Abstract
Formalin-fixed, paraffin-embedded tissues from elk (Cervus elaphus), goats, and camelids with case histories and lesions suggestive of Parelaphostrongylus tenuis were examined by histology to characterize lesions that could aid in definitively diagnosing P. tenuis infection. Additionally, sections of paraffin-embedded tissue were used in a nested polymerase chain reaction (nPCR) using Parelaphostrongylus-specific primers to determine how PCR results corresponded with histological findings. Histological changes in brain and spinal cord consisted of linear tracks of hemorrhage; tracks or perivascular accumulations of hemosiderin-laden macrophages; acute foci of axonal degeneration and/or linear glial scars; and perivascular, parenchymal, or meningeal accumulations of eosinophils and/or lymphocytes and plasma cells. Of the 43 samples with histologic lesions consistent with neural larval migrans, 19 were PCR positive; however, only 8 were confirmed Parelaphostrongylus by DNA sequencing. Additionally, 1 goat was identified with a protostrongylid that had a 97% identity to both Parelaphostrongylus odocoilei and a protostrongylid nematode from pampas deer (Ozotoceros bezoarticus celer) from Argentina. None of the histologic lesions individually or in combination correlated statistically to positive molecular tests for the nematode. The results indicate that it is possible to extract Parelaphostrongylus DNA from formalin-fixed, paraffin-embedded tissue, but extended fixation presumably can cause DNA crosslinking. Nested PCR provides another diagnostic tool to identify the cause of neurologic disease in camelids and elk with histologic lesions consistent with neural larval migrans. Furthermore, potential novel protostrongylid DNA was detected from a goat with lesions consistent with P. tenuis infection, suggesting that other neurotropic Parelaphostrongylus species may occur locally.
Introduction
Parelaphostrongylus tenuis is a nematode commonly found in the meninges of white-tailed deer (Odocoileus virginianus), lending it the common name meningeal worm. While P. tenuis rarely causes clinical signs or lesions in the definitive host, it creates severe neurological problems in aberrant or dead-end hosts such as elk (or red deer; Cervus elaphus), moose (or Eurasian elk; Alces alces), llamas (Lama glama), alpacas (Vicugna pacos), and numerous other cloven hoof stock, including goats (Capra aegagrus).1,9,19 Clinical signs include ataxia, circling, head tilt, hind limb paresis, arched neck, blindness, and eventual death.9,12,17,19 Dead-end hosts become affected by ingesting terrestrial snails and slugs infected with third-stage larvae. The larvae penetrate the gastrointestinal wall and travel up lateral spinal nerves to the spinal cord and/or brain, causing damage as they migrate. 7
Currently, there is no commercially available antibody test to detect P. tenuis antemortem. Most diagnoses are based on clinical signs, antemortem detection of high levels of eosinophils in cerebrospinal fluid, and response to treatment with anthelmintics and nonsteroidal anti-inflammatory drugs.14,15 The likelihood of finding a nematode during gross necropsy and/or histology is remote, and therefore diagnoses are often inconclusive. Histologic lesions of the central nervous system (CNS), that characterize P. tenuis infection are similar to Baylisascaris procyonis in human, nonhuman primate, and avian hosts and include axonal degeneration and swelling, perivascular cuffing, the presence of hemosiderin-laden macrophages, and increased numbers of eosinophils.7,9,12,19 However, these lesions may also indicate other neurologic conditions, such as encephalitis due to a variety of viral or parasitic causes. A conclusive test for P. tenuis is critical for proper diagnosis.
In 2011, 13 a nested polymerase chain reaction (nPCR) method was developed to definitively identify Parelaphostrongylus.3–5,10,13,18 Using nPCR allows for increased sensitivity to detect Parelaphostrongylus spp. DNA at quantities that conventional PCR would not detect. Having a positive identification could provide owners and/or wildlife managers a definitive cause of death, allowing them to implement targeted, specific mitigation protocols. To determine if a definitive diagnostic method for P. tenuis could be developed, formalin-fixed, paraffin-embedded tissues from ruminants with case histories and lesions suggestive of P. tenuis were examined by histology to characterize lesions that could aid in definitively diagnosing P. tenuis infection. Additionally, nPCR using Parelaphostrongylus-specific primers was utilized to determine how PCR results corresponded with histological findings.
