Abstract
There has been a developing interest in the use of oral fluid for the diagnosis of different pathogens such as Porcine reproductive and respiratory syndrome virus (PRRSV). PRRSV and PRRSV-specific antibodies have been shown to be present in oral fluid samples, but the correlation between diagnostic results in oral fluid and serum samples has been insufficiently addressed. Studies investigating this correlation focused on boars older than 6 months and type 2 strains, but it is known that the outcome of a PRRSV infection is age and strain dependent. To address this gap, the current study reports on the detection of PRRSV and PRRSV-specific antibodies in serum and oral fluid samples collected over a 6-week period after an experimental infection of 8-week-old individually housed pigs with Lelystad virus, the type 1 prototype strain. Quantitative reverse transcription polymerase chain reaction analysis showed that significantly more serum samples were PRRSV RNA–positive than oral fluid until 5 days postinfection (dpi). Between 7 and 21 dpi, PRRSV RNA detection was similar in both samples but higher detection rates in oral fluid were found from 28 dpi. Compared with existing literature, this highlights that detection rates at particular time points postinfection might vary in function of strain virulence and animal age and provides useful information for the interpretation of pen-based oral fluid results. An excellent agreement between the oral fluid and serum enzyme-linked immunosorbent assay results was observed at every time point, further supporting the usefulness of oral fluid as a diagnostic sample for antibody detection.
Keywords
Introduction
Porcine reproductive and respiratory syndrome virus (PRRSV; order Nidovirales, family Arteriviridae, genus Arterivirus) is the etiologic agent of PRRS, a chronic, infectious disease of swine that, despite the availability of different types of vaccines, is still difficult to control. 8 In Europe, PRRSV is endemic in nearly all countries with major pig production, whereas countries with smaller swine stocks such as Finland, Norway, Sweden, and Switzerland are considered PRRSV free. 18 PRRSV is differentiated into 2 genetically distinct genotypes: type 1, or European genotype, and type 2, or North American genotype. Based on DNA sequencing of open reading frame (ORF)5 and ORF7, 3 genetic subtypes are recognized in type 1 PRRSV, and the presence of a fourth subtype has been proposed. 26 In Western and Central Europe, including Belgium, only subtype 1 (Lelystad virus–like) is circulating, while all subtypes are present in Eastern Europe. 26 Type 2 PRRSV has been reported sporadically in Europe, and most isolates share a high degree of identity with the modified live vaccine strain that is based on the type 2 VR2332 isolate.15,19,26
On natural infection via the respiratory route, the virus has a specific tropism for cells of the monocyte–macrophage lineage. 7 After replication in local macrophages, the virus rapidly spreads to lungs and lymphoid organs. 2 Other tissues may also be infected, but not as commonly. Viremia develops in pigs 12–24 hr postinfection and remains detectable for approximately 20–30 days after infection.2,7,28 In the oropharyngeal region of growing pigs, however, PRRSV and viral RNA have been detected for several months (up to 251 days postinfection [dpi]). 28 PRRSV infection passes often unnoticed because many animals do not show clinical signs, and infection may only be detected during diagnosis by the presence of viral nucleic acids or anti-PRRSV antibodies in serum samples. 6
Oral fluid collected with a rope has been described as an alternative diagnostic sample for PRRSV detection 22 and has the advantages that it is less intrusive than blood sampling and that samples can be collected at group level from pigs that are housed in pens. Therefore most studies have focused on pen-based oral fluid samples, and only a few of the studies systematically compared the diagnostic sensitivity in serum samples versus oral fluid samples.13,20,24,25 Furthermore, most studies investigated the sensitivity of PRRSV detection in oral fluid samples after infection with type 2 field strains8,21,24 while only a limited number of studies have focused on infection with type 1 strains.8,14 Despite the fact that studies have been performed in individually housed boars older than 6 months,8,14,21 the correlation between diagnostic results in serum and oral fluid of individual animals at different time points postinfection is currently insufficiently addressed. Such correlation is, however, important to allow a better understanding and interpretation of results obtained during pen-based sampling. Therefore, it was our objective to compare the detection of PRRSV via enzyme-linked immunosorbent assays (ELISAs) and quantitative reverse transcription polymerase chain reaction (qRT-PCR) tests in serum and oral fluid samples collected from individually housed animals over a 6-week time period after infection with Lelystad virus, the type 1 prototype PRRSV strain.
