Abstract
Bluetongue is a vector-borne viral disease that affects domestic and wild ruminants. The epidemiology of this disease has recently changed, with occurrence in new geographic areas. Various real-time quantitative reverse transcription polymerase chain reaction (real-time qRT-PCR) assays are used to detect Bluetongue virus (BTV); however, the impact of biologic differences between New World camelids and domestic ruminant samples on PCR efficiency, for which the BTV real-time qRT-PCR was initially validated are unknown. New world camelids are known to have important biologic differences in whole blood composition, including hemoglobin concentration, which can alter PCR performance. In the present study, sheep, cattle, and alpaca blood were spiked with BTV serotypes 10, 11, 13, and 17 and analyzed in 10-fold dilutions by real-time qRT-PCR to determine if species affected nucleic acid recovery and assay performance. A separate experiment was performed using spiked alpaca blood subsequently diluted in 10-fold series in sheep blood to assess the influence of alpaca blood on performance efficiency of the BTV real-time qRT-PCR assay. Results showed that BTV-specific nucleic acid detection from alpaca blood was consistently 1–2 logs lower than from sheep and cattle blood, and results were similar for each of the 4 BTV serotypes analyzed.
Keywords
Bluetongue (BT) is a vector-borne disease caused by a virus belonging to the family Reoviridae, genus Orbivirus. 19 The infection occurs in susceptible ruminant hosts in areas where suitable vectors are present. Bluetongue virus (BTV) is currently undergoing a dramatic change in its global epidemiology and distribution, attributed in part to recent climate changes, 14 anthropogenic and social factors, and associated shifts in vector-capacity and distribution of the vector of BTV (midges of the Culicoides genus). 12
Molecular tools for BTV detection, especially real-time quantitative reverse transcription polymerase chain reaction (real-time qRT-PCR) techniques, are recommended, based largely on the high analytical sensitivity of the method. 4,10,13 Because BTV associates primarily with blood cells during infection, the most common specimen for detecting the virus using real-time qRT-PCR is ethylenediamine tetra-acetic acid (EDTA) blood collected from live animals. 11,20 However, biological matrices, and blood in particular, are known to contain well-documented PCR inhibitors, 2,3,26 and biologic differences between species can be expected to variously impact nucleic acid extraction and PCR assay efficiency. Camelid blood composition is substantially different from the blood of other ruminants, especially regarding the concentration of hemoglobin, 2,3,6 and so can be expected to alter PCR performance to some unknown degree. The objective of the present study was to evaluate potential differences in BTV real-time qRT-PCR assay performance efficiency between sheep, cattle, and alpaca blood.
In order to assess the performance efficiency of the BTV real-time qRT-PCR in the different species, EDTA blood was obtained from clinically healthy animals; 1 alpaca and 2 sheep tested in one pool, and 2 cattle were tested in one pool. Bluetongue virus field strain serotypes 10, 11, 13, and 17 originally recovered from sheep residing in California and propagated in Vero cells 7 were stored at −70°C. a One hundred µl of the virus stock was added to 900 µl of alpaca, cattle, or sheep blood to obtain a 1:10 dilution (10−1). To assess the linearity of the real-time qRT-PCR for each species, the spiked 10−1 dilutions were incubated at 4°C overnight on a gentle rocking device, then further diluted with the homologous species blood in a 10-fold dilution series to a final dilution of 10−8, prior to testing. To assess the potential impact of endogenous constituents in alpaca blood on BTV real-time qRT-PCR efficiency, 1 part of spiked alpaca blood in 9 parts of non-spiked sheep blood was prepared, followed by 10-fold dilutions in sheep blood through a 10−8 dilution. The sample RNA was extracted from each of the dilutions using a commercial kit b according to manufacturer’s recommendations, and the real-time qRT-PCR assay for BTV detection was performed as previously described. 17,18 A well-characterized BTV-positive sheep blood sample was used as a positive extraction control, and non-spiked blood from the same species blood pool as used for the dilution series served as negative control. Experiments were performed in triplicate on each of 3 separate days for each of the serotypes 10, 11, 13, and 17 using a single lot of extraction and PCR reagents. Inter-run repeatability, using real-time qRT-PCR threshold cycle (Ct) values as the result, for each species and each serotype was calculated and presented as a mean and standard deviation.
Repeated measures analysis of variance was used to assess between-species BTV real-time qRT-PCR performance differences using Student’s paired t-test (P < 0.05). The serial dilutions that were within the BTV real-time qRT-PCR diagnostic detection range for each species and serotype were analyzed using R2 of the mean of the 3 replicates in order to assess the linearity of the assays.
Alpaca BTV-spiked blood showed significantly higher Ct values, indicating a lesser assay efficiency, when compared to BTV-spiked sheep and cattle blood for all 4 BTV serotypes. The lower detection efficiency of the assay may be due to the alpaca blood interfering with the extraction, PCR amplification, or both steps.
