Abstract
Bovine respiratory disease (BRD) causes severe economic losses to the cattle farming industry worldwide. The major bacterial organisms contributing to the BRD complex are Mannheimia haemolytica, Histophilus somni, Mycoplasma bovis, Pasteurella multocida, and Trueperella pyogenes. The postmortem detection of these organisms in pneumonic lung tissue is generally conducted using standard culture-based techniques where the presence of therapeutic antibiotics in the tissue can inhibit bacterial isolation. In the current study, conventional and real-time polymerase chain reaction (PCR) assays were used to assess the prevalence of these 5 organisms in grossly pneumonic lung samples from 150 animals submitted for postmortem examination, and the results were compared with those obtained using culture techniques. Mannheimia haemolytica was detected in 51 cases (34%) by PCR and in 33 cases (22%) by culture, H. somni was detected in 35 cases (23.3%) by PCR and in 6 cases (4%) by culture, Myc. bovis was detected in 53 cases (35.3%) by PCR and in 29 cases (19.3%) by culture, P. multocida was detected in 50 cases (33.3%) by PCR and in 31 cases (20.7%) by culture, and T. pyogenes was detected in 42 cases (28%) by PCR and in 31 cases (20.7%) by culture, with all differences being statistically significant. The PCR assays indicated positive results for 111 cases (74%) whereas 82 cases (54.6%) were culture positive. The PCR assays have demonstrated a significantly higher rate of detection of all 5 organisms in cases of pneumonia in cattle in Northern Ireland than was detected by current standard procedures.
Bovine respiratory disease (BRD) is a global problem causing severe economic losses to the cattle farming industry through mortality, loss of production, and treatment costs. 12 In particular, BRD is the most frequent cause of mortality in calves and adult cattle in Northern Ireland, and based on postmortem submissions to the Veterinary Sciences Division of the Agri-Food and Biosciences Institute (VSD-AFBI) during 2012, was recorded as accounting for 43% of deaths in juveniles (1–5 months), 40% of deaths in 6–12-month weanlings, 16% of deaths in adult cattle, and was the second most frequently diagnosed condition (13%) in neonatal calves (All-island Animal Disease Surveillance Report 2012, http://www.afbini.gov.uk/index/publications/featured-publications.htm). The major bacterial pathogens associated with the BRD complex are Mannheimia haemolytica, Histophilus somni, Mycoplasma bovis, Pasteurella multocida, and Trueperella pyogenes.9,12 All these organisms are ubiquitous in the cattle population but can infiltrate into the lungs following stress or viral infection. 9
The diagnostic detection of these pathogens in pneumonic lung tissue taken at necropsy at AFBI is by standard culture techniques and, in the case of Myc. bovis, by a capture–enrichment sandwich enzyme linked immunosorbent assay (sELISA) confirmed by culture. 4 Isolation of these pathogens can be negatively influenced by the presence of antibiotics in the tissue administered during the terminal stages of pneumonia, or by overgrowth of less fastidious bacteria present during culture. A study conducted at AFBI on Myc. bovis detection using a real-time polymerase chain reaction (qPCR) assay showed that infection rates for this organism were underdiagnosed by the existing sELISA method. 6 The aim of the current study was to apply PCR assays to the detection of the bacteria most commonly associated with pneumonia in cattle from lung samples taken at postmortem examination, and to compare the results with those obtained with the currently used diagnostic techniques.
Grossly pneumonic bovine lung samples (n = 150) were obtained from animals submitted to AFBI for postmortem examination between February and July 2013. In all cases, standard bacteriological culture was done for M. haemolytica, H. somni, P. multocida, and T. pyogenes.10,11 Approximately 1 g of lung sample was homogenized in mycoplasma transport medium, 8 and a 100-µl aliquot was taken for Myc. bovis testing by the capture–enrichment sELISA. In addition, DNA was isolated from all lung samples using a commercial DNA isolation kit a and stored at −20°C for testing by conventional PCR or qPCR.
