Abstract
Avian bornaviruses (ABVs) are a group of genetically diverse viruses within the Bornaviridae family that can infect numerous avian species and represent the causative agents of proventricular dilatation disease, an often fatal disease that is widely distributed in captive populations of parrots and related species. The current study was designed to assess the antigenic variability of the family Bornaviridae and to determine its impact on ABV diagnosis by employing fluorescent antibody assays. It was shown that polyclonal rabbit sera directed against recombinant bornavirus nucleoprotein, X protein, phosphoprotein, and matrix protein provided sufficient cross-reactivity for the detection of viral antigen from a broad range of bornavirus genotypes grown in cell culture. In contrast, a rabbit anti-glycoprotein serum and 2 monoclonal antibodies directed against nucleoprotein and phosphoprotein proteins reacted more specifically. Antibodies were readily detected in sera from avian patients infected with known ABV genotypes if cells persistently infected with a variety of different bornavirus genotypes were used for analysis. For all sera, calculated antibody titers were highest when the homologous or a closely related target virus was used for the assay. Cross-reactivity with more distantly related genotypes of other phylogenetic groups was usually reduced, resulting in titer reduction of up to 3 log units. The presented results contribute to a better understanding of the antigenic diversity of family Bornaviridae and further emphasize the importance of choosing appropriate diagnostic tools for sensitive detection of ABV infections.
Introduction
Avian bornaviruses (ABV) are negative-sense, single-stranded RNA viruses belonging to the family Bornaviridae. In 2008, these viruses were identified in parrots and related species (order Psittaciformes) suffering from proventricular dilatation disease (PDD)13,14 and later experimentally confirmed to represent the causative agents of the disease.6,16,19,20,28 Proventricular dilatation disease causes gastrointestinal tract dysfunction with weight loss, regurgitation, and shedding of undigested seeds with the feces as the most common presenting signs. Affected birds have also been described to suffer from neurological symptoms like ataxia or seizures. In general, the disease is associated with lymphoplasmacytic infiltrates in central and peripheral neuronal tissues.7,20,23
Until 2008, 2 genotypes of the mammalian Borna disease virus (BDV) were the only known members of the family Bornaviridae. 17 Since then, however, the family has grown considerably as has the number of documented host species (Fig. 1). To date, 7 ABV genotypes (ABV-1 to -7) have been described in psittacine birds.14,29,34 These genotypes can be located to 2 different phylogenetic groups (Fig. 1). Additional ABV genotypes were discovered in 2 other host orders, passerines (Passeriformes) and waterfowl (Anseriformes). Three distinct ABV genotypes (ABV-C1 to -C3) were described in common canaries (Serinus canaria forma domestica).30,35 These 3 genotypes are closely related to a genotype identified in Bengalese finches (or white-rumped munia, Lonchura striata forma domestica) in Japan. 30 Further ABV isolates obtained from estrildid finches form a new genotype (ABV-EF), which is only distantly related to the genotypes described previously. 31 Yet another genotype (ABV-CG) was isolated from wild waterfowl of different species, such as Canada geese (Branta canadensis) and mute swans (Cygnus olor) in North America.8,18 The 2012 detection of reptile bornavirus sequences in tissues from a Gabon viper (Bitis gabonica) suggests that reptiles may also be natural hosts of bornaviruses. 3
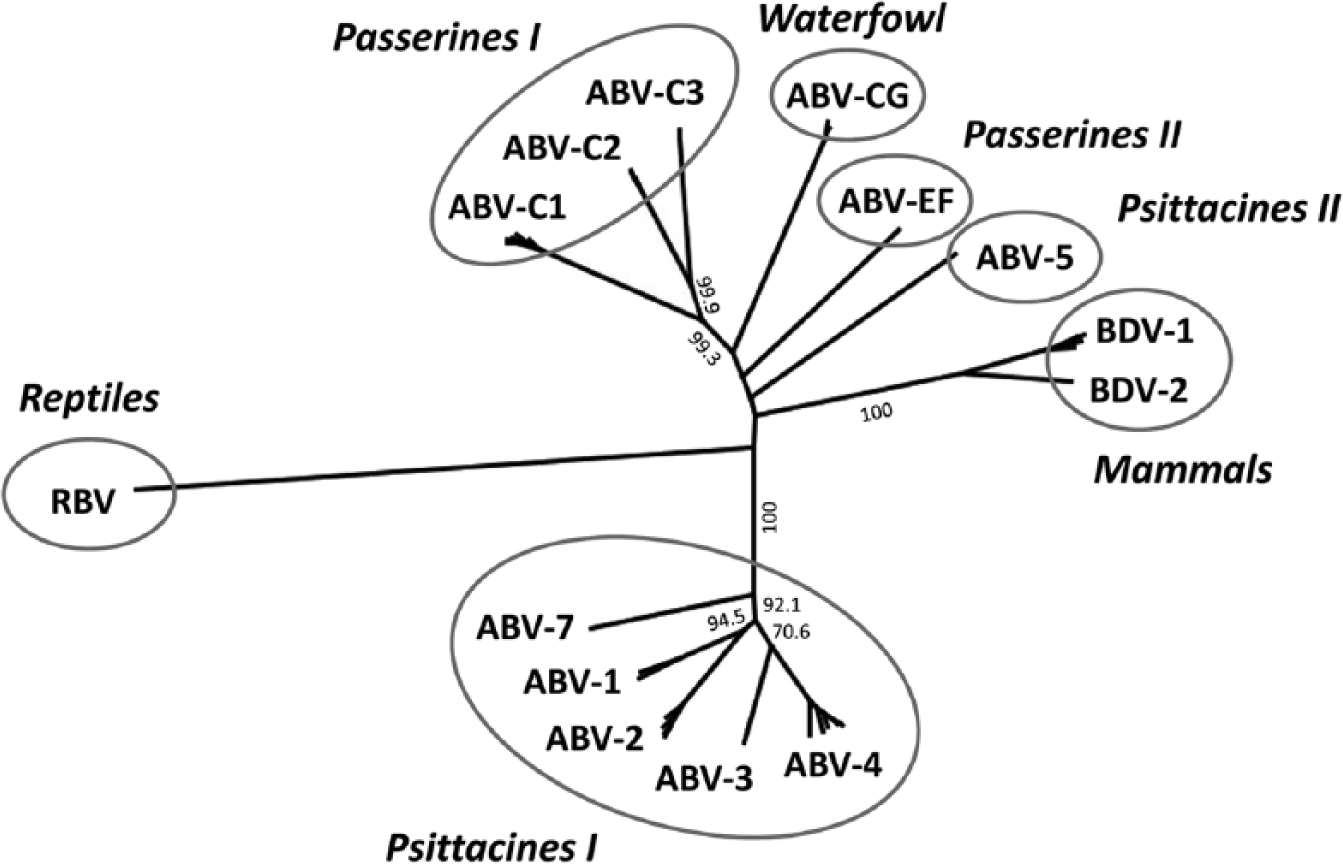
