Abstract
The use of swabbing to sample tissue samples, prior to nucleic acid extraction and performance of a real-time polymerase chain reaction (PCR) assay, was investigated for the detection of the viral pathogens
Tissue is a common sample taken for animal disease testing. When testing by polymerase chain reaction (PCR) assay, tissue samples are typically processed by dissecting a small section of tissue from the submitted specimen, mechanically disrupting the tissue (using methods such as freeze thawing and/or grinding) and digesting in lysis buffer prior to nucleic acid purification and subsequent PCR amplification. 8 Although well established, this process is time consuming, particularly when large numbers of samples need to be processed. In addition, even though complete digestion and lysis of tissue can occur after a few hours, many samples require overnight incubation to achieve this. This means that, for routine testing, laboratories often include a standard overnight tissue lysis step to allow samples to be processed together in an efficient manner, but even then, incomplete digestion of tissue can sometimes be an issue that can cause problems with downstream nucleic acid extraction steps. 8
In the current work, evaluation of an approach, originally developed for detection of equine influenza,
2
where swabbing of tissue is used as an alternative sampling method to the traditional tissue dissection and lysis approach described above, was undertaken. The evaluation was performed using 2 viruses that are important economically and for which PCR testing is commonly requested:
A panel of 100 tissue samples previously submitted to the Animal Health and Veterinary Laboratories Agency (United Kingdom) for routine PCR analysis was used in the current study. The samples had been tested for BVDV or PRRSV using the tissue lysis method and PCR and then stored at −20°C. The majority of the panel had tested PCR positive previously, but also included BVDV- and PRRSV-negative samples (38 positive and 12 negative for BVDV; 33 positive and 17 negative for PRRSV). Samples were representative of the tissue types typically submitted for PCR analysis from tissue (for BVDV: 39 spleen, 10 thymus, and 1 fetal tissue sample; for PRRSV: 38 spleen, 2 thymus, 7 lung, and 3 bronchial lymph node samples). The samples were allowed to thaw and were then sampled and tested using both the current tissue lysis method and the swabbing method, followed by PCR analysis.
For the current tissue dissection and lysis method, 1–10 mg of tissue was removed using disposable scalpels, added to a 1.5-ml microfuge tube, and then frozen at −70°C. Using dry ice or a cooled metal block to keep the sample cold, the sample was then disrupted using a sterile micropestle. a After this step, 200 μl of lysis buffer (prepared as described in the instructions of the commercial nucleic acid extraction kit b used) was added to the ground sample, and the tube was vortexed. Following incubation overnight at 60°C, 100 μl of the resulting tissue lysates were then processed on a nucleic acid extraction instrument c as described in the manufacturer’s instructions using a nucleic acid extraction kit b with an elution volume of 200 μl. The resulting nucleic acid extracts were then amplified using protocols based on established one-step real-time reverse transcription PCR assays for either BVDV 6 or PRRSV 3 using a real-time PCR instrument. d
To sample the tissue with swabs, 2–3 fresh cuts were made in each tissue sample using a sterile, disposable scalpel. Typically, cuts were 5 mm long and deep. A swab e was inserted into the cuts and agitated vigorously with a back and forth motion (typically 6 times) to disrupt the tissue. The same swab was used to visit each cut site. The head of the swab was then cut into a 3-ml bijou bottle containing 250 μl of 10 mM Tris–HCl and 1 mM ethylenediamine tetra-acetic acid buffer (pH 8.0). f The contents of the bijou were mixed by vortexing for 10 sec and then allowed to stand at room temperature for 10 min. After an additional vortexing for 10 sec, 100 μl of the liquid was mixed with 150 μl of lysis buffer (as described above) and incubated either overnight or 2 hr at 60°C. Following incubation, 100 μl of the resulting material was then processed using the same nucleic acid extraction instrument and nucleic acid extraction kit as described above.