Individual conventional PCR assays using previously described primers b were performed for M. haemolytica, 13 H. somni, 2 P. multocida, 14 and T. pyogenes. 16 Each PCR was carried out using a commercial PCR kit. c Each 25-µl reaction contained 1× PCR buffer, 200 µM of each deoxyribonucleotide triphosphate, 0.5 µM of each primer, 2.5 U of DNA polymerase, and 5 µl of template DNA. Amplification was carried out on a conventional thermocycler d with the following cycling parameters: activation at 95°C for 15 min followed by 35 cycles of denaturation at 94°C for 1 min, annealing at 67°C for 1 min for M. haemolytica, 57°C for 1 min for H. somni, 59°C for 1 min for P. multocida, 62°C for 1 min for T. pyogenes, extension at 72°C for 1 min, and a final extension of 72°C for 10 min. A qPCR using previously described primers and hydrolysis probe b was performed for Myc. bovis. 15 The qPCR reactions were performed using a commercial PCR kit. e Each 25-µl reaction contained 1× master mix, 0.4 µM of each primer, 0.2 µM of hydrolysis probe, 1× internal control, and 2 µl of template DNA. Amplification was done on a real-time thermocycler f using the following conditions: 5 min at 95°C then 40 cycles of 15 sec at 95°C and 30 sec at 60°C. DNA was extracted from laboratory strains of the 5 target organisms, using a commercial DNA isolation kit, a and was used as positive controls along with negative controls in all amplification runs for each PCR.
The Myc. bovis sELISA was performed as previously published.4,7 The sELISA results were recorded as obtained but cases were only reported as positive if they were confirmed positive by culture.
Immunohistochemical staining (IHC) for Myc. bovis was carried out on formalin-fixed lung sections from 6 cases that were qPCR positive/sELISA negative, and 10 cases that were qPCR positive/sELISA positive. Sections were prepared with monoclonal antibody 5A10 that is specific to Myc. bovis and Myc. agalactiae 5 and a commercial antibody detection kit. g
Statistical analysis software h using the chi-square test and 2 × 2 tables was used to explore any proportional differences and association on the results obtained by PCR and culture. For the qPCR assay, samples were taken as positive if the threshold cycle value was less than 37, which defined the limit of detection of the assay, determined by serial dilutions of known concentration of Myc. bovis DNA as 60 copies/µl.
Table 1 shows the PCR or qPCR results obtained and the corresponding results from diagnostic culture and sELISA. All of the PCR tests detected a significantly higher number of positive cases than culture (M. haemolytica, Myc. bovis, P. multocida, and T. pyogenes: P < 0.001; H. somni: P = 0.01). The results for M. haemolytica and H. somni reflect the findings of other researchers.3,17 However, culture has been reported to detect a higher number of samples positive for P. multocida than PCR. 3
Summary of polymerase chain reaction (PCR) and culture/enzyme-linked immunosorbent assay results from 150 terminal cases of bovine pneumonia.
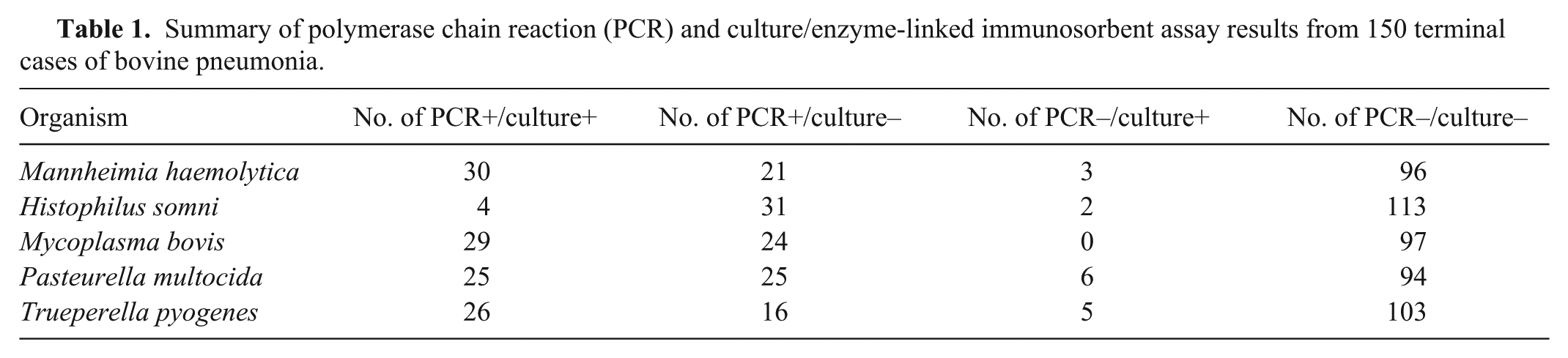
Although there was histological evidence of bacterial involvement in the majority of cases, only 82 of the 150 cases (54.6%) had bacterial culture–positive results. The PCR assays demonstrated positive results for 111 cases (74%) identifying pathogens in 32 more cases than culture or sELISA, thus better supporting the postmortem diagnosis.