The Bornaviridae family displays a broad genetic variability. To date, 2 mammalian Borna disease virus (BDV) genotypes, 13 Avian bornavirus (ABV) genotypes of 5 phylogenetic groups, and 1 reptile bornavirus (RBV) have been identified. Partial nucleoprotein (N), X, and phosphoprotein (P) gene sequences (max. 960 base pairs; corresponding to positions 896–1,825 of the full genome of strain ABV-2 #6609, GenBank accession no. FJ620690) were obtained from GenBank and inferred using the neighbor-joining algorithm and a Jukes–Cantor distance model in Geneious Pro 6.1.6. l The tree was rooted to RBV Bitis gabonica. Values at branches represent support in 1,000 bootstrap replicates, and only values above 70 at major branches are shown. Sequences of ABV-6 (group Psittacines II) and ABV Lonchura striata forma domestica (group Passerines I) are not available for this genomic region.
Avian bornavirus strains of the same genotype usually exhibit at least 95% nucleotide identity with each other. For genotypes belonging to the same phylogenetic group, sequence identity is approximately 80–85%, whereas viruses from different phylogenetic groups are only 65–75% identical. Reptile bornavirus represents, to date, the most divergent member of family Bornaviridae, with a maximum of 57% nucleotide sequence identity to all other known genotypes. It has been demonstrated that these differences may result in reduced sensitivity of reverse transcription polymerase chain reaction (RT-PCR) assays commonly used for ABV detection.30,31
Detection of bornavirus antigen and bornavirus-specific antibodies is routinely performed by serological assays, such as indirect fluorescent antibody test (IFAT), enzyme-linked immunosorbent assay (ELISA), or Western blot. It was shown that antigens from different bornavirus genotypes could be detected using polyclonal anti–ABV-2 or anti–BDV-1 rabbit sera.20,27,29–31,34,35 Bornavirus-specific antibodies in the sera of various psittacine birds could be detected by IFATs using BDV-infected Madin–Darby canine kidney (MDCK) cells or ABV-infected quail fibroblast (CEC-32) cells. 12 However, the extent to which antigenic heterogeneity of different bornavirus genotypes affects the sensitivity of these diagnostic tools has not yet been evaluated systematically.
The current study was designed to assess bornavirus antigen detection using polyclonal rabbit sera and mouse monoclonal antibodies (mAbs). Furthermore, the effect of antigenic variability on the sensitivity of antibody detection by IFAT was analyzed by testing sera obtained from naturally or experimentally bornavirus-infected animals against various cell lines infected with different ABV and BDV genotypes.
Material and methods
Cell lines and viruses
The CEC-32, quail smooth muscle (QM7), chicken hepatoma (LMH), African green monkey kidney (Vero), and MDCK cell lines were used for IFATs. Cells were persistently infected with bornavirus isolates ABV-1 #16364, 29 ABV-2 #6609, 27 ABV-4 #6758, 27 ABV-7 #16667a, 29 ABV-C1 AS-20, 30 ABV-C2 #15864, 30 ABV-EF VS-4709, 31 BDV-1 H215, 26 BDV-1 He/80/FR, 21 or BDV-2 No/98. 17
Anti-bornavirus antibodies and serum samples from infected animals
The following polyclonal rabbit sera directed against individual bornavirus proteins or monoclonal mouse antibodies were used for the detection of bornavirus antigens in cell cultures: rabbit anti–ABV2-N (dilution 1:500), 24 rabbit anti–BDV1-N (1:1,000), rabbit anti–BDV1-X a (1:700), rabbit anti–BDV1-P a (1:500), rabbit anti–BDV1-M b (1:1,000), 2 rabbit anti–BDV1-G c (1:200), mouse anti–BDV1-N Bo18 d (1:100), 26 and mouse anti–BDV1-P 30H8 e (1:500). Antibody concentrations were chosen to give optimal signal-to-background ratios for the respective homologous target virus.
