0.05% protease for 2 min; rinsed 3 times in deionized water; incubated at room temperature for 1 hr with 200 μl of primary anti- M. bovis antibody
e
; rinsed with Tris-buffered saline (TBS); incubated at room temperature for 10 min with 200 μl of secondary antibody; rinsed with TBS; incubated at room temperature for 5 min with 200 μl of 3,3'-diaminobenzidine
f
; rinsed with distilled water; and counterstained with an automatic counterstainer.
g
Primers and probes were designed using Primer Premier Software
h
to the uvrC gene of M. bovis (GenBank accession no. AF003959). Primers Mbov F2024: TCTAATTTTTT CATCATCGCTAATGC, Mbov R2135: TCAGGCCTTT GCTACAATGAAC, and probe Mbov uvrC: FAMAACTGCATCATATCACATACT-MGB were purchased from commercial sources.
i
Primers amplify a 112-base pair (bp) uvrC gene fragment and show specificity for M. bovis using GenBank BLAST analysis (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi). A 66-bp oligonucleotide was designed to serve as an internal control (IC) sequence to monitor each reaction for the presence of inhibition (Mbov IC: TCTAATTTTTTCATATCGCTAATGCTCTTGTAC ACACCGCCC-AGTTCATTGTAGCAAAGGCCTGA and Mbov IC Probe: VIC-CTTGTACACACCGCCCMGB). The IC is flanked by the same primer sequences as the uvrC target sequences used for M. bovis detection; however, the central probe sequence was based on the 16S rRNA sequence to avoid recognition by the M. bovis -specific probe.
Samples for real-time PCR testing were prepared as previously described.
4
Briefly, 1–3 g of tissue was mixed with 5 ml of sterile saline; homogenized with a stomacher; and resultant fluid was collected. Two hundred microliters of the tissue or saline homogenate, fluid sample, broth culture, or milk sample was used for DNA extraction using a commercial kit
j
employing the kit's protocol according to the manufacturer's instructions. All sample types were resuspended in 200 μl of elution buffer (10 mM Tris-Cl, 0.5 Mm ethylenediamine tetra-acetic acid; pH 9.0). Real-time PCR was performed in a 25-μl reaction volume containing 1X universal TaqMan master mix,
e
1 μM each primer, 0.15 μM Mbov uvrC probe, 0.05 μM Mbov IC probe, 1 μM Mbov IC, and 2.5 μl of DNA template. Reactions were performed in a real-time PCR instrument.
h
The PCR reaction was carried out using the following cycling parameters: polymerase-activation for 10 min at 95°C followed by 40 cycles of denaturing at 95°C for 15 sec, and annealing and amplification at 60°C for 1 min. Fluorescence was acquired at the end of each round of the annealing step (single mode) for both FAM (6-carboxyfluorescein; Channel A) and VIC (2′-chloro-7′-phenyl-1,4-dichloro-6-carboxyfluorescein; Channel B). Commercial analytical software
h
was used to analyze the real-time PCR results after amplification was complete.
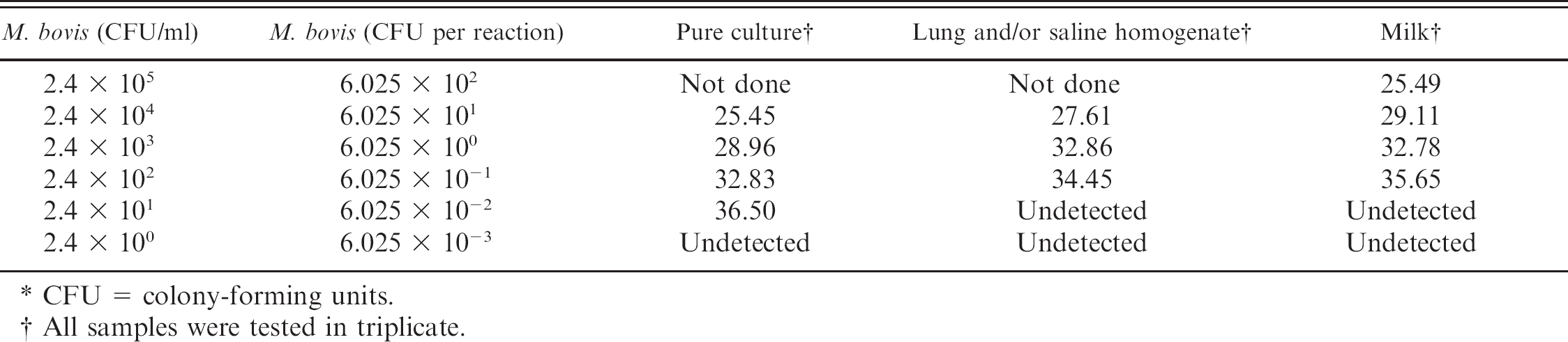