Mycoplasma bovis was positive by IHC in 6 of the 10 PCR-positive/sELISA-positive cases and in 4 of the 6 PCR-positive/sELISA-negative cases demonstrating that Myc. bovis had reached the chronic stage of infection in 10 of these animals. The 4 IHC-positive cases in which the sELISA was negative highlight incidences were culture-based techniques fail to diagnose the presence of pathogens in the lung. The failure to demonstrate Myc. bovis by IHC in 6 PCR-positive cases likely indicates the secondary contribution of this organism in these cases. Without the availability of IHC for the other 4 pathogens examined in the study, the relevant contribution of each organism in cases demonstrating multiplicity of infection is impossible to determine, but during the terminal stages of disease it can probably be assumed that the immune system is unable to counteract invasion by secondary opportunistic pathogens.
The difference in detection rates between culture and PCR for H. somni highlights the difficulties in isolating this organism. Histophilus somni can readily be overgrown by other organisms such as P. multocida, Klebsiella, Escherichia coli, and Proteus spp., which were isolated in many of the H. somni PCR-positive/culture-negative cases examined in this study and may explain why H. somni was never isolated alongside M. haemolytica, P. multocida, or T. pyogenes (Table 2) under the cultural conditions used. Histophilus somni was only isolated once when another bacterium (E. coli, data not shown) was also present. In contrast, the PCR results demonstrate the comparable frequency of H. somni coinfection with the other pathogens. Of the 35 cases in which H. somni was detected by PCR, M. haemolytica was also detected 13 times (37%), P. multocida was detected 18 times (51.4%), and T. pyogenes was detected 14 times (40%). While BRD is a multifactorial disease in which multiple pathogens can infect the lung, it is not possible to determine the relative primary or secondary significance of the pathogens by the PCR or culture methods used in the current study. Table 3 shows the number of cases in which multiple pathogens were detected. The PCR assays identified coinfection in 45.3% of total cases, better illustrating the occurrence of multifactorial infection in comparison to the 22.7% of cases identified by culture.
Frequency of coinfection between pathogens.*
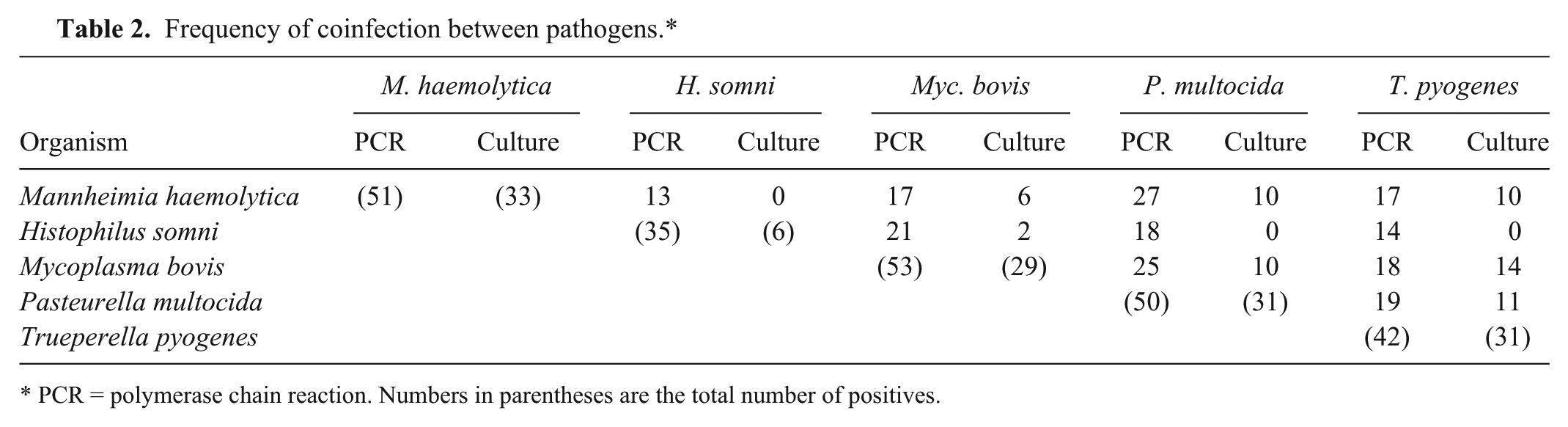
PCR = polymerase chain reaction. Numbers in parentheses are the total number of positives.