In addition, serum samples obtained from bornavirus-infected psittacines, canaries, and mice were tested for the presence of bornavirus-specific antibodies. Psittacine sera originated from birds of different species that were naturally infected with genotypes ABV-1 (n = 2), ABV-2 (n = 3), or ABV-4 (n = 2), or from cockatiels experimentally infected with ABV-4 (n = 2). 28 Sera of common canaries were collected after experimental infection with either genotype ABV-C1 (n = 4) 28 or ABV-C2 (n = 4). 30 Borna disease virus genotype 1–specific sera were obtained from experimentally infected mice (n = 4).
Indirect fluorescent antibody test
Indirect FATs were performed to test bornavirus antigen detection by different polyclonal rabbit sera and monoclonal mouse antibodies as well as to detect bornavirus-specific antibodies in sera from infected animals. For this purpose, persistently infected cells were mixed at a ratio of 1:20 with uninfected cells of the same cell line and seeded in 96-well microtiter plates. f The mixture of infected and uninfected cells allowed for a better discrimination between positive signals and background fluorescence. Wells with uninfected cells of each cell type served as additional controls. Following overnight culture, cells were fixated with 3% paraformaldehyde g and permeabilized with 50 µl of 0.5% Triton X-100 h in phosphate buffered saline (PBS). For antigen detection, permeabilized cells were incubated with 50 µl per well of diluted polyclonal rabbit sera or monoclonal mouse antibodies for 90 min at room temperature (RT), followed by incubation with 50 µl of either cyanine (Cy)3-labeled goat anti–rabbit-immunoglobulin (Ig)G i (1:300) or goat anti–mouse-IgG i (1:200).
For detection of bornavirus-specific antibodies from avian and mouse sera, permeabilized cells were incubated with 50 µl of three-fold serial dilutions of psittacine, canary, or mouse sera for 90 min at RT. Subsequently, cells were incubated for 90 min with 50 µl of rabbit anti–gray parrot-IgG (diluted 1:300) 28 or rabbit anti–canary-IgG (1:5,000) 30 for the detection of psittacine or canary sera, respectively. Thereafter cells were incubated with 50 µl of goat anti–rabbit-IgG-Cy3 (1:300) overnight at RT in the dark. Cells incubated with mouse sera were similarly treated with goat anti–mouse-IgG-Cy3 (1:200). All sera were diluted in PBS supplemented with 2% normal goat serum, j and cells were thoroughly washed 3 times with PBS after each incubation step. Cells were analyzed using fluorescence microscopy, and, for each sample and dilution step, bornavirus-positive and bornavirus-negative wells were compared. Wells were considered positive if the expected 5% bornavirus-positive cells were markedly brighter than the background staining of uninfected cells in the same well and in the corresponding bornavirus-negative control well. Endpoint titers were calculated for each serum with each target cell line. For each individual serum, titers obtained with the homologous target virus were set to 100%, and cross-reactivity to heterologous genotypes is presented as a percentage of this titer.
Isolate BDV-1 He/80/FR was used for IFAT with monoclonal mouse antibodies and monospecific rabbit sera, while isolate BDV-1 H215 was used for antibody detection from patient sera. Further information about the viruses and cell lines used for the respective analysis is provided in the results section and the respective figures.
Sequence analysis
RNA extracted from cell cultures persistently infected with BDV-1 H215 was amplified by RT-PCR, and the partial genome sequence was determined by Sanger sequencing. k The sequence was deposited at GenBank with accession number KJ950616. Amino acid sequences of complete bornavirus nucleoprotein (N) or matrix (M) protein genes were derived from GenBank. Sequence alignments were performed by MUSCLE using commercial software. l
Results
Avian bornaviruses can be detected using cross-reactive antibodies
Detection of ABV antigens in infected cell cultures, tissue sections, or by Western blot is performed with antibodies that are directed against proteins of BDV-1 or psittacine ABV genotypes. These antibodies have proven to provide sufficient cross-reactivity with other bornavirus genotypes.20,24,27,29–31,34,35 In the current study, a panel of different polyclonal rabbit sera and monoclonal mouse antibodies for antigen was systematically tested for the detection of 4 bornavirus genotypes (ABV-2, ABV-C2, ABV-EF, and BDV-1) from different phylogenetic groups (Fig. 2). Some antibodies were additionally tested with further selected bornavirus genotypes.