In the first part of this study, the sampling method was compared (swabbing vs. tissue lysis) but the overnight digestion step was maintained for all samples. Nucleic acid extraction and the subsequent PCR assays were carried out in 3 separate experiments: 2 batches of 20 mixed positive and negative samples and 1 batch of 10 negative samples. Following extraction, all nucleic samples were stored at −80°C until used. Each PCR batch involved the making up of a new master mix but used the same batch of reagents and primers. The PCR assays were carried out in triplicate for the PRRSV assay but singly for BVDV (which reflects the current routine testing regime); an average of the threshold cycle (Ct) value for the PPRSV result was taken forward for analysis. One technician carried out all testing of batches 1 and 2 (extraction and PCR for both diseases); a different technician carried out all testing of batch 3.
The subsequent results (Figs. 1, 2) demonstrate that equivalent PCR results were obtained using the tissue and swabbing approaches for both BVDV and PRRSV. Of the 100 samples tested (38 positive and 12 negative BVDV samples; 33 positive and 17 negative PRRSV samples when tested using the tissue lysis method) only 1 BVDV sample showed a discrepant result, with swabbing failing to detect a tissue lysis method BVDV-positive sample. Unfortunately, there was insufficient material to retest this sample, meaning it is difficult to determine if this was a result of testing error or a genuine failure of the swabbing method. All negative samples tested negative with both methods.