Materials and methods
A retrospective search of the University of Tennessee, College of Veterinary Medicine (Knoxville, Tennessee) pathology archive spanning the past 10 years (2003–2013) was conducted to find all necropsy cases of suspected P. tenuis infection or unexplained neurological lesions in elk, llamas, alpacas, and goats. Archived histology slides of all selected cases were reviewed by a board-certified pathologist (SJ Newman) to determine if characteristic lesions, including axonal degeneration, migration tracts, encephalitis, meningitis, hemorrhage, hemosiderin-laden macrophages were present; lesions were recorded for each case. Tissue samples were cut from paraffin blocks from all cases with characteristic P. tenuis lesions for DNA extraction and subsequent PCR testing as previously described.4,13 An additional histology slide was cut after the 10-µm tissue PCR slices were taken and subsequently stained to determine if nematodes had been uncovered at the depth of the corresponding PCR sample. Extraction of DNA was completed according to the manufacturer’s instructions, a and PCR was completed as previously described.4,13 Separate microtome blades were used for each block, and the blades were cleaned thoroughly between blocks and microtomes using DNA degrading solution. Both DNA extraction and PCR-negative controls were used in nPCR reactions. Nested PCR reactions were attempted in duplicate. The PCR products were examined using electrophoresis in a 2% agarose gel. Positive DNA amplicons were removed and purified using gel extraction following the manufacturer’s instructions. a Bidirectional sequencing of amplicons was performed using nested amplification primers at the University of Tennessee sequencing facility (Knoxville, Tennessee). The obtained sequences were compared to those in GenBank using Basic Local Alignment Search Tool (BLAST). In samples where a nematode was present but Parelaphostrongylus was not detected using the aforementioned primers, nematode-wide, short subunit ribosomal RNA primers were used to determine if other nematode species were responsible for the verminous migration. A chi-square analysis of individual variables and logistic regression analysis of multiple variables were used to compare histologic lesion types that were PCR positive or had evidence of a nematode on histological sections. Statistical significance was determined at p = 0.005.
Results