Materials and methods
Virus strain
Lelystad virus, the type 1 prototype PRRSV strain (fourth passage), was cultivated and titrated on Marc-145 cells as described previously. 3 Aliquots of 1 × 105 50% tissue culture infectious doses (TCID50)/ml were stored at −80°C.
Animals, inoculation, and sample collection
Sixteen Belgian Landrace piglets (average age of 8 weeks), without PRRSV immunoglobulin (Ig)G antibodies as tested via ELISA a and negative for PRRSV RNA via an ORF7-based qRT-PCR, b were used in the current study. Animals were divided into 2 groups (group A, n = 10; group B, n = 6) and were housed individually at air-filtered level-2 biosecurity facilities (CODA-CERVA, Machelen, Belgium). During 1 week of acclimatization, the animals were trained to chew on ropes that were soaked with apple juice. Thereafter, each piglet from group A was manually restrained and challenged intranasally with 2 ml of viral inoculum containing 1 × 105 TCID50/ml of the PRRSV Lelystad strain (1 ml in each nostril). Pigs from group B were mock infected with phosphate buffered saline and kept as negative controls.
Blood and oral fluid samples were taken in the morning on days 0, 3, 5, 7, 10, 14, 21, 28, 35, and 42 postinfection (dpi) for evaluation by qRT-PCR and ELISA. Oral fluid sampling was conducted before blood sampling to reduce the chance that animals would refuse to bite on the ropes after physical manipulation. Blood was collected by jugular venipuncture and centrifuged at 1,000 × g for 10 min; the serum was then aliquoted and stored at −80°C until assayed. At each time point, oral fluid samples were first collected with nontreated cotton ropes c and subsequently with polyester ropes c to evaluate a) potential differences between ropes in retention of virus and/or viral RNA or b) if the presence of potential inhibitors of the downstream PCR reaction would influence PRRSV RNA detection. In brief, 1 rope (length: 1 m; diameter: 14 mm) was suspended in each pen and was left in place for 30 min, during which the animal could chew on it and moisten it with oral fluid. To recover the oral fluid samples, ropes were manually wrung, and the oral fluids were collected in 50-ml conical centrifuge tubes. d Oral fluid samples were immediately chilled on ice, centrifuged at 1,800 × g for 10 min at 4°C, and stored as aliquots at −80°C until use. Animal experiments were performed in accordance with the guidelines of European Union and Belgian regulations on animal welfare in experimentation, under the supervision of the Ethical Welfare Committee of the IPH-VAR, Belgium (procedure agreement no. 120112-01).
Detection of PRRSV by quantitative reverse transcription PCR
PRRSV RNA was detected via a qRT-PCR that targets an ORF7 fragment. Briefly, total RNA from serum samples was extracted e according to the manufacturer’s protocol. RNA from oral fluid was extracted as previously described. 2 Following extraction, the RNA was amplified b in a 25-µl reaction mixture containing 12.5 µl of 2× multiplex RT-PCR buffer, 2.5 µl of 10× PRRSV primer probe mix, 1.25 µl of multiplex enzyme mix, 0.75 µl of nuclease-free water, and 8 µl of extracted RNA. All PCRs were run on a real-time PCR system. f For each assay, positive and negative control samples were tested with the unknowns. Samples with a threshold cycle (Ct) value less than 37 were considered positive.
Detection of PRRSV-specific antibodies by ELISAs
Commercially available ELISAsa,g were used to evaluate the presence of PRRSV-specific IgG antibodies in serum and oral fluid samples, respectively. Both assays were performed according to the manufacturer’s instructions. ELISA results were expressed as sample-to-positive (S/P) ratios (optical density [OD] value of sample – OD value of negative control)/(OD value of positive control – OD value of negative control). Samples that had an S/P value equal or greater than the cutoff value (0.40) were considered positive for PRRSV-specific IgG antibodies.