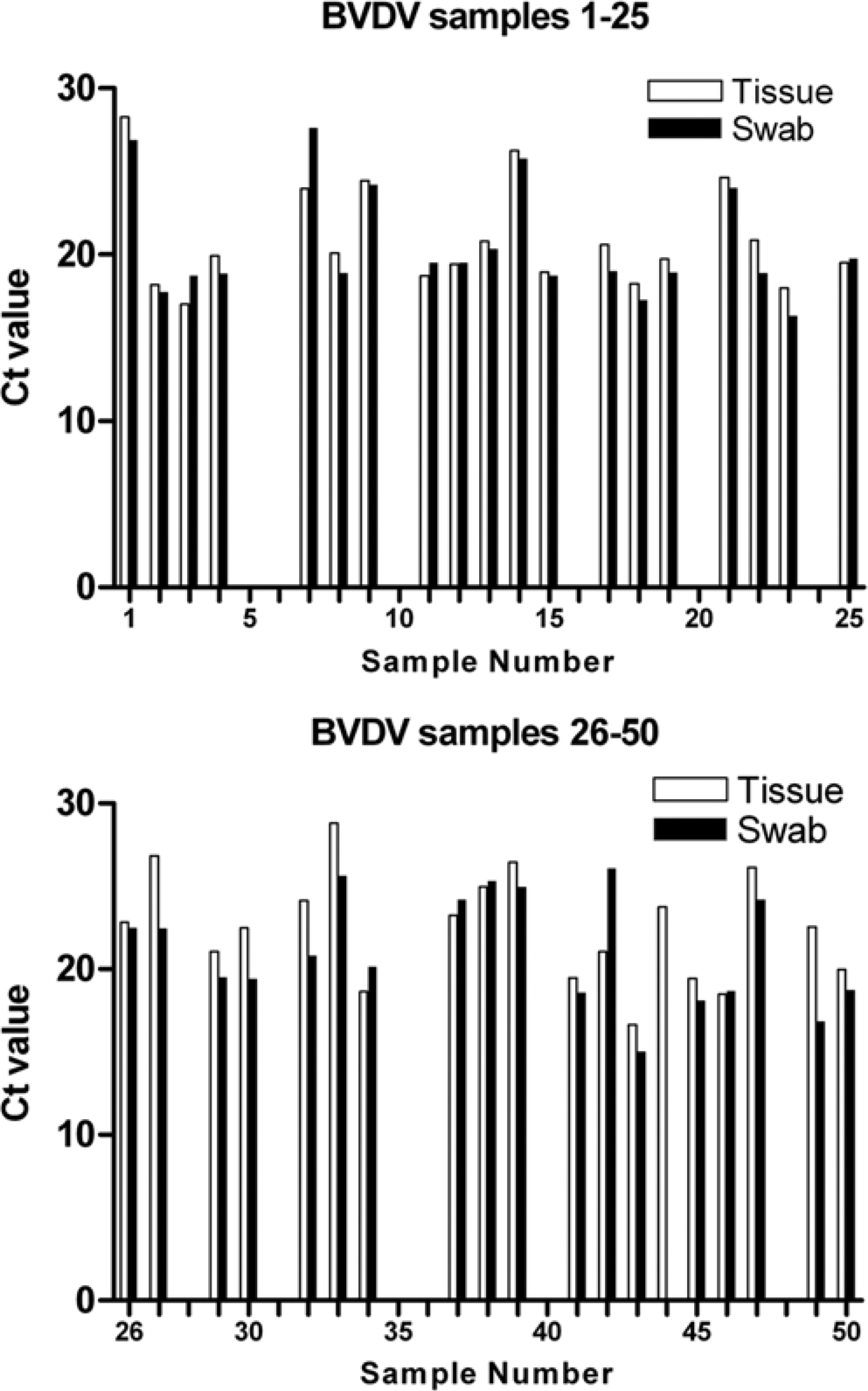
Swabbing produces equivalent results to tissue sampling for real-time polymerase chain reaction (PCR) detection of
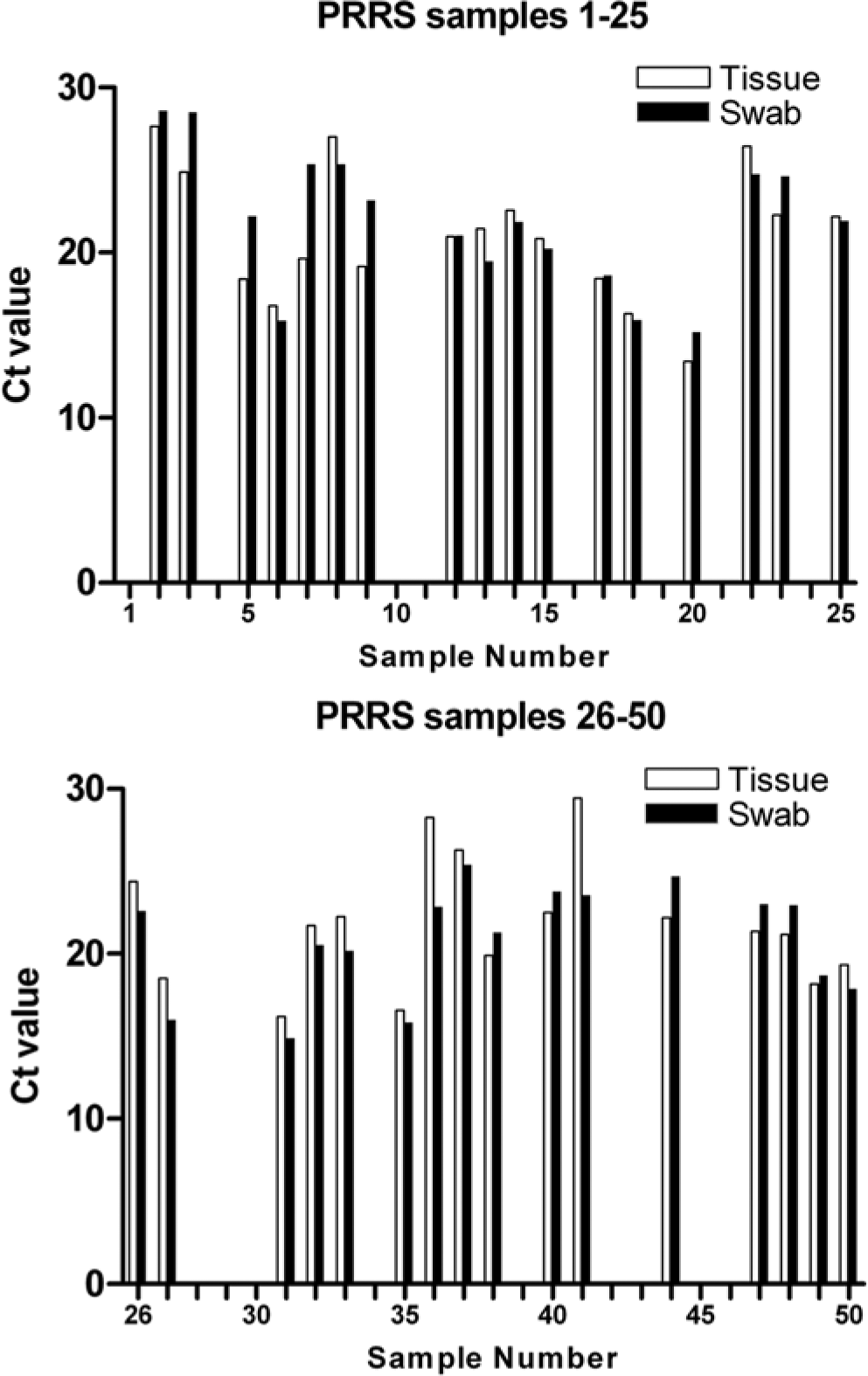
Swabbing produces equivalent results to tissue sampling for real-time polymerase chain reaction (PCR) detection of
Calculation of Cohen kappa coefficient (using the DAGStat program 5 ) using the tissue lysis method as the gold standard, and assigning swabbing results as positive or negative, produced values of 0.95 (95% confidence interval: 0.8–1.0) and 1 for BVDV and PRRSV, respectively, indicating excellent agreement between the 2 methods. In addition, there was good correlation of Ct values demonstrating that swabbing appeared to be capable of reflecting the viral load in a sample in an equivalent manner to traditional tissue sampling and lysis. The average difference between the Ct values when carried out on the same tissue sample using lysis and swabbing was 1.6 for BVDV and 1.9 for PRRSV demonstrating good agreement between Ct values obtained with the 2 methods.
In order to compare methods agreement across a range of Ct values, differences in the Ct value in a pair were plotted against their average (Fig. 3). 7 For PRRSV, there appeared to be no bias in which method produced the lower Ct value (indicative of more sensitive detection), with 54% of positive samples (18/33) producing a lower Ct with the swab method. Interestingly, for BVDV, there did appear to be a pattern of swabbing