A total of 43 cases (individual animals) were identified as possible Parelaphostrongylus infections based on the necropsy reports. Of these, 38 cases had characteristic histologic lesions but only 22 samples (individual tissue sections) that were either PCR positive and/or had nematodes present in histologic section were included in further study (Table 1). In 9 elk, 2 samples were from spinal cord and the other 8 were from brain (10). In alpacas, 3 samples were from spinal cord and 3 were from the brain (6). In llamas, 4 samples were from brain and 1 sample was from spinal cord (5). The single goat had spinal cord involvement only (1).
Polymerase chain reaction (PCR) results, sequence results, worm presence, and characteristic histologic lesions of neural larval migrans.*
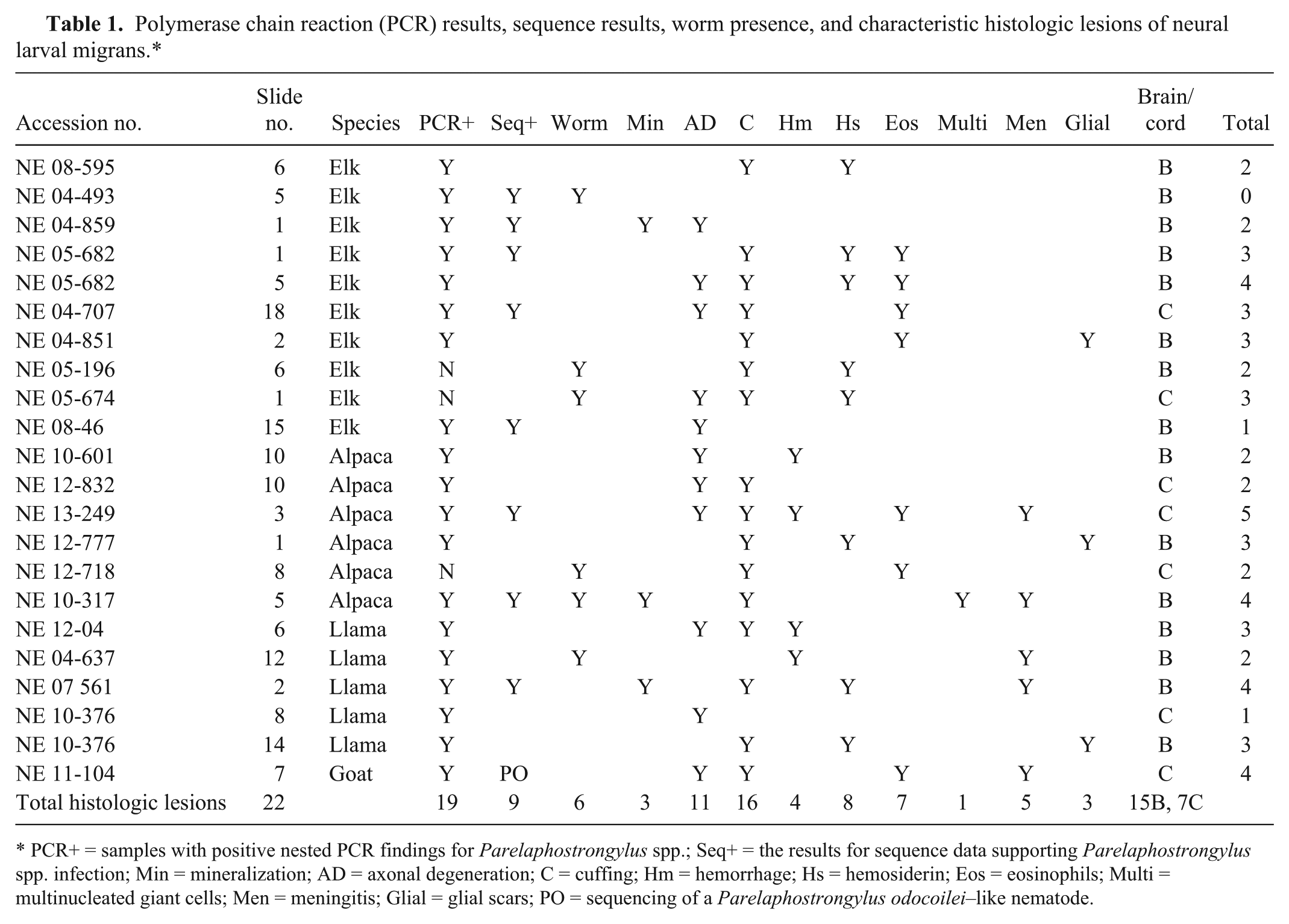
PCR+ = samples with positive nested PCR findings for Parelaphostrongylus spp.; Seq+ = the results for sequence data supporting Parelaphostrongylus spp. infection; Min = mineralization; AD = axonal degeneration; C = cuffing; Hm = hemorrhage; Hs = hemosiderin; Eos = eosinophils; Multi = multinucleated giant cells; Men = meningitis; Glial = glial scars; PO = sequencing of a Parelaphostrongylus odocoilei–like nematode.
Nineteen out of 22 (86%) samples were PCR positive (see Table 1). The samples were from 17 animals, including 7 elk, 4 llamas, 5 alpacas, and 1 goat. Of the 19 samples, 8 were sequenced as P. tenuis: 5 from elk, 1 from a llama, and 2 from alpacas. Six of the 22 (27%) samples had a nematode present within the histologic tissue section considered to be consistent with P. tenuis; however, only 2 of the 6 were sequenced as P. tenuis. The remaining amplicons failed to produce usable sequences on multiple attempts, and thus, confirmation of the presence of Parelaphostrongylus specifically was unobtainable. GenBank BLAST analysis of the 83 nucleotide bp consensus partial second ribosomal internal transcribed spacer sequence of the goat disclosed a 97% identity and 100% coverage to multiple P. odocoilei sequences, which is a nematode found in mule deer (Odocoileus hemionus) and black-tailed deer (Odocoileus hemionus columbianus) of the Pacific Northwest, 9 and 97% identity and 95% coverage to an unidentified member of the Protostrongylidae family from an endangered pampas deer (Ozotoceros bezoarticus celer) from Argentina (GenBank accession no. JN092130). In contrast, the goat sequence had 100% coverage and a 94% identity to 2 P. tenuis sequences. The nematode-wide small subunit ribosomal RNA primers did not produce any PCR-positive amplicons.