A suspension of pure culture as well as serial dilutions (2.4 × 100 to 2.4 × 105 colony-forming units (CFU)/ml) of M. bovis cells spiked in milk and lung tissue homogenate fluid were extracted and tested in triplicate to determine real-time PCR assay sensitivity. Bacterial species used to assess specificity of the real-time PCR assay included the following: 3 M. bovis strains (M23, Jasper, spm PM23 8/7 W23) and the positive extraction control; Mycoplasma alkalescenes, Mycoplasma arginini G230, Mycoplasma bovoculi C52, Mycoplasma bovirhinis 262i P13, Mycoplasma bovigenitalium, Mycoplasma dispar, Mycoplasma canadense 275C P4, Mycoplasma canis, Ureaplasma U233, Acholeplasma spp.; and 21 other bacterial species, including P. multocida, Streptococcus agalactiae, Streptococcus dysgalactiae, Arcanobacterium pyogenes, Streptococcus uberis, H. somni, M. haemolytica, Enterobacter spp., Enterococcus faecalis, coagulase-negative Staphylococcus spp., Staphylococcus aureus, Clostridium perfringens, Klebsiella pneumoniae, Serratia marcescens, group G Streptococcus, Proteus spp., Pseudomonas aeruginosa, Escherichia coli, and Morganella morganii. In addition, 19 clinically normal culture-negative lung samples were evaluated by the same real-time PCR method. Because of the small size of the PCR product, sequencing was not performed; however, size of the product was consistent among samples tested.
Sensitivity (Se), specificity (Sp), and kappa coefficients of agreement (κ) between the 2 test methods were calculated for the real-time PCR assay performance compared with results of culture on the various sample matrices using WinEpiscope 2.0.
k
Levels of agreement were determined using guidelines provided previously.
11
A subset of 54 cases in which lung and joint tissue samples had gross and histopathologic examinations, M. bovis culture, and IHC testing for M. bovis antigens were examined to more definitively assess disease status than culture alone. Samples with tissue damage consistent with M. bovis infections that were positive for M. bovis by both culture and IHC staining were defined as true positive samples; those with gross and histopathologic tissue lesions more consistent with other agents that were negative for M. bovis by culture and IHC staining were classified as true negative samples. These samples were tested by the real-time PCR assay, and Se, Sp, and K were determined, respectively.
Equal volumes of lung tissue or saline homogenate or milk and M. bovis broth culture were mixed, and colony counts were performed for use in assessing the detection limits. Results on broth culture samples and spiked matrices (saline from lung homogenate and milk) are listed in Table 1.
Concentrations of 2.4 × 102 CFU/ml were detected in all spiked samples tested. The assay showed no false-positive results when tested on culture samples from non-M. bovis bacterial species, including other common bovine bacterial respiratory pathogens. Thirty-two out of 109 (29.4%) of the M. bovis-positive validation lung tissues were infected with 1 additional pathogen; 7 out of 109 (6.4%) were infected with 2 additional bacterial species; and 2 out of 109, (1.8%) with 3 additional pathogens. Ten out of 45 (22.2%) M. bovis-negative samples used for validation were culture-positive for at least 1 other bacterial pathogen. All specimens containing other bacterial species showed complete agreement between M. bovis culture and real-time PCR results. Seven clinical specimens were colonized with other Mycoplasma species, including M. ovipneumoniae, M. arginini, and M. dispar; and all were M. bovis PCR negative.
