Abstract
A multiplex amplification refractory mutation system reverse transcription polymerase chain reaction (ARMS RT-PCR) was developed for the differential diagnosis of
Introduction
Since the initial discovery of the link with feline lymphosarcoma in 1964,
Virus isolation remains the gold standard for the diagnosis of FeLV infection. Both a fluorescent antibody test (FAT) and an enzyme-linked immunosorbent assay (ELISA) against the p27 core protein are used, but the cats have to have been infected for 6 and 4 weeks to test positive, respectively. 13 Polymerase chain reaction (PCR) and real-time PCR (qPCR) assays targeting the FeLV proviral DNA or RNA were developed for the diagnosis of FeLV. 11 Because positive results can be obtained less than 2 weeks after exposure, these molecular tests are considered sensitive approaches. 16 The progressive infection is characterized by positive ELISA and FAT results and high plasma viral RNA or proviral DNA loads, whereas only a positive result for proviral DNA is often seen in the regressive infection. 27 Under certain conditions, including those found in discordant cats that present positive ELISA and negative FAT results and in those kittens that receive a maternal antibody that interferes with the detection of the FeLV antigen, the animals should be further considered sources for disease spread if they present positive PCR results. 27
To date, the vaccination of cats remains an important strategy against FeLV-associated diseases. 18 Although different efficacies have been reported, recombinant subunit vaccines, canary-vectored recombinant vaccines, and inactivated whole virus vaccines are the most recommended. 15 Even though vaccines cannot protect cats from an integration of proviral DNA and viral replications, less persistent viremia and FeLV-associated diseases and longer survival times have been observed. 15 Therefore, repeated vaccination plans combined with a screening program appear to be beneficial to cat owners. 30 Accordingly, as well as potency, the control of the safety of inactivated FeLV vaccines has become a significant issue for both users and producers due to the unique lifecycle of FeLV.
While the FeLV-positive rate of sick cats based on ELISA and FAT was 13.1%, a rate of only 2.3% was found for random healthy cats in North America. 13 The average positive rate for sick cats in Brazil and Canada is 16.4% and 6%, respectively, by ELISA.5,20 The use of PCR for screening FeLV in Brazil revealed positive rates for sick cats and healthy cats of 47.2% and 47.4%, respectively. 5 The positive rate obtained in a survey conducted in Spain was 35.7% by ELISA. 10 The FeLV-positive rate of cats in Brazil by PCR is consistent with the average prevalence of 30–40% found worldwide because PCR can detect the proviral DNA in FeLV-infected cats, which may not exhibit soluble antigens during the test. 17 At present, no information is available on the prevalence of FeLV in Taiwan.
Single nucleotide polymorphisms (SNPs) are found in different individual isolates, and these changes account for sequence variations among these isolates. Such SNPs are useful for differential diagnosis because most DNA markers are based on genetically stable areas of the genome.
33
Since 2011, both multiplex amplification refractory mutation system reverse transcription polymerase chain reaction (ARMS RT-PCR) and high-resolution melt analysis (HRMA) have been used for SNP analysis. These procedures exhibit several advantages, including the ability to check at least 104 samples in 5 working days, wide availability, and rapid turnaround time.19,33 When specific terminal 3′-nucleotides are included in the primers, ARMS RT-PCR can selectively amplify the respective alleles.4,24 This procedure has been used for the differentiation of wild-type and vaccine strains of
The aim of the current study was to analyze the nucleotide sequences of the long terminal repeat (LTR) and the p27 genes of Taiwan isolates of FeLV and 2 contemporary vaccines. Based on point mutations inferred from the above analysis, both ARMS RT-PCR and ARMS qRT-PCR with HRMA protocols were developed for the differential diagnosis of field isolates and vaccine strains. Such approaches can be used to detect the replication of a vaccine strain that has not been properly inactivated.
Materials and methods
Sample collection and analysis
A total of 55 blood samples from sick cats admitted to local veterinary clinics around Taiwan were transported to the molecular diagnostic laboratory of the Department of Veterinary Medicine of National Chung Hsing University. The LTR gene, which is located in an exogenous FeLV element, and the p27 gene were sequenced for phylogenetic analysis.
P27 immunochromatography analysis
One hundred and fifty microliters of blood in 0.25 mg of ethylenediamine tetra-acetic acid (EDTA) were mixed with the horseradish peroxidase–labeled anti-FeLV antigen conjugate, and the mixture was then analyzed using a commercial kit a according to the manufacturer’s instructions.