Statistical analysis
Two-sided Fisher exact tests were used to evaluate differences in PRRSV detection ratios in serum and oral fluid by qRT-PCR at each time point. The Ct values associated with the peaks of PRRSV RNA detected in serum and oral fluid were compared using a Student t-test. The agreement between the detection of PRRSV-specific antibodies in serum and saliva by ELISA was measured using Cohen kappa coefficient. The following ranges were used for interpretation of the kappa coefficient 16 : poor agreement: 0.00; slight agreement: 0.00–0.20; fair agreement: 0.21–0.40; moderate agreement: 0.41–0.60; substantial agreement: 0.61–0.80; almost perfect: 0.81–1.00. The independent samples t-test was used to evaluate differences in the mean number of samples collected per animal and per rope type between PRRSV-infected and mock-infected animals. Data was analyzed using commercial h software, and P values <0.05 were considered to be significant for all tests.
Results
Sample collection
A total of 160 serum samples, 60 oral fluid samples collected with cotton ropes, and 45 oral fluid samples collected with polyester ropes were obtained at different time points after infection. Two animals from the PRRSV-infected group (pigs 7 and 8) showed no interest in the presented ropes. Five animals (2 from the infected group and 3 from the control group) showed preference for either cotton (pigs 6, 15, and 16) or polyester (pigs 4 and 11) ropes (Table 1). The success rate of oral fluid collection by ropes from individually housed piglets of 8–14 weeks old was variable, with successful oral fluid collection of 37.5–87.5% at a particular time point. The mean number of samples collected per animal did not significantly differ between PRRSV-infected and mock-infected pigs for cotton ropes (meaninfected, mock infected = 3.6, 4.0; P = 0.792) or for polyester ropes (meaninfected, mock infected = 3.3, 2.0; P = 0.474), indicating that PRRSV infection did not influence the biting behavior of the pigs. One sample from the cotton rope–collected oral fluids and 2 samples of the polyester rope–collected oral fluids released insufficient oral fluid to perform qRT-PCR. However, all oral fluid samples could be tested by ELISA.
Successful oral fluid collections by ropes from individually housed piglets (nos. 1–16) aged 8–14 weeks.

The results represent the number of successful collections on 10 different sampling days.
Detection of PRRSV by quantitative reverse transcription PCR
When the qualitative qRT-PCR results for oral fluid obtained by both cotton and polyester ropes were compared for pigs that had consecutively chewed on both ropes (n = 22), no differences were observed. These results are presented in Table 2. Based on serum PCR results, all inoculated animals developed viremia within 3 days after PRRSV inoculation. Viremia lasted for 14–28 days, with 100% positive animals from 3 to 14 dpi, 30% at 21 dpi, and only 10% at 28 dpi.