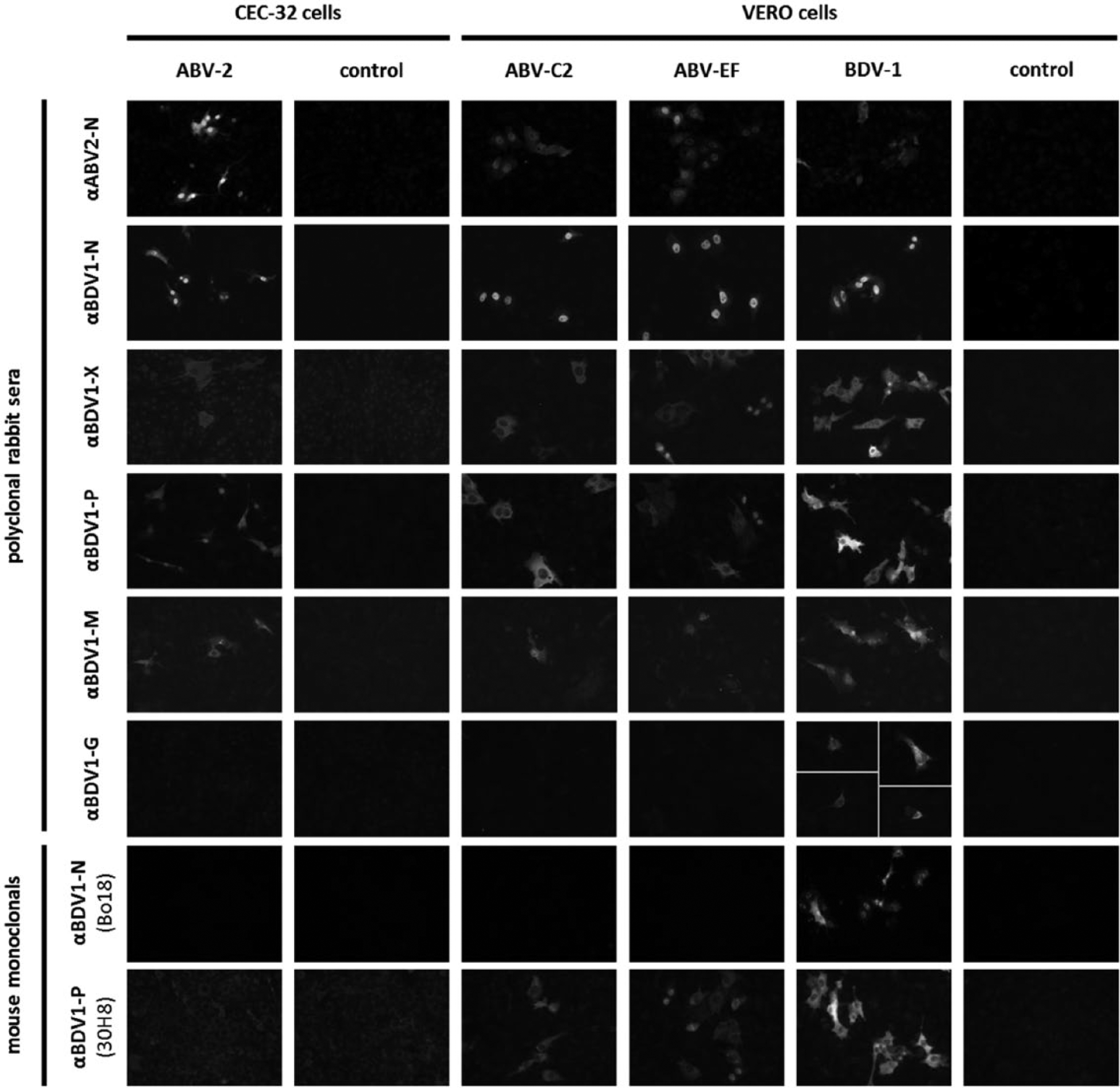
Detection of bornavirus antigen by fluorescent antibody test. Cells persistently infected with the indicated bornavirus strains from 4 different phylogenetic groups were mixed with uninfected cells to contain approximately 5% infected cells, except for serum αBDV1-G for which completely infected cultures were used. Polyclonal rabbit sera and mouse monoclonal antibodies directed against different bornavirus proteins were used together with cyanine (Cy)3-labeled secondary antibodies.
All antibodies showed highest signal intensity when detecting their homologous genotype. Polyclonal rabbit sera directed against N, X, phosphoprotein (P), and M proteins also detected all tested heterologous genotypes with sufficient signal intensity (Fig. 2). In contrast, the mAb 30H8 (anti–BDV1-P) cross-reacted with the passerine genotypes ABV-C2 and ABV-EF but not with the psittacine genotype ABV-2. A similar lack of cross-reactivity of antibody 30H8 was also observed for other members of this phylogenetic group, specifically ABV-1, ABV-4, and ABV-7 (data not shown). Mouse monoclonal Bo18 (anti–BDV1-N) detected genotypes BDV-1 (Fig. 2) and BDV-2 (data not shown), but none of the ABV strains tested. Similarly, rabbit anti–BDV1-G did not cross-react with ABV genotypes (Fig. 2).
In agreement with studies on cell cultures persistently infected with BDV, ABV N proteins were detected mainly in the nucleus and often formed nuclear dots. In contrast, P, M, and X proteins were located predominantly in the cytoplasm, but to some extent also appeared as nuclear dots (Fig. 2).2,3,32,33 Detection of glycoprotein (G) was restricted to only a small proportion of BDV-infected cells in which it was mainly visible in cytoplasmic structures (Fig. 2). This is in congruence with previous findings in cell cultures and in brains of infected rats and horses.5,22,25
The epitope of the mAb Bo18 is located at amino acid positions 19–27 of the BDV-1 N protein. 1 Alignment of 72 BDV sequences confirmed the Bo18 epitope to be conserved in almost all strains, including BDV-2 No/98 (GenBank accession no. AJ311524). The notable exception was strain BDV-1 H215 (KJ950616), which is consistent with the previously observed failure of Bo18 to bind the N protein of this strain.4,26 None of the analyzed ABV N protein sequences contained the conserved Bo18 epitope (Fig. 3A). Five linear epitopes were previously identified for the polyclonal anti–BDV1-M serum. 2 In agreement with the good cross-reactivity of this serum, 3 of these epitopes are highly conserved among bornaviruses (Fig. 3B). The epitopes detected by the remaining antibodies and sera used in the present study are not known.