Isolation of DNA and RNA from white blood cells
Genomic DNA was extracted using a commercial kit b according to the manufacturer’s instructions. Three hundred microliters of blood in 0.5 mg of EDTA were mixed with 1 ml of TRIzol reagent. c After a 5-min incubation, 200 μl of chloroform was added to the lysate. After centrifugation, the supernatant was mixed with an equal volume of isopropanol and then stored at −70°C overnight. After centrifugation, the RNA was pelleted and dissolved in diethylpyrocarbonate (DEPC)-treated double-distilled water (DDW).
Polymerase chain reaction primers
The FeLV-U3-F(1), FeLV-G-R(1), FeLV-U3-F(2), and FeLV-G-R(2) primers were exactly as described in a previous report (Table 1). 23 The P27 OF 744, P27 OR 744, P27 IF 744, and P27 IR 744 primers were designed after multiple alignments of the p27 gene of FeLV. After the p27 genes from vaccine SP and P strains and wild-type strains of FeLV (TW1–TW12) were sequenced, a point mutation was identified between the vaccine and wild-type strains. The P27 vaccine-F and P27 wild-R primers were designed to specifically target the vaccine and wild strains, respectively. Detailed information on the primers is shown in Figure 1 and Table 1.
Primers for the detection of
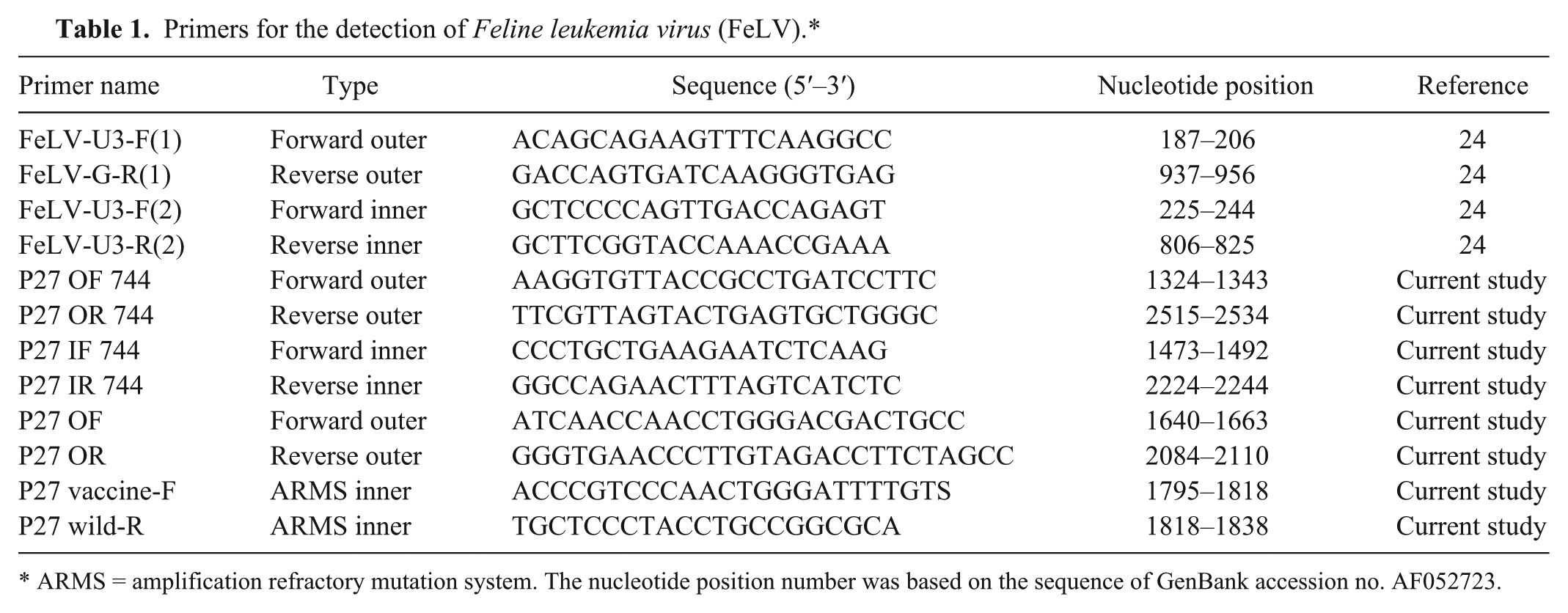

ARMS = amplification refractory mutation system. The nucleotide position number was based on the sequence of GenBank accession no. AF052723.