Qualitative results of Porcine reproductive and respiratory syndrome virus (PRRSV) detection by quantitative reverse transcription polymerase chain reaction in serum and oral fluid samples of individually housed pigs, intranasally inoculated with the Lelystad strain (European prototype PRRSV).*
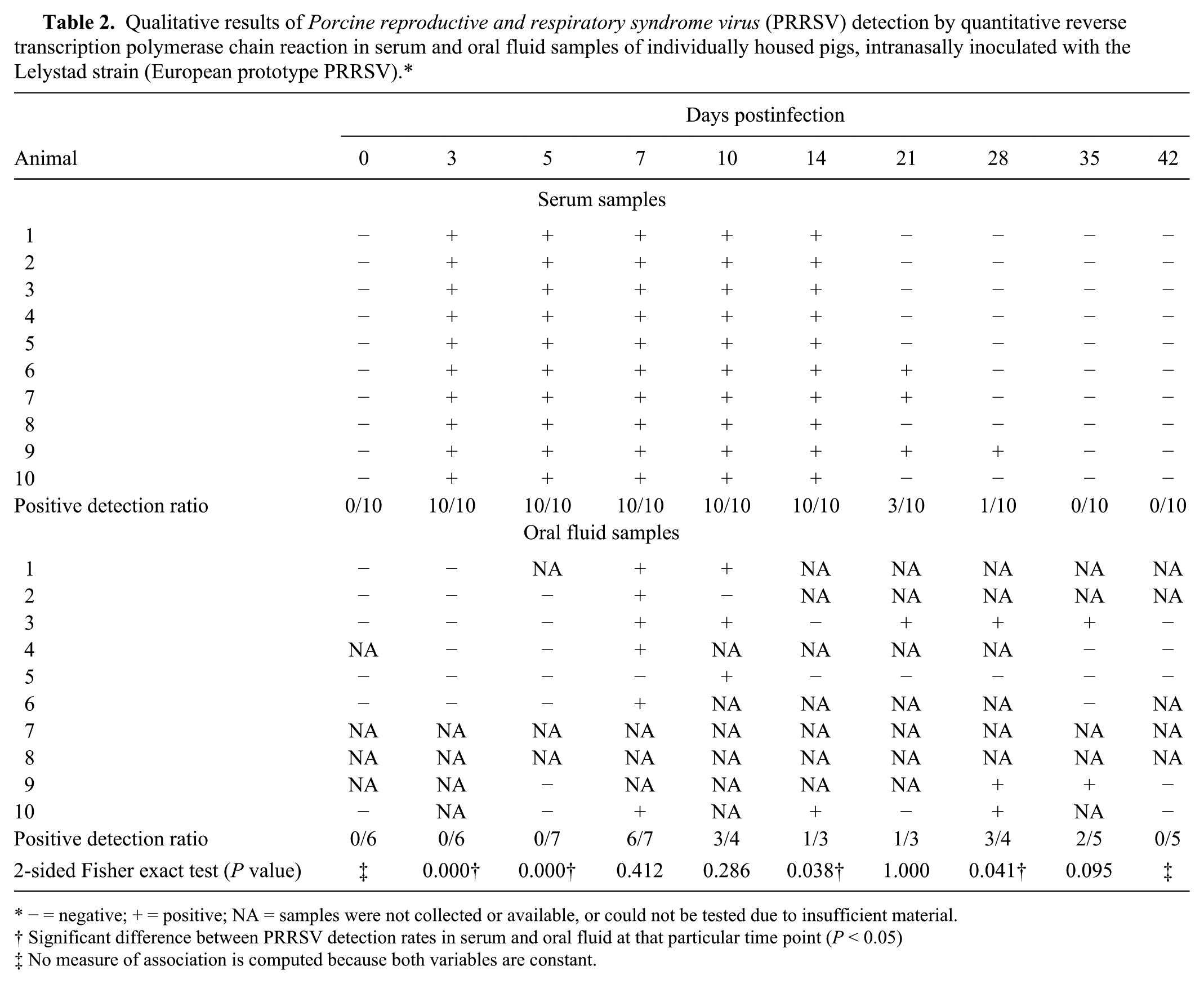
− = negative; + = positive; NA = samples were not collected or available, or could not be tested due to insufficient material.
Significant difference between PRRSV detection rates in serum and oral fluid at that particular time point (P < 0.05)
No measure of association is computed because both variables are constant.
At each time point, at least 3 and mostly between 5 and 7 oral fluid samples were collected that could be tested by PCR. Presence of viral RNA in oral fluid samples could be detected in inoculated pigs from 7 to 35 days after inoculation. PRRSV RNA, however, was not consistently detected in oral fluid of all animals during that time period: detection ratios varied from 85.7% at 7 dpi to 75% at 10 dpi, 33.3% at 14 and 21 dpi, 75% at 28 dpi, and 40% at 35 dpi. Fisher exact tests showed that significantly more serum samples were PRRSV positive by qRT-PCR than oral fluid samples at 3, 5, and 14 dpi while the opposite was true at 28 dpi. At day 7, 10, 21, and 35, there were no significant differences in the detection rate in serum and oral fluid. The P value of 0.095 at 35 dpi suggests, however, that PRRSV might be detectable for a longer amount of time in oral fluid than in serum (Table 2).