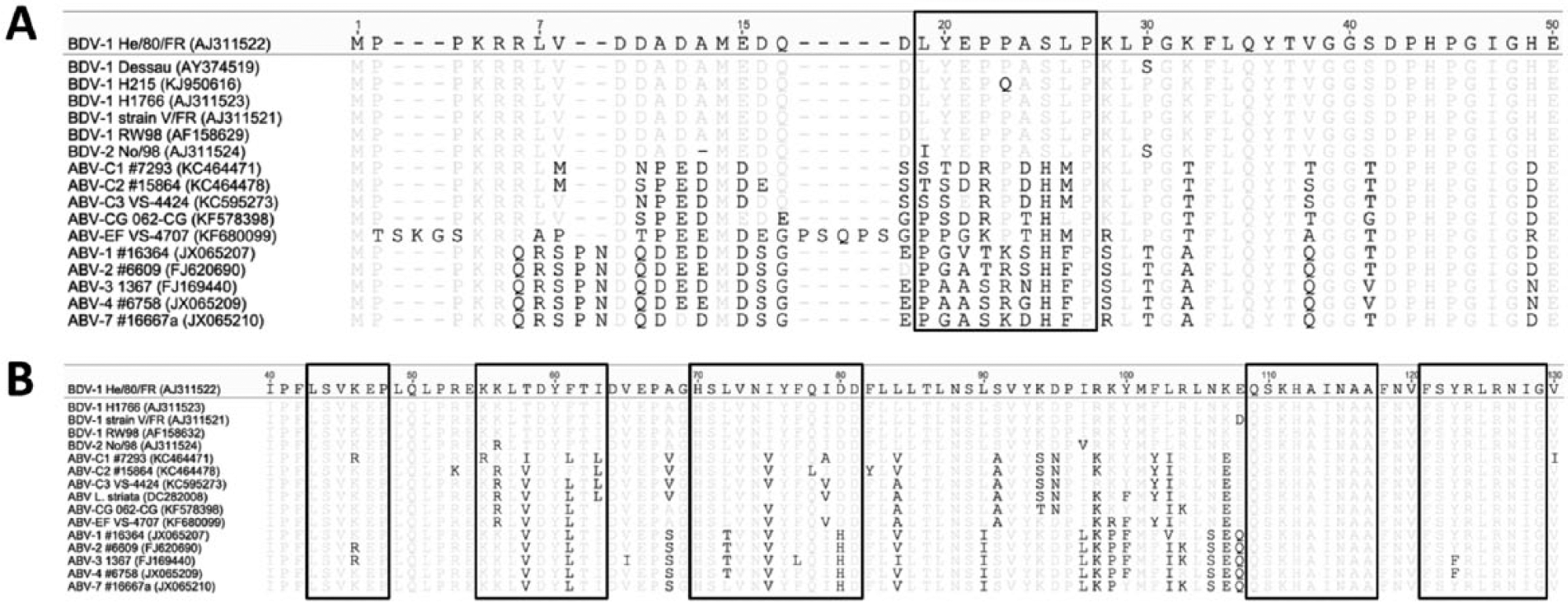
Conservation of the epitopes of 2 anti-bornavirus antibodies. Amino acid sequences of complete nucleoprotein (N; panel
Antigenic diversity affects sensitivity of bornavirus-specific antibody detection by IFAT
In order to test the effect of antigenic variability on the detection of bornavirus-specific antibodies by IFAT, sera from animals infected with known bornavirus genotypes were tested for their cross-reactivity with target viruses from 8 different genotypes (Fig. 4). Persistently infected target cells were seeded with an excess of uninfected cells to obtain approximately 5% infected cells. The small proportion of brightly stained bornavirus-positive cells in each well allowed for an easy discrimination between specific signals and background fluorescence. This was particularly important for sera collected from canaries, which, in the current study, generally exhibited more extensive nonspecific staining as compared to psittacine or mouse sera (Fig. 5).