In summary, nematodes (6/22, 27%), mineralization (3/22, 13%), axonal degeneration (11/22, 50%), perivascular cuffs (16/22, 72%), hemorrhage (4/22, 18%), hemosiderin-laden macrophages (8/22, 36%), eosinophils (7/22, 31%), multinucleated giant cells (1/22, 4%), meningitis (5/22, 22%), and glial scars (3/22, 13%) were identified in samples. In elk, the number of histologic changes ranged from 0–4, in alpacas 2–5, in llamas 1–4, and there were 4 in the single goat.
A summary of histologic lesions for animals along with P. tenuis PCR and sequencing results is presented in Table 1. Characteristic lesions are shown in Figures 1 and 2. All 9 elk had lesions in 1 or both sections that would be considered consistent with P. tenuis infection and 8 were PCR positive. Three elk had evidence of verminous encephalitis; however, only 1 of the 3 animals was PCR and sequence positive. All 6 alpacas had lesions in 1 or both sections that would be considered consistent with P. tenuis infection and 5 were PCR positive; 1 alpaca had evidence of a nematode, but was PCR negative. All 5 llamas had lesions in 1 or both sections that would be considered consistent with P tenuis infection and all 5 were PCR positive; in 1 alpaca, there was also evidence of a nematode. The single goat had lesions consistent with P. tenuis and was PCR positive, but further sequencing revealed a potentially novel Parelaphostrongylus spp. closely related to P. odocoilei.