Quantitative qRT-PCR results (Fig. 1) showed that a peak amount of PRRSV RNA was detected at 5 dpi in serum and at 7 dpi in oral fluid. The mean Ct values at those peaks were not significantly different (P = 0.36), indicating that similar amounts of PRRSV RNA were present. In oral fluid, the detected amount of PRRSV RNA in positive samples seemed to decrease slowly until no more positive samples were found at 42 dpi. The Ct values in serum samples showed more fluctuation. Although not enough positive oral fluid samples were present to allow a statistical comparison, comparable amounts of PRRSV RNA seemed to be present in positive oral fluid and serum samples at most time points postinfection when viral RNA was detected in both samples. When oral fluid was successfully collected from a particular animal with both cotton and polyester ropes, the Ct values of positive animals were continuously approximately 1 Ct value higher in oral fluid–collected polyester ropes than with cotton ropes (data not shown), suggesting more virus or viral RNA is retained by polyester than by cotton. No PRRSV RNA was detected in any of the control animals during the experiment.
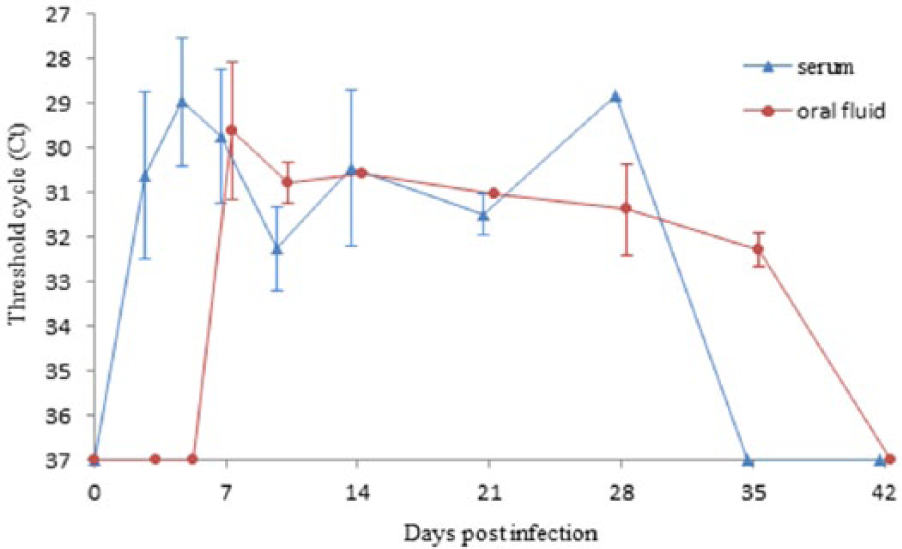
Mean (± standard deviation) threshold cycle values of quantitative reverse transcription polymerase chain reaction Porcine reproductive and respiratory syndrome virus–positive serum and oral fluid (collected with cotton ropes) at different time points postinfection.
Detection of antibodies directed against PRRSV by ELISAs
When the qualitative ELISA results for oral fluid obtained by both cotton and polyester ropes were compared for pigs that had successively chewed on both ropes (n = 23), only 1 discordant result (animal 5, day 10) was found. The qualitative results are presented in Table 3.