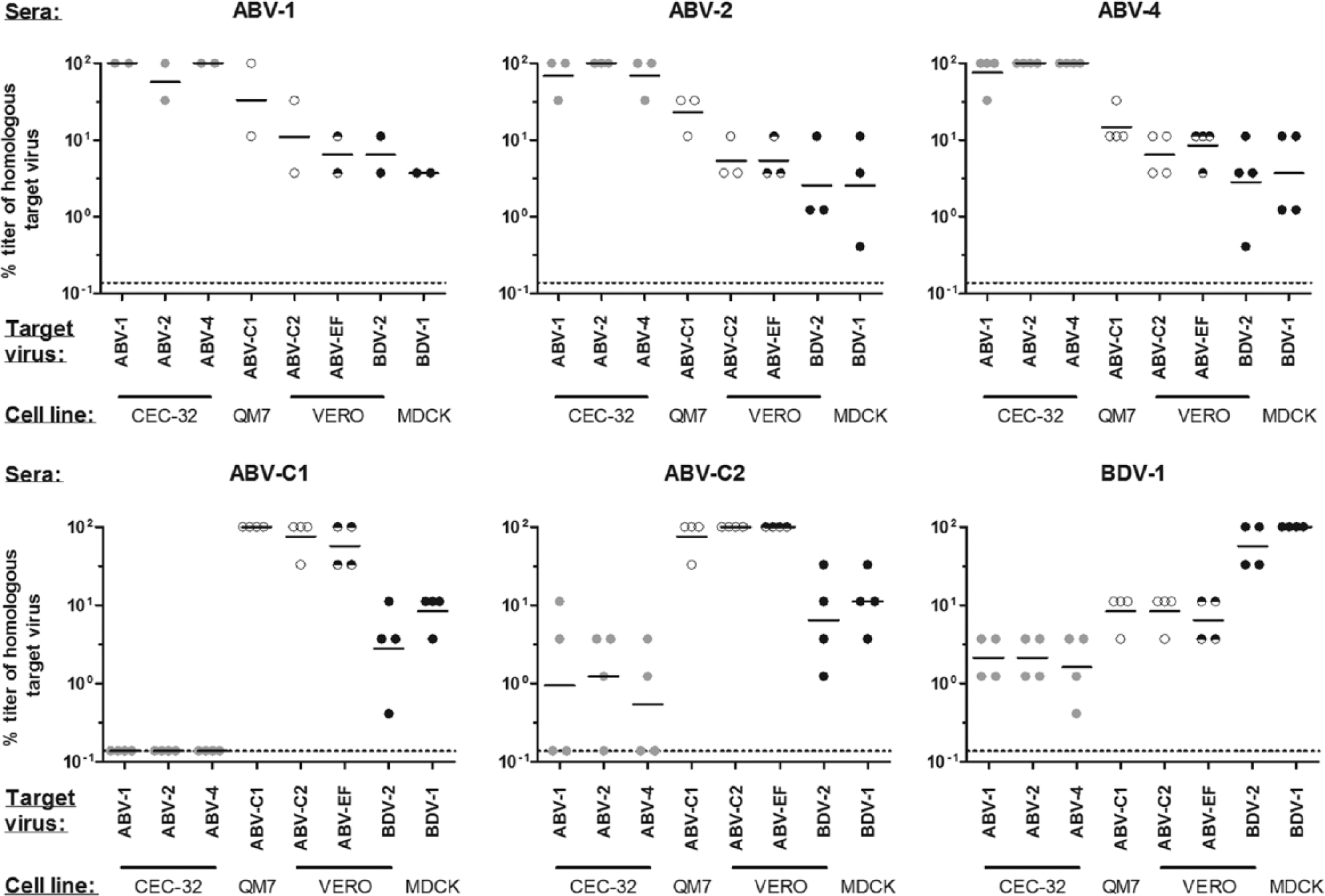
Genetic variability of bornaviruses influences sensitivity of antibody detection by indirect fluorescent antibody test. Sera from animals infected with 6 different bornavirus genotypes were tested for their cross-reactivity using target cells infected with 8 different bornavirus genotypes. Endpoint titers were determined and, for each serum antibody, titers are presented as percentages of the titer obtained with the homologous target virus. Dotted lines represent the detection limit of the assay.
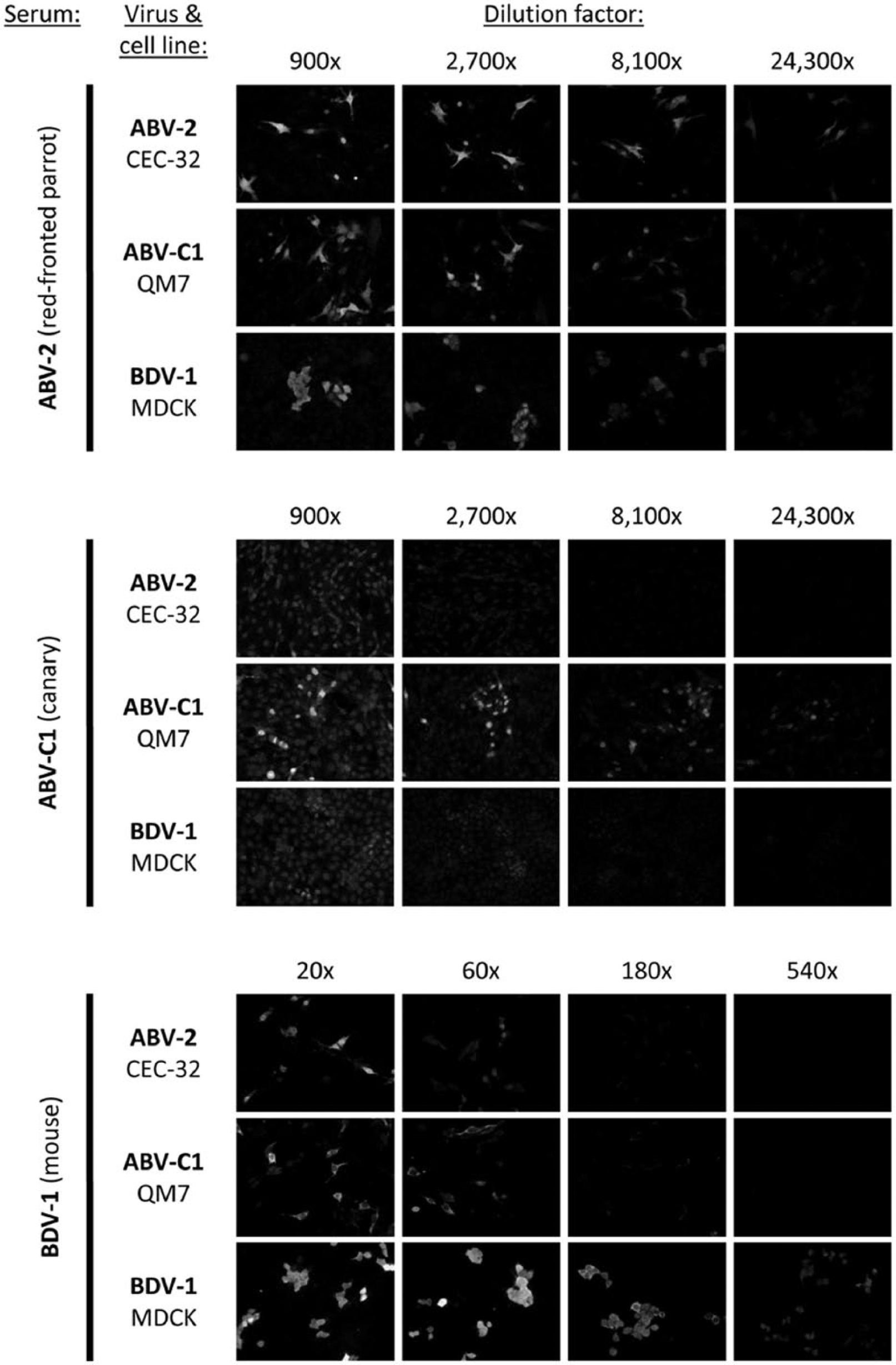
Detection of bornavirus-specific antibodies using indirect fluorescent antibody test. Serial dilutions of 3 representative sera from bornavirus-infected animals (directed against genotypes ABV-2, ABV-C1, or BDV-1) were tested on target cells persistently infected with the same 3 bornavirus genotypes. Wells containing approximately 5% infected cells were used to allow for easier discrimination between bornavirus-specific staining and background fluorescence.