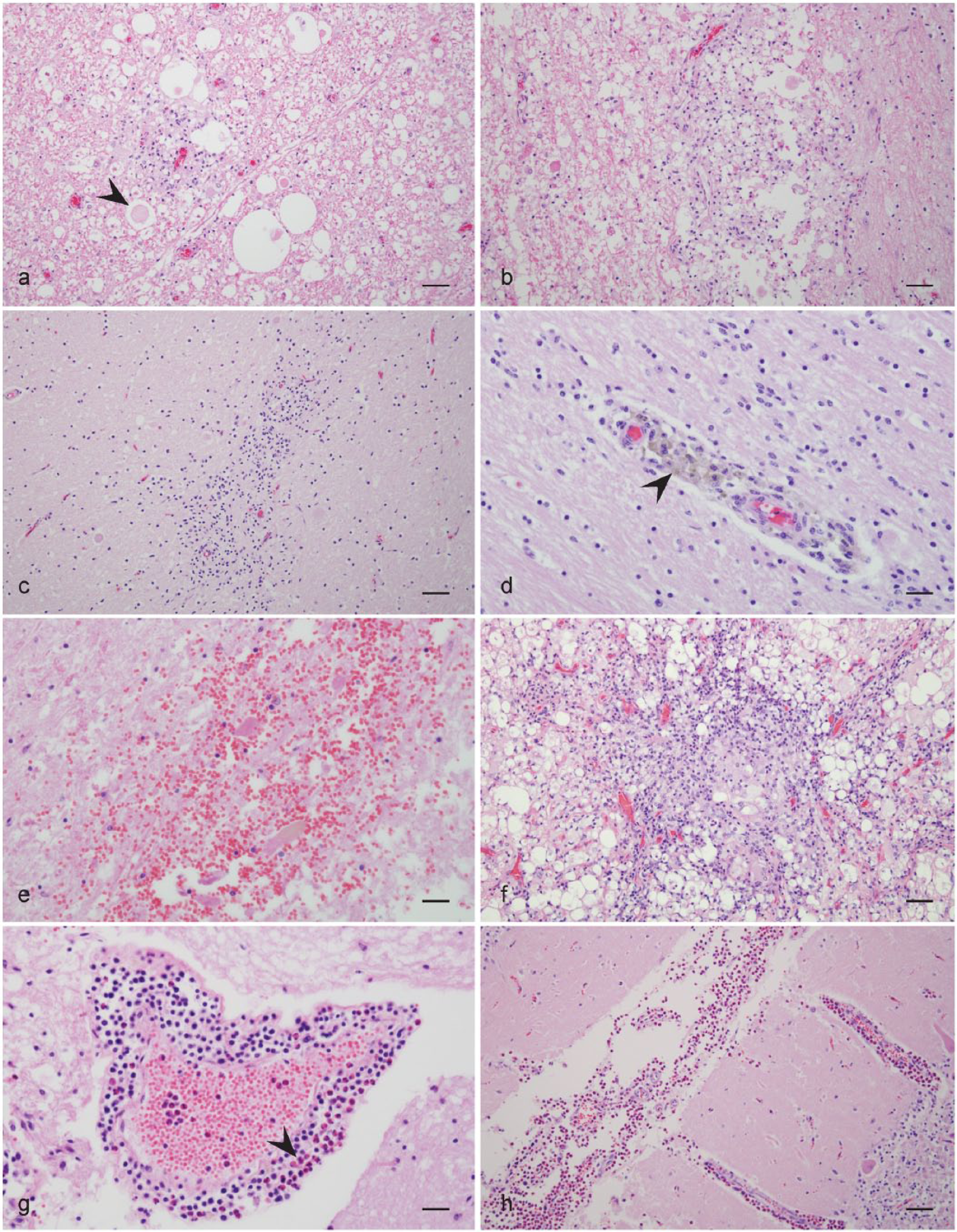
Characteristic histologic lesions of suspected neural larval migrans.
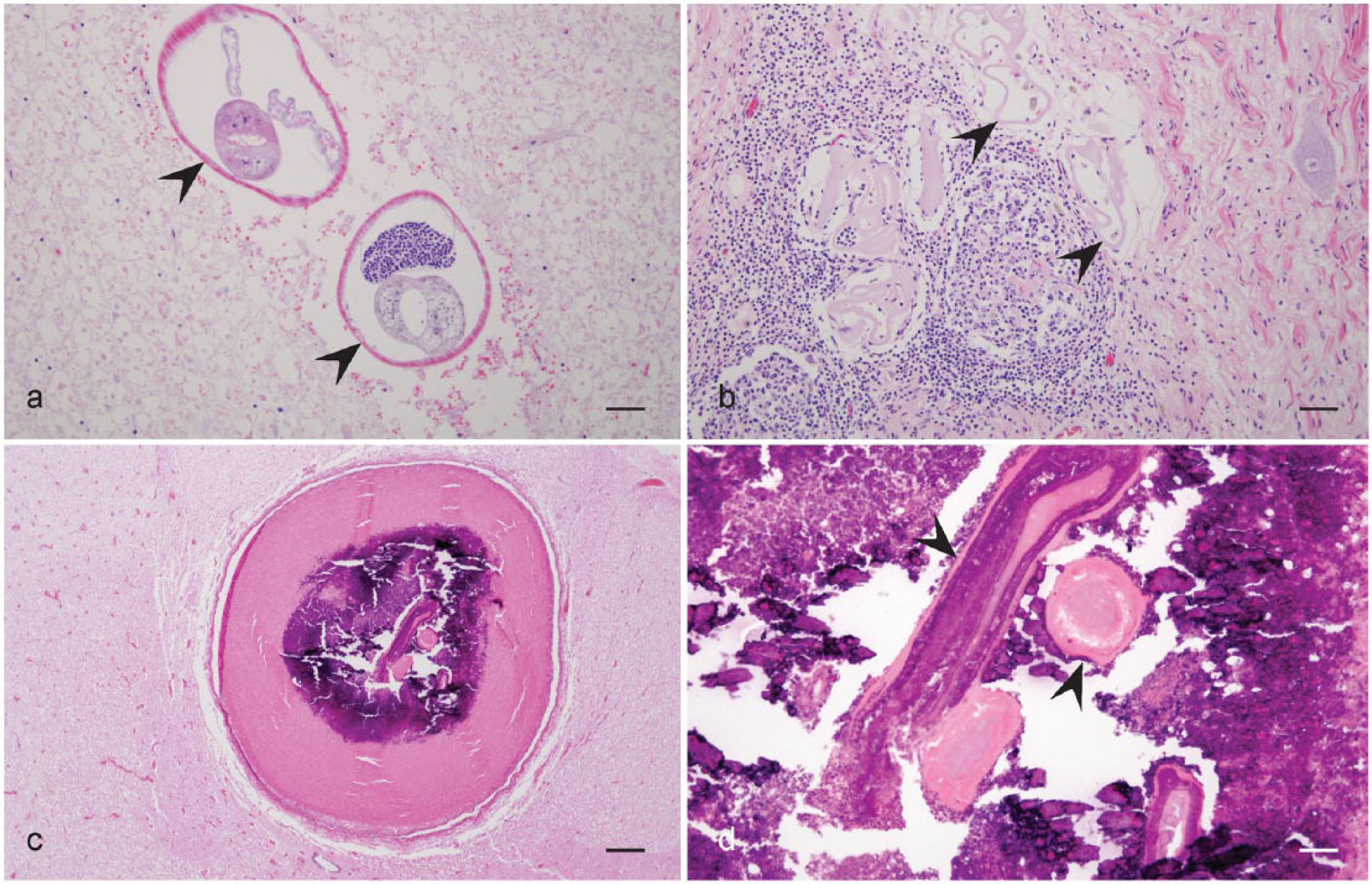
Characteristic histologic lesions of neural larval migrans.
Statistical analysis performed on the 22 samples did not show a strong relationship between individual histologic lesions and the ability to detect P. tenuis by PCR and sequencing. In the chi-square analysis of individual histologic variables, the strongest associations were between the presence of the nematode (P = 1.106) and the presence of eosinophils (P = 0.099), but neither were significant. In order to reduce the chance of type I error (false positive) and applying a Bonferroni adjustment to an alpha of 0.05, produces a new significance level of 0.005. The logistic regression analysis of multiple variables determined that no combination of histologic variables could positively predict the presence of P. tenuis.
Discussion
The nematodes from the 4 PCR-negative cases had histologic lesions consistent with neural larval migrans. Parasites identified in affected sections had variable characteristics, but degeneration and mineralization often hampered examination of internal features of the larvae. When recognizable, histological characteristics, including the musculature, hypodermal chords, and large multinucleate intestinal cells, were consistent with the superfamily Metastrongyloidea. 1 Both the diameter and the anatomical location of this nematode were consistent with P. tenuis and Elaphostrongylus rangiferi. 9 Only P. tenuis has been previously found in the areas where these animals originated. Although E. rangiferi has not been specifically tested with the nPCR primer pair used in the current study, the primary reaction primers were found not to amplify E. rangiferi. 3 A possible explanation for the negative PCR results in the samples with visible nematode cross sections is that extended formalin fixation can crosslink the DNA, which inhibits amplification, an outcome seen in other studies.2,11 Undoubtedly, in 1 sample (Fig. 2c, 2d), the nematode was probably too mineralized and degenerative to allow DNA extraction.
Six of the 8 sequence-positive samples did not have a nematode present in the histologic tissue examined initially or following additional tissue sectioning. These results indicate that P. tenuis DNA can be detected with nPCR from formalin-fixed, paraffin-embedded tissue even if a nematode is not visible in the sample. The DNA of the nematode appears to remain in the tissue of the host in some instances and, when in large enough amounts, can be detected with nPCR. While the potential for crosslinking inhibits identifying some samples, this technique provides an additional means to diagnose P. tenuis infection in suspicious cases.