Detection of Porcine reproductive and respiratory syndrome virus (PRRSV)-specific immunoglobulin G antibodies by enzyme-linked immunosorbent assay in serum and oral fluid samples collected by cotton and polyester ropes of individually housed pigs, intranasally inoculated with the Lelystad strain (European prototype PRRSV).*
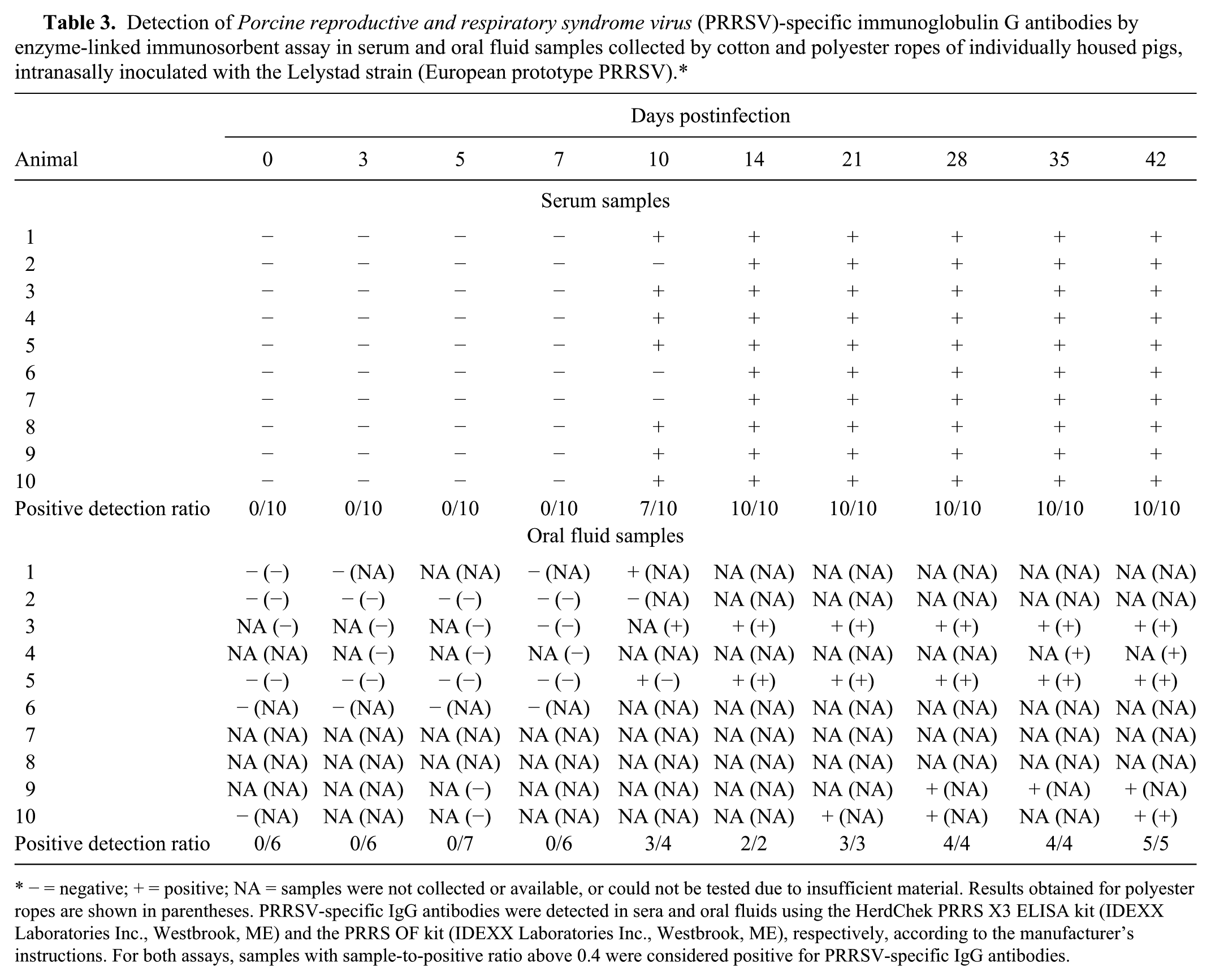
− = negative; + = positive; NA = samples were not collected or available, or could not be tested due to insufficient material. Results obtained for polyester ropes are shown in parentheses. PRRSV-specific IgG antibodies were detected in sera and oral fluids using the HerdChek PRRS X3 ELISA kit (IDEXX Laboratories Inc., Westbrook, ME) and the PRRS OF kit (IDEXX Laboratories Inc., Westbrook, ME), respectively, according to the manufacturer’s instructions. For both assays, samples with sample-to-positive ratio above 0.4 were considered positive for PRRSV-specific IgG antibodies.