For all sera, calculated antibody titers were highest when the homologous target virus was used for the assay (Figs. 4, 5). Good cross-reactivity between closely related genotypes belonging to a common phylogenetic group was observed with hardly any loss in sensitivity (Fig. 4). For instance, antibodies directed against ABV-1 were detected well using target cells infected with ABV-2 or ABV-4 and vice versa. Detected titers decreased markedly by 1–2 log units when target cells were infected with less closely related bornaviruses, such as ABV-C2, ABV-EF, BDV-1, or BDV-2.
Similarly, good cross-reactivity was observed between the closely related canary genotypes ABV-C1 and ABV-C2, as well as between the mammalian genotypes BDV-1 and BDV-2, whereas cross-reactivity with genotypes of other phylogenetic groups was largely reduced, resulting in titer reduction of up to 3 log units (Fig. 4). Surprisingly, detection of antibodies directed against ABV-C1 or ABV-C2 was not markedly diminished when using ABV-EF as the target virus. In general, the cross-reactions between the canary group (ABV-C1 and -C2) and the mammalian group (BDV-1 and -2) appeared to be stronger than the cross-reactions of both groups with the psittacine group (ABV-1, -2, and -4; Fig. 4).
Antigen expression by different cell lines does not markedly influence antibody detection
Because ABV strains do not replicate equally well in all cell types,29,30 different avian and mammalian cell lines were used in parallel during the present study. To test whether the choice of the target cell line influences antibody detection, permissive cell lines were infected with ABV-4, ABV-C1, or BDV-1 isolates and subsequently tested with homologous sera. For ABV, no effect on the sensitivity was observed. When the avian cell lines CEC-32 and QM7 were infected with BDV-1 and used as target cells, antibody titers of homologous sera were slightly reduced compared to mammalian Vero or MDCK cells (Fig. 6).
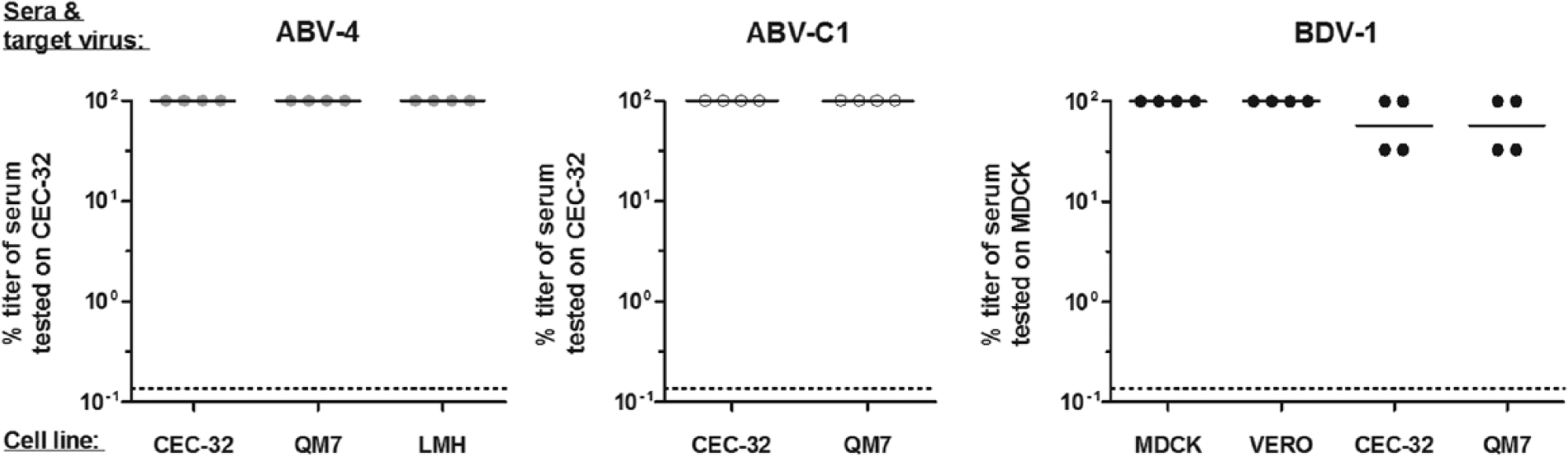
The choice of target cell line used for indirect fluorescent antibody test does not influence sensitivity of antibody detection. Sera from animals infected with ABV-4, ABV-C1, or BDV-1 were tested on different cell lines infected with the homologous target virus. Antibody titers were calculated as percentages of the titer obtained with either CEC-32 cells (ABV-2 and ABV-C1) or MDCK cells (BDV-1). Dotted lines represent the detection limit of the assay.