Interestingly, evidence was found of a goat infected with a protostrongylid that had a 97% identity to both P. odocoilei and an unpublished sequence of an unidentified member of the Protostrongylidae family from an endangered pampas deer. Although the spinal cord of the goat had lesions consistent with P. tenuis infection including perivascular cuffing, eosinophilic encephalitis, and axonal degeneration, the sequence had only a 95% identity to multiple P. tenuis sequences in GenBank. Parelaphostrongylus odocoilei is found in the muscle of infected hosts, and it is not considered to have a CNS migration phase 9 ; however, it is possible that aberrant migration can occur. Unfortunately, sections of worms, larvae, or eggs were not observed histologically in this case, and hence it is unknown if this is a previously unrecognized protostrongylid species or if it is a unique sequence group for P. odocoilei. Previously, P. odocoilei has been found in wild cervids and caprines from northwestern North America.6,8,9,14,16
In the current study, the number of different histologic lesions per case varied from 1 to 5; however, there was no statistically significant relationship with samples positively identified as P. tenuis based on the number and/or type of lesions. Perivascular cuffing and axonal degeneration were the most common lesions in samples positively identified as P. tenuis; however, they were also the most common lesions in PCR- or nematode-negative samples. These lesions are not considered definitive for neural larval migrans but, when identified, consideration of P. tenuis is warranted in appropriately infected species. Hemosiderin-laden macrophages around vessels or in tracks, are supportive of previous hemorrhage, and may be found in some cases of neural larval migrans. Eosinophils within the CNS have always been linked to migrating parasites or in pigs to salt toxicity. Eosinophils were identified in many cases but were limited to the elk, alpaca, and goat cases. Antemortem cerebrospinal fluid taps consider eosinophils as a strong indicator of possible neural larval migrans, in particular P. tenuis. 13 Meningitis overall was relatively uncommon, and not seen in any elk. The cause of this is not known, as there is widespread evidence of inflammatory infiltrates, but infiltrates are not localized to the meninges. Hemorrhage, especially in the form of an acute linear track lesion, is only expected in acute cases and was not common in the current series of lesions. Glial scars represent old healed regions of migration and were relatively rare in the cases described herein. Mineralized foci and multinucleated giant cells were the least common change, and were only found in sections with a worm or a degenerative worm fragment present. Additionally, multiple types of lesions were identified in most cases ranging from a total of 0–4 lesions in elk, 2–5 lesions in alpacas, 1–4 lesions in llamas, and 4 lesions in the goat. The elk case that had no histologic reaction despite the presence of parasites in cross-sections was anomalous (Fig. 2a). Building support for visceral larval migrans shows that examination of multiple representative sections may reveal a complex of histologic lesions that might lend support for stronger consideration of P. tenuis migration, and hence if nematodes were not available in section, diagnostic nPCR could be further recommended based on the results in the present study. More brain samples were available for characterization, likely due to the easier collection of brain rather than spinal cord in these animals. Cord was collected in those cases in which it was deemed warranted from the history and physical exam. Fewer elk had cord available because of their large size and often longer decomposition period at necropsy. There is no reason to suspect that the location within the CNS is important unless clinical signs are able to localize lesions to specific anatomical locations. Unfortunately, the sample size was small for each species, and further examination of representative histologic lesions in species of interest is needed. The nPCR method provides an additional means to diagnose P. tenuis infection in suspicious cases. Furthermore, in cases of suspected P. tenuis infection, it is important to limit formalin fixation time to minimize excessive cross linking of the nematode DNA. This is particularly important with P. tenuis due to the frequent inability to detect the nematode on histological exam of suspect cases; however, being able to amplify Parelaphostrongylus DNA by nPCR is demonstrated in this study.
Footnotes
Acknowledgements
Dr. Ann Reed is acknowledged for statistical support. Ms. Anik Vasington provided graphic assistance. Ms. Misty Bailey provided technical editing support.
a.
Qiagen GmbH, Hilden, Germany.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
Support for this project was provided by the Fund for Education Advancement and Research (RW Gerhold and SJ Newman) from the Department of Biomedical and Diagnostic Sciences, and the Center of Excellence in Livestock Diseases and Human Health (C Dobey), College of Veterinary Medicine, University of Tennessee, Knoxville, TN.