Seven of the 10 inoculated pigs became serologically positive by ELISA by 10 dpi. The remaining infected pigs seroconverted by 14 dpi and remained seropositive for the duration of the experiment (Table 3). Similar results were found in oral fluid as PRRSV antibodies were first detected at 10 dpi (3/4), and all animals were positive from 14 dpi until the end of the study. The control pigs were seronegative for antibodies against PRRSV throughout the study.
The sensitivity of the oral fluid ELISA was determined relative to the positive ELISA results in serum from the PRRSV-infected animals (ncotton = 19; npolyester = 15). Eighteen out of 19 and 14 out of 15 seropositive pigs were positive in oral fluid samples collected with cotton ropes and polyester ropes, respectively, corresponding with a relative sensitivity of 94.7% and 93.3%. To determine the specificity of both ELISAs, serum samples and oral fluid samples from the mock-infected control animals were used. Because all control piglets tested continuously negative, results in serum and oral fluid were 100% concordant, and the specificity of both tests was 100%. The overall agreement beyond chance between results in oral fluid and serum, expressed by kappa values, was excellent with kappa values of 1.00 for cotton-derived samples (p < 0.0001; 95% CI: 1.00, 1.00) and 0.95 for polyester-derived samples (p < 0.0001; 95% CI: 0.90, 1.00) when all samples were included (ncotton = 60; npolyester = 45).
Discussion
Rope sampling is a noninvasive method of oral fluid collection from pigs that is being used for the detection of several infectious agents including PRRSV.4,17,23,24 The oral fluid collection from individual young piglets showed, however, not to be straightforward and was largely dependent on individual animal behavior. Although the animals were trained to chew on the ropes in 1 week, and oral fluid sampling was conducted before physically manipulating the animals, in the current study, only 2 pigs chewed at every time point, while 2 other pigs would not chew on the ropes throughout the entire experiment. The low success rate in oral fluid collection was unexpected and is probably related to the young age of the pigs, as a previous study described 97% successful oral fluid collections over a 21-day study period in animals of 6 months of age. 14 This is supported by studies showing that young animals tend to use enrichment objects, such as ropes, to a lesser extent 5 and that older pigs seem to approach novel objects more quickly. 27 Also, the individual housing of the piglets is most probably another influencing factor that explains the low oral fluid collection rate because animals held in a group are more motivated to perform exploratory behavior together. 5 The fact that no clinical signs were observed on infection with the PRRSV Lelystad strain and that no differences in willingness to chew on the ropes were found between PRRSV-infected and control animals indicates that infection does not influence the number of collected samples at particular time points. However, this finding could be different for infections with more virulent strains (e.g., the type 1 subtype 3 Lena strain, 11 known to induce high fever, anorexia, depression, and even mortality).
The qRT-PCR results showed that PRRSV RNA could only be detected in oral fluid samples from 7 dpi while all animals were already positive in serum at 3 dpi. This differs from previous studies in individual animals where PRRSV was already detected in oral fluid in 10–60% of the infected animals at 1 dpi.8,13 In both previous studies, however, 6-month-old boars were inoculated with highly virulent strains. The contradictory results can most likely be explained by differences in virulence between strains used because it has already been demonstrated that more virulent strains replicate faster (beginning before day 3 after infection) and reach a peak on day 7 while attenuated isolates show a moderate increase (beginning after day 3) and reach a lower peak titer between days 7 and 15.9,10 This hypothesis is reinforced by other studies showing that PRRSV infections with type 1 strains are associated with a delayed acute infection of lung macrophages with peak amounts of positive cells only at 2–3 weeks postinfection, 2 while infections with highly virulent isolates8,9 result in high numbers of infected alveolar macrophages at 3 dpi. Our results also suggest that the PRRSV RNA detected in oral fluid most likely does not originate from virus circulating in serum, but rather from virus that has replicated in the lung or other local tissues.