Multifactorial nature of infection.*
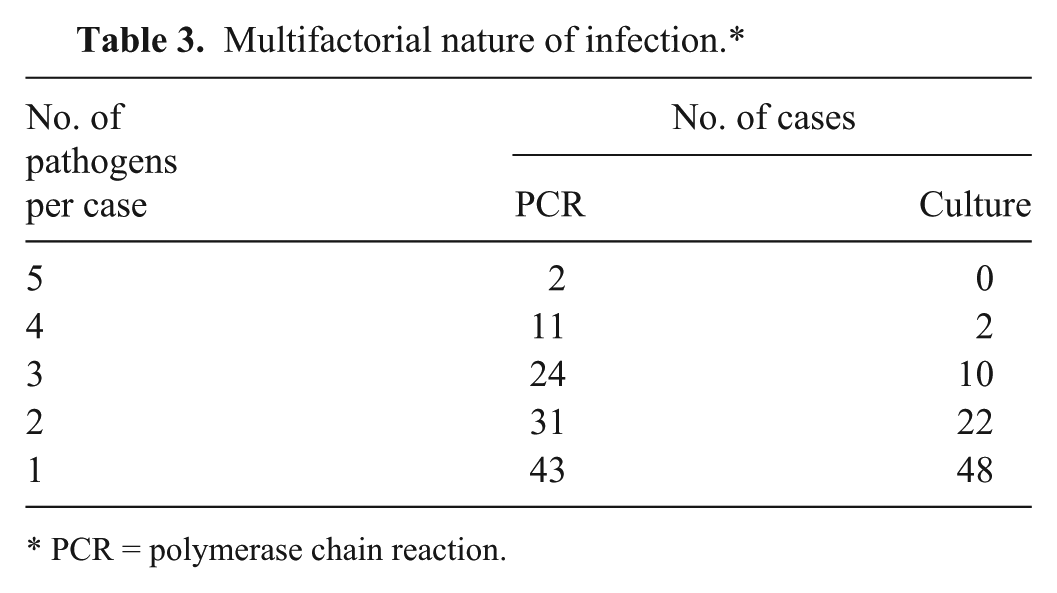
PCR = polymerase chain reaction.
The higher number of positive PCR results obtained compared to the diagnostic use of culture may not be surprising given the use of antibiotics to treat pneumonia cases on the farm. An advantage of PCR is its ability to detect growth-inhibited and dead bacteria following antibiotic therapy.1,17,18 In 57 of the 150 cases (38%), the administration of antibiotic treatment was recorded on the postmortem submission form but this figure most likely does not reflect the true level of antibiotic use as full histories are not always given. The results of the current study demonstrate that employing PCR assays as ancillary tests in the diagnostic laboratory would greatly improve the diagnosis of bacteria contributing significantly to the terminal stages of pneumonia and would provide relevant information for the prospective treatment or vaccination program in the respective herd.
Footnotes
a.
DNeasy blood and tissue kit, Qiagen Ltd, Manchester, United Kingdom.
b.
Primers and hydrolysis probes, Sigma-Aldrich Company Ltd, Dorset, United Kingdom.
c.
HotStarTaq DNA polymerase kit, Qiagen Ltd, Manchester, United Kingdom.
d.
Applied Biosystems GeneAmp PCR system 9700, Life Technologies Ltd, Paisley, United Kingdom.
e.
QuantiFast pathogen PCR + IC kit, Qiagen Ltd, Manchester, United Kingdom.
f.
Rotor-Gene Q, Qiagen Ltd, Manchester, United Kingdom.
g.
DakoReal, Envision detection system, Dako Denmark A/S, Glostrup, Denmark.
h.
STATA 11.1 software, Stata Corp., College Station, TX.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