Discussion
Avian bornavirus–induced disease imposes a major threat to captive psittacine populations and possibly also to wild bird populations. To date, at least 13 ABV genotypes have been discovered in different species.15,18,29–31,34,35 It is therefore of great importance to develop and evaluate reliable tools for the detection of ABV infections that are able to cope with this high genetic variability. Intra vitam diagnosis is at present based mainly on a combination of RNA detection by RT-PCR and detection of ABV-specific antibodies by either IFAT or ELISA. Viral antigen may be detected by immunohistochemical staining directly from tissue samples or by FAT following virus isolation in cell culture.20,23,24,27,29–31,34,35 Reverse transcription PCR assays detecting a broad range of bornaviruses are available, but sensitivity may vary considerably among different genotypes.30,31 The current study was designed to investigate the impact of antigenic variability of bornaviruses on antigen and antibody detection.
In a first approach, antibodies directed against distinct antigens of genotypes BDV-1 and ABV-2 were tested by FAT with 4 bornavirus genotypes from different phylogenetic groups. Polyclonal rabbit sera directed against N, X, P, and M proteins showed good cross-reactivity with all 4 genotypes tested. This is in agreement with previous studies.20,24,27,29–31,34,35 In contrast, a polyclonal anti–BDV-G protein serum and the mAbs anti–BDV1-N Bo18 and anti–BDV1-P 30H8 are not suitable to detect a broad spectrum of bornaviruses, as they showed reactivity only with the homologous virus or a small number of additional genotypes.
Serologic analyses were performed by IFAT, and patient sera were titrated using a broad range of bornavirus genotypes as target viruses. Sera directed against specific bornaviruses did cross-react with heterologous target viruses. However, the sensitivity of detection was markedly reduced when target viruses were not closely related genetically. This is in contrast to the results of a previous study 12 that reported antibody titers in sera from naturally ABV-infected psittacines to be similar when using ABV-infected CEC-32 or BDV-infected MDCK cells. The results presented herein emphasize that knowledge on the range of bornavirus genotypes expected in the tested animals is important for the choice of the most appropriate diagnostic test. Although the present study was solely based on FATs, a similar impact on diagnostic sensitivity is expected for antibody detection by ELISA or Western blot.
Borna disease virus antigens are frequently used worldwide for serological screenings of animals and human beings, and positive reactions are often considered indicative of BDV infection.9–11,36,37 The current study clearly demonstrates that tests using BDV antigens may also detect cross-reactive antibodies induced by other bornaviruses, though with a lower sensitivity. It is therefore strongly recommend that antibodies detected with any target bornavirus should be regarded as bornavirus-reactive rather than genotype-specific.
Interestingly, analysis of sera from canaries experimentally infected with either ABV-C1 or ABV-C2 showed more prominent cross-reactivity with mammalian BDV strains as compared to genotypes of the phylogenetic group “Psittacines I.” Similarly, sera from BDV-1–infected mice and the mAb 30H8, which is directed against the BDV-1 P protein, revealed stronger signals with genotypes ABV-C1, ABV-C2, and ABV-EF than with psittacine genotypes. These results indicate that the known phylogenetic ABV groups detected in passerines are more closely related to mammalian BDVs than to the ABV group “Psittacines I.”
In summary, the present study demonstrated a considerable antigenic diversity among the Bornaviridae family, which may markedly influence the detection of anti-bornavirus antibodies. Considering the variety of bornaviruses detected since 2008, it is likely that new genotypes will continue to be identified. A better understanding of antigenic relationships within the virus family and the availability of reliable and standardized diagnostic assays will be important to cope with this genetic diversity.
Footnotes
Acknowledgements
The authors like to thank Felix Röttele for excellent technical assistance and Wolfgang Garten, Jürgen Richt, Keizo Tomonaga, and Lothar Stitz for providing antibodies against BDV proteins.
a.
Davids Biotechnologie, Regensburg, Germany.
b.
Kindly provided by Keizo Tomonaga, Kyoto, Japan.
c.
Kindly provided by Wolfgang Garten, Gießen, Germany.
d.
Kindly provided by Jürgen Richt, Tübingen, Germany.
e.
Kindly provided by Lothar Stitz, Tübingen, Germany.
f.
BD, Franklin Lakes, NJ.
g.
Roth, Karlsruhe, Germany.
h.
Sigma-Aldrich Chemie GmBH, Munich, Germany.
i.
Jackson Immunoresearch Laboratories, Newmarket, United Kingdom.
j.
Vector Laboratories Inc., Burlingame, CA.
k.
GATC Biotech AG, Constance, Germany.
l.
Geneious Pro 6.1.6, Biomatters, Auckland, New Zealand.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
All work was funded directly by the institutes involved.