Once PRRSV was detected in oral fluid at 7 dpi, similar detection rates in oral fluid and serum samples were found until 21 dpi, with the exception of 14 dpi. At 14 dpi, the detection rate dropped to 33.3% in oral fluid where it increased again to 75% at 28 dpi. A similar pattern has been described earlier on PRRSV infection of individual boars. 8 Although interesting, no straightforward explanation for this observation can be provided at this moment. It should, however, be taken into consideration that only a limited amount of samples were gathered at 14 dpi.
Also remarkable were the higher detection rates in oral fluid at 28 and 35 dpi compared with serum samples. Again, no conclusive explanation about the origin of the viral RNA in oral fluid at that time point can be given, but the reported positive lung macrophages found 2–3 weeks after infection with type 1 isolates 1 lead to the hypothesis that these might account for the positive results. Yet, replication in local tissues such as salivary glands and tonsils can at this moment not be excluded as a potential source of PRRSV in oral fluid samples.
Overall, our PCR results suggest that, depending on the viral strain, oral fluid might not be an ideal sample when the early detection of PRRSV in individually housed animals is targeted, but it does allow for a prolonged detection period. Extrapolation of these results to oral fluid sampling of group-housed pigs, being the main intent of the technique, suggests that circulating PRRSV could be detected for a longer period of time via monitoring of oral fluid than via serum collected from individual animals. To correctly perform such an extrapolation, the minimal number of PRRSV-infected animals per pen necessary to allow virus detection in a pen-based oral fluid sample remains to be determined.
Comparison of ELISA results of serum samples and matched oral fluid samples showed that both ELISAs produce highly similar results. Because both tests detect anti-PRRSV antibodies of the IgG subtype, the good agreement between both tests provides an extra indication that IgG in oral fluid originates primarily from the passive transfer of IgG antibodies from serum to oral fluid. 2 These results in individual animals reinforces conclusions of previous studies 12 suggesting that oral fluid is a suitable and easy-to-collect sample to monitor the infection status of a herd by evaluating the antibodies in oral fluid samples.
The current experimental study shows that oral fluid holds potential as a diagnostic sample to detect type 1 PRRSV strains in individually housed pigs. Highly comparable ELISA results were obtained in oral fluid and serum, showing that oral fluid can be used as a simple, rapid, and reliable detection of PRRSV-specific antibodies and might be a useful sample to monitor PRRSV antibody status of individually housed animals. While some studies already have shown that this is true for samples collected at group level for type 2 PRRSV strains, it would also be useful to validate this claim for type 1 strains. In qRT-PCR, somewhat different results were found compared with previous studies, as oral fluid seemed to be a less suitable sample than serum to detect PRRSV infection early after infection. Interestingly, oral fluid allowed a prolonged detection of the virus compared with serum. Because the results did not correspond with previous studies, it would be interesting to perform similar studies with more virulent type 1 subtype 1 strains or strains belonging to other subtypes and to evaluate the detection of type 1 strains at group level. In addition, the origin of the PRRSV RNA in the oral fluids remains unknown and should be a focus of future studies.
Footnotes
Acknowledgements
The authors would like to acknowledge the veterinarians Willem Van Campe and Laurent Mostin and the animal caretakers of CODA-CERVA Machelen for their assistance in carrying out the biological sample collection. A.B. Cay and N. De Regge share the last authorship.
a.
HerdChek PRRS 3X ELISA, IDEXX Laboratories Inc., Westbrook, ME.
b.
TaqMan NA and EU PRRSV reagents, Life Technologies, Carlsbad, CA.
c.
Barrois-Cebef, Neder-Over-Heembeek, Brussels, Belgium.
d.
BD, Franklin Lakes, NJ.
e.
Applied Biosystems MagMAX Pathogen RNA/DNA kit, Life Technologies, Carlsbad, CA.
f.
LightCycler 480 real-time PCR system, F. Hoffmann-La Roche AG, Basel, Switzerland.
g.
PRRS OF, IDEXX Laboratories Inc., Westbrook, ME.
h.
SPSS Statistics version 22.0, IBM Corp., Armonk, NY.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This study was funded by the Federal Public Service of Health, Food Chain Safety and Environment (RF 10/6235), Belgium.