producing lower Ct values (74%; 29/39 samples) but confirmation of this would require further evaluation with higher numbers of swabs. There was a tendency for samples with higher Ct values to produce more variation between samples (lysis vs. swabs), but this is probably related to the PCR component of the assay where it is recognized that lower levels of target nucleic acid lead to greater variation in PCR results. 9
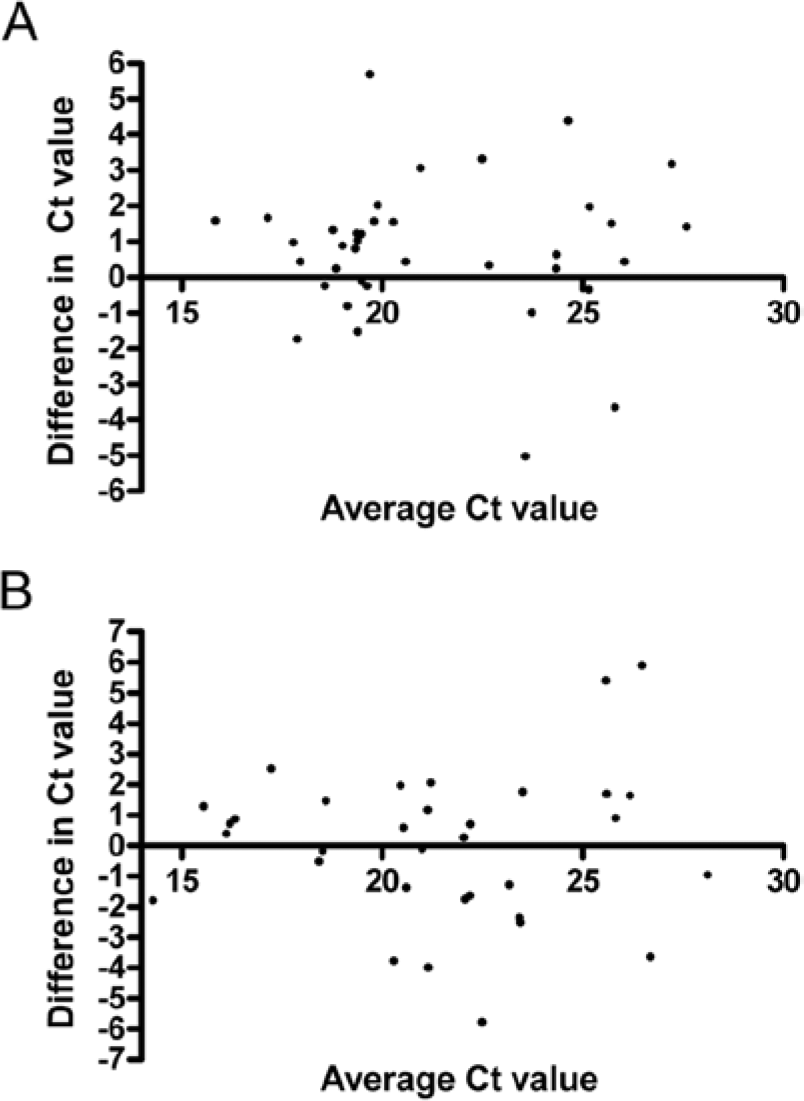
The difference between the threshold cycle (Ct) values obtained using the tissue lysates and tissue swabs (L – S) on the same samples plotted against their average (L + S)/2 to visualize method agreement.
Swabbing tissues offers a technically more straightforward method than dissection for nucleic acid testing for viral pathogen detection. Although superficially similar, the 2 approaches are actually very different in terms of hands-on time and ease of use. For example, dissecting small pieces of tissue, freezing the tissue, and then individually disrupting them is time consuming. This contrasts with the rapid sampling offered by the swabbing approach. The use of swabs may also provide useful advantages for pooling of samples, as swab eluates can easily be combined for testing. Swabbing may also be useful for sampling larger areas than is practical using tissue sampling. In the current study, wire-stemmed swabs with a rayon viscose bud were used. However, it is possible that alternative swabs may lead to improved results and this could be an area for further evaluation.
Following the results described above, preliminary experiments were also carried out to explore if use of the swabbing method would allow for use of a 2-hr lysis step compared with the overnight lysis in the original method. This shortened lysis step could be advantageous in diagnostic situations where rapid turnaround of samples was required, potentially allowing same-day testing of tissue samples. Using BVDV to test this approach, an additional small panel of 11 positive samples (9 spleen and 1 thymus sample) was tested with the current approach (tissue dissection and overnight 60°C lysis) and the swabbing method using both a 2-hr and overnight lysis. For each swabbing event, separate swabs were used, and separate cuts were made in the same tissue sample. Following lysis, samples were processed immediately (after 2 hr or overnight lysis) using the same commercial DNA extraction kit and instrument as described above. The use of swabs and a 2-hr lysis resulted in successful detection of all samples (Fig. 4); in fact, the lower Ct values seen in all samples indicated more sensitive detection. This is an intriguing finding and suggests that overnight incubation may be resulting in reduction in sensitivity, perhaps because of virus nucleic acid degradation. By removing the need for protracted tissue lysis, and allowing more rapid processing of samples, the use of swabs may allow an improvement in test sensitivity. More work is needed to confirm these findings and build on these preliminary results, including the testing of negative samples.