The real-time qRT-PCR results obtained from alpaca blood were on average 4 and 5.4 Ct values higher than those from cattle and sheep blood, respectively. The findings corresponded to approximately 2 logs lower analytic sensitivity (Fig. 1) considering that 3 Ct units are approximately equivalent to a 10-fold dilution 22 and 1 dilution approximates 1 log of virus. The Ct values for BTV-spiked sheep and cattle blood were not significantly different. In contrast, when BTV-spiked alpaca blood was diluted in sheep blood (10−2), the mean Ct values detected were lower compared to the unaltered alpaca blood spiked with a 10-fold greater virus concentration. All subsequent dilutions of the alpaca blood, which contained more than 90% sheep blood, gave similar results to the BTV-spiked sheep blood for all serotypes studied (Fig. 1). The BTV real-time qRT-PCR was linear over 6 log dilutions for all serotypes when tested in cattle and sheep blood. With the exception of BTV-17, the R2 values for assay linearity were >0.95 for dilutions from 10−1 through 10−6, which is within the BTV real-time qRT-PCR Ct diagnostic range. In all experiments, BTV-17 demonstrated a shorter linear range, which was presumed to be the result of BTV-17 cell culture material having a lower initial virus concentration, therefore initially higher Ct values, and an earlier loss of BTV RNA detection. For all serotypes and species, the repeatability of the assays and R2 values obtained for the serial dilutions suggest that differences in the BTV RNA detection limits were a result of the assay performance for different species’ blood rather than an effect of intralaboratory variation.
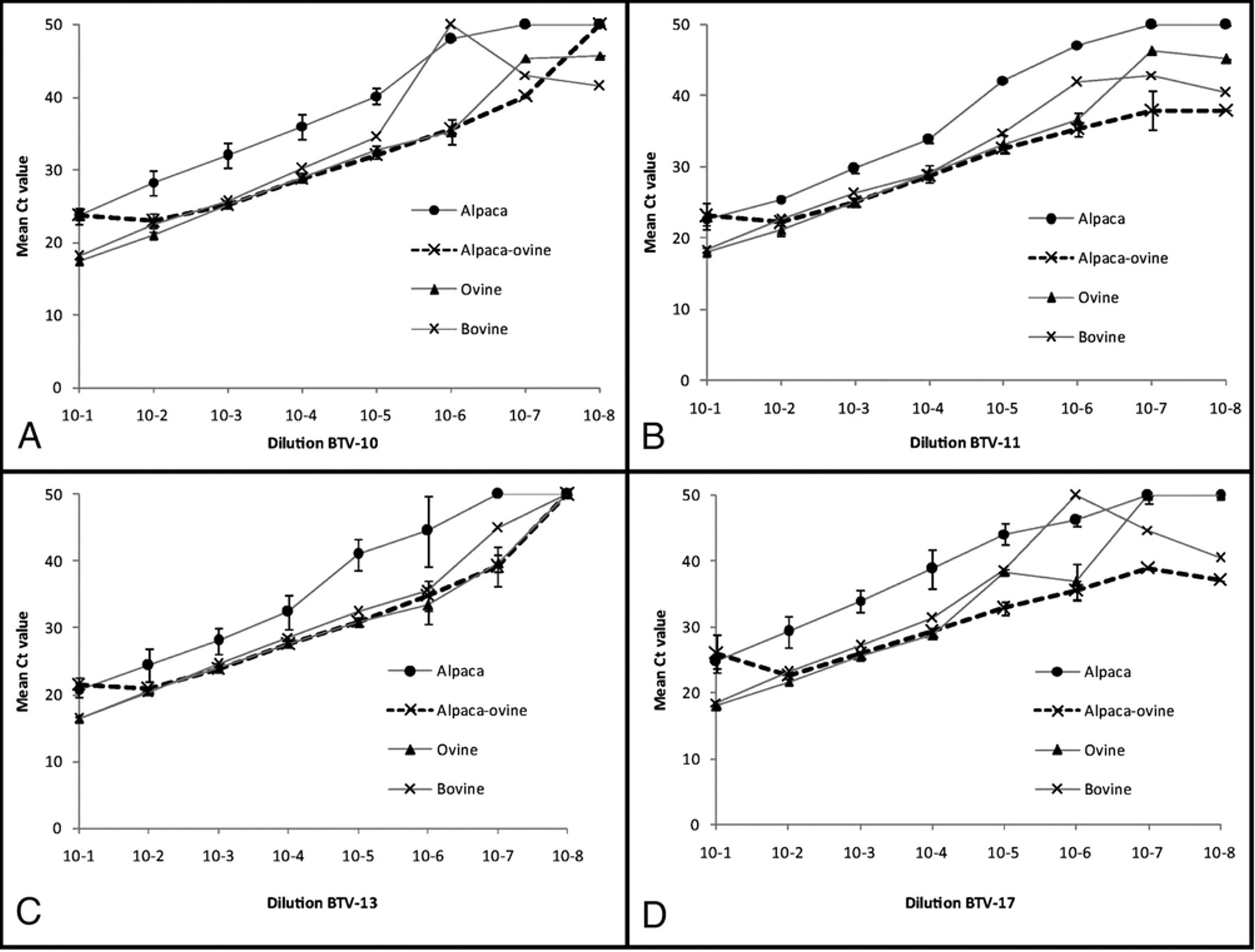
Detection of Bluetongue virus (BTV) by real-time quantitative reverse transcription polymerase chain reaction in different species blood. BTV serotypes 10 (A), 11 (B), 13 (C), and 17 (D) were 10-fold serially diluted in ethylenediamine tetra-acetic acid blood collected from healthy alpaca, ovine, and bovine. 10−1 spiked alpaca blood was also subsequently diluted in ovine blood. Error bars represent the standard deviation of the 3 runs of the assay. Ct = threshold cycle.