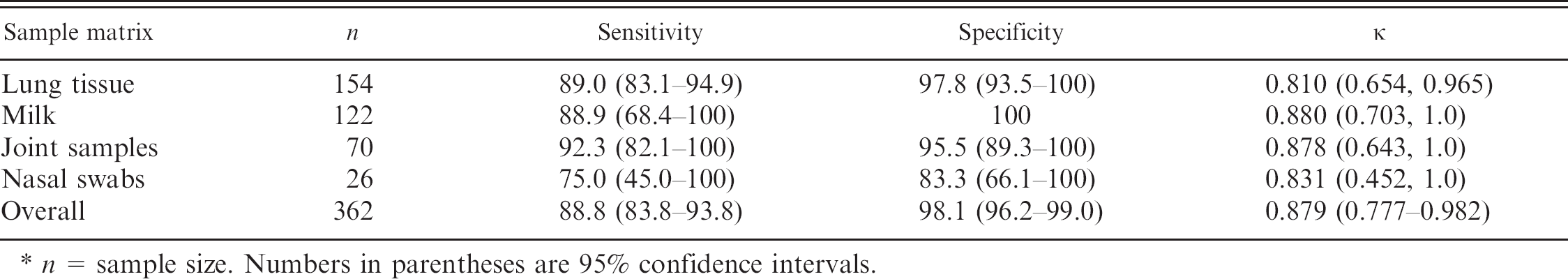
The Se, Sp, and κ based on bacterial culture accompanied by their 95% confidence intervals for the various matrices are presented in Table 2. Results on the most common matrices included lung tissue homogenate Se 89.0% (83.1-94.9%), Sp 97.8% (93.5-100), κ 0.810 (0.654-0.965, near-perfect agreement); milk Se 88.9% (68.4-100%), Sp 100%, κ 0.880 (0.703-1.0, near-perfect agreement); joint fluid/tissues Se 92.3% (82.1-100%), Sp 95.5% (89.3-100%), κ 0.878 (0.643-1.0, near-perfect agreement). Overall the real-time PCR assay demonstrated a Se of 88.8 (83.8-93.8), a Sp of 98.1 (96.2-99.0), and a κ of 0.879 (0.777-0.982; near-perfect agreement). Testing on the limited number of other matrices (BAL, tracheal wash, and ear swabs) showed near-perfect agreement with the real-time PCR assay used.
Results on the subset of 54 isolates with additional confirmation of disease status are listed in Table 3. Lung tissue samples comprised the majority of these sample types because other matrices often do not have accompanying tissue for IHC testing. Measures of assessment for lung homogenates were Se 97.4 (92.3-100), Sp 62.5 (29.0-96.0), κ 0.664 (0.379, 0.949); for joint samples, the measures of assessment were Se 80.0 (44.9-100), Sp 100, κ 0.696 (0, 1.0); and the overall measures of assessment were Se 97.7 (93.2-100), Sp 70.0 (41.6-98.4), κ 0.733 (0.466-1.0).
Coinfection and contamination with other bacterial agents is a common occurrence in clinical M. bovis infections and may interfere with Mycoplasma culture.
17
Testing of the real-time PCR target evaluated in the current study against other commonly recovered ruminant pathogens (P. multocida, M. haemolytica, H. somni, A. pyogenes) as well as tissues containing these agents demonstrated excellent specificity.
In addition to the requirements associated with M. bovis culture, several other difficulties can arise in establishing a definitive diagnosis of this agent in clinical samples. Mycoplasma bovis is frequently associated with the presence of other bacterial (M. haemolytica, H. somni, P. multocida) or viral pathogens (Bovine viral diarrhea virus, Infectious bovine rhinotracheitis virus, Parainfluenza virus type 3, and Bovine respiratory syncytial virus). Mycoplasma bovis can also be isolated from clinically healthy animals and from normal tissues, suggesting that its identification may not be sufficient to establish an etiology of disease. In addition, the inherent ability of M. bovis to alter its surface antigens can interfere with antibody recognition needed for accurate IHC confirmation and may account for the lower specificity seen in the samples with IHC diagnostic criteria.
13
Additional information such as clinical signs, gross and histopathologic tissue evaluation, and assessments for other infectious agents is needed to establish an association between agent infection and disease.
A review of cases submitted to the ISU VDL in 2009 was conducted to assess the utility of the real-time PCR assay based on the uvrC target. With the availability of a PCR diagnostic test, culture and IHC testing for M. bovis were performed infrequently in 2009; therefore, a case definition that included compatible clinical disease with gross and histopathologic evaluation of tissues was used to assess the value of the real-time PCR assay. The majority of samples tested were lung tissue (247/300, 82.3%), yielding Se of 81.8 (73.8-89.9), Sp of 85.7 (79.2-92.2), and κ of 0.675 (0.537, 0.814, very good agreement). Other matrices were tested infrequently and did not meet the established case definition.
Methods to improve the speed and accuracy of M. bovis identification will enhance the ability to recognize this agent in clinical disease and may provide insight into its role in pathogenesis.
14
The TaqMan-based assay described in the current study provides a rapid and sensitive method to identify M. bovis in field samples without the requirements for a specific thermocycler. The presence of an IC assured that any difficulties with reaction inhibition would be detected. The assay demonstrated similar sensitivity and specificity in a range of sample matrices to currently available diagnostic methods, including culture, but could be completed more quickly without the need for specialized culture media and conditions. As a real-time assay, it also eliminates the requirement of separate amplification and detection reactions associated with gel-based PCR techniques. Although the identification of this agent alone does not equate with disease etiology, rapid detection that can be accomplished through the real-time PCR assay described, which can be used on a variety of platforms, can expedite M. bovis identification and be a useful diagnostic aid.