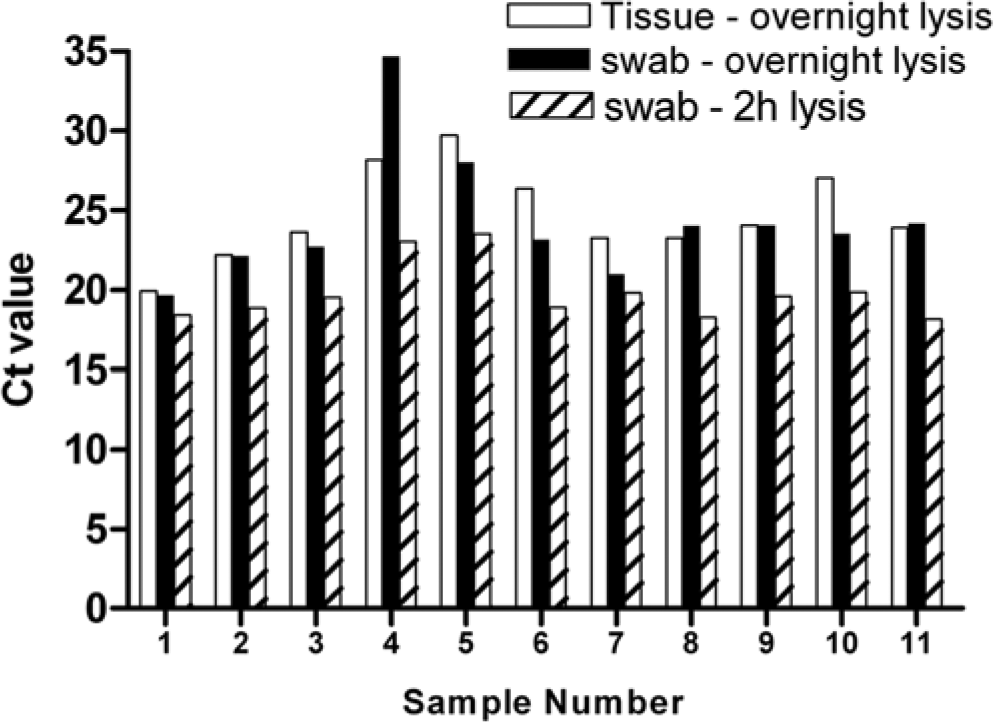
Swabbing combined with a 2-hr lysis step produces equivalent results to tissue sampling and swabbing with an overnight lysis step for real-time polymerase chain reaction (PCR) detection of
The data presented in the current study suggests that tissue swabbing offers an interesting alternative to the traditional tissue lysis approach when using PCR to test tissues for the presence of 2 common veterinary pathogens. Further work will be necessary with larger numbers of samples before the approach could be adopted for routine use or for other viruses. However, the approach appears to be promising and may prove useful to laboratories facing the day-to-day challenges of routine veterinary testing.
Footnotes
Acknowledgements
The authors thank Mats Isaksson from the Swedish National Veterinary Institute, Uppsala, for raising the possibility of this approach and providing an initial protocol and Marie Lightfoot, Nick Torrens and Joe Kirk for expert technical assistance.
a.
Anachem Ltd, Luton, Bedfordshire, United Kingdom.
b.
DNA II tissue kit (external proteinase K protocol), Roche Diagnostics Ltd, Burgess Hill, United Kingdom.
c.
MagNAPure Classic instrument, Roche Diagnostics Ltd, Burgess Hill, United Kingdom.
d.
MX3000P, Agilent Technologies UK Ltd, Wokingham, Berkshire, United Kingdom.
e.
Dryswab ENT, Medical Wire and Equipment, Corsham, United Kingdom.
f.
Promega UK Ltd, Southampton, Hampshire, United Kingdom.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was supported by funds from the AHVLA Test Development Programme, Project RD0015.