The real-time qRT-PCR has been described as the most sensitive and time efficient diagnostic method for identifying BTV infection in ungulates. 24,25,28 In general, although blood characteristics differ from one species to another, most assay validations are performed only on the more common domestic livestock species, which does not include New World camelids. Alpacas are known to be susceptible to BTV infection; however, their role in the epidemiology of this pathogen has yet to be investigated. 11,18 Bluetongue virus serotype 1 has been isolated from healthy and diseased llamas, 15 and isolated reports document detection of BTV by real-time qRT-PCR in alpacas with acute clinical disease in both California and Germany. 8,18 Conflicting findings were reported in Switzerland where a serological survey indicated that exposed animals were serologically negative by competitive enzyme-linked immunosorbent assay, suggesting that New World camelids had not played an important role in BTV-8 spread since 2007. 27 There is however little information regarding the impact of different BTV serotypes, as well as different arthropod vectors, on the manifestation and susceptibility of New World camelids to BT exposure and infection.
Multiple studies have focused on the identification and characterization of PCR inhibitors. Examples include studies showing variable sensitivity of polymerases to hemoglobin, leading to the general conclusion that hemoglobin has an inhibitory effect in PCR assays. 1,3,5 Other studies have demonstrated that purified immunoglobulin (Ig)G from human plasma can cause Taq polymerase inhibition. 2 Immunoglobulin G–related polymerase inhibition has not been studied in animal blood, and no data have been published on the impact of the dimeric IgG found in camelid species on PCR efficiency. 16 Unfortunately, PCR inhibitors have not been critically studied or defined for New World camelids, or in reality for most animal species despite the broad use of PCR in veterinary diagnostics. Further investigations are needed to identify the specific blood and tissue components affecting PCR assay performance in alpaca and other ruminants susceptible to BTV, so that appropriate modifications in the methodology can be used to optimize viral detection. 21,23,26
A limitation of the present study is that the blood values (complete blood cell count and chemistry panel) of the convenience blood samples taken from clinically healthy animals were not measured prior to the analyses, so any comparisons made assumed normal reference values as published for healthy animals. It is also unknown whether the results of the study performed for BTV, can be implied for other agents and real-time qRT-PCR assays. However, a similar set of experiments was performed comparing real-time qRT-PCR performance for Bovine viral diarrhea virus (BVDV) in blood of sheep, cattle, and alpacas utilizing the same enzymes and an assay internal control that is routinely used with the BVDV real-time qRT-PCR assay. In the preliminary BVDV real-time qRT-PCR trial, no significant differences were found in the performance of the assay when using blood from different species (data not shown). This finding highlights the importance of validating each assay for every species, and that no assumptions can be made about equivalent efficiency and performance of real-time qRT-PCR assays in different sample matrices. 9
In conclusion, it is recommended that assay validation should be performed for each species and each sample matrix before the assay is routinely used for diagnostic purposes. The present study demonstrated that blood from alpaca results in a different BTV real-time qRT-PCR assay performance, thus interfering with detection efficiency, and potentially resulting in an increased risk of false-negative test results. The lower real-time qRT-PCR detection sensitivity for BTV in alpaca blood may result in undiagnosed BTV infection in alpacas, and may also underestimate prevalence data if used without full knowledge of the actual BTV real-time qRT-PCR performance in this species.
Footnotes
Acknowledgements
The authors thank Dr. Jim MacLachlan for his advice and critical reading of the manuscript.
a.
California Animal Health and Food Safety Laboratory System, Davis, CA.
b.
MagMAX™-96 for Microarrays Total RNA Isolation Kit, Ambion, Austin, TX.
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Post-graduate fellowship support was provided to Barbara Brito by Becas Chile, Ministry of Education, Chilean Government.