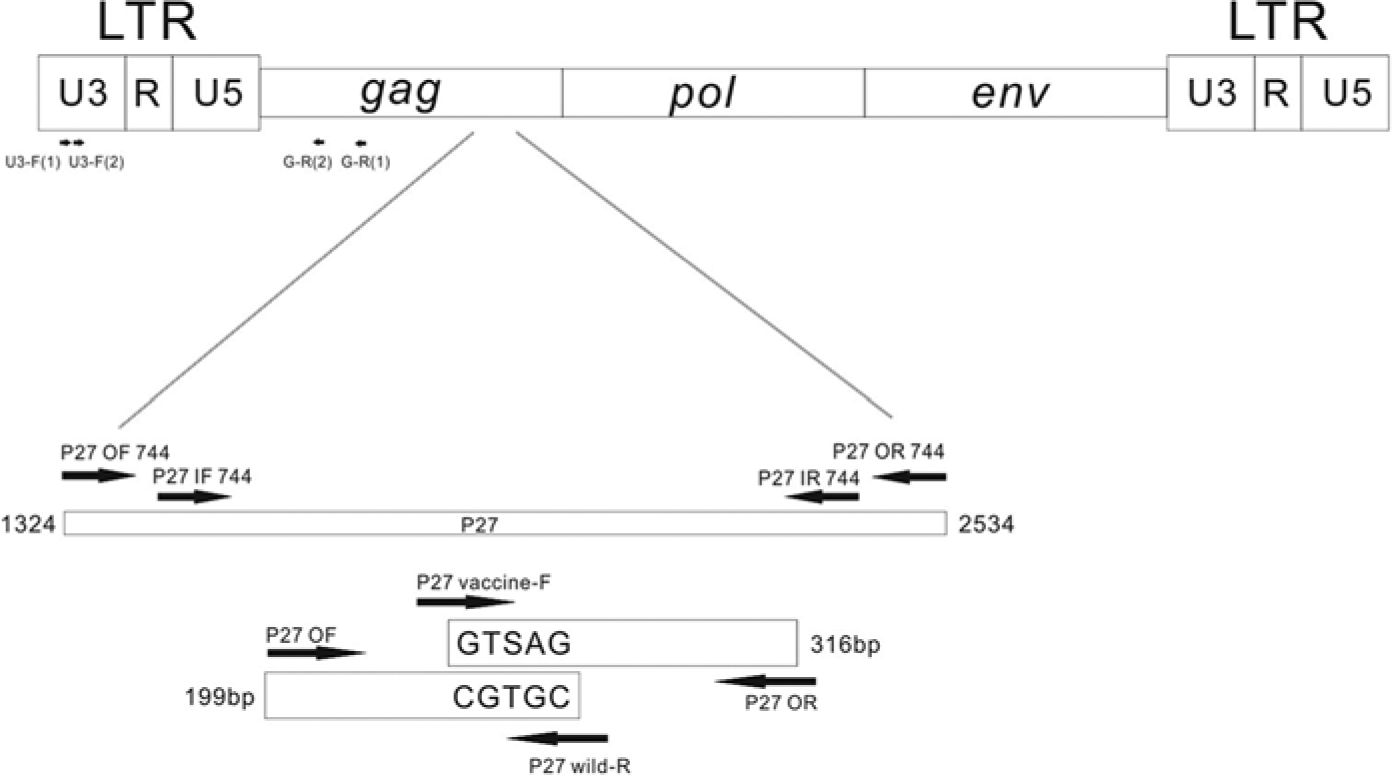
Regions of the
Polymerase chain reaction reverse transcription PCR, and cloning of sequence
The extracted RNA was heated to 65°C and then placed on ice. The first-strand complementary (c)DNA synthesis was performed in a total reaction volume of 12 μl containing 4 μl of 5× first-strand buffer, 2 μl of 0.1 M dithiothreitol, 1 μl of ribonuclease inhibitor (40 units/μl), 1 μl of M-MLV reverse transcriptase
d
(200 units/μl), and 0.1 μM of oligo-dT primer at 37°C for 50 min and then inactivated at 70°C for 15 min. The PCR reaction was performed in a total reaction volume of 25 μl consisting of 2.5 μl of 10× buffer, 10 μM of each deoxyribonucleotide triphosphate (dNTP), 1.5 mM MgCl2, 0.1 μM of the FeLV-U3-F(1) and FeLV-G-R(1) primers or the P27 OF 744 and P27 OR 744 primers, 0.5 μl of Taq polymerase
e
(5 units/μl), and 3 μl of genomic DNA or cDNA. The PCR conditions were 95°C for 5 min, 35 cycles of denaturation at 95°C for 30 sec, annealing at 50°C for 30 sec, and extension at 72°C for 1 min, and a final extension at 72°C for 10 min. The second PCR was performed using the aforementioned conditions with the following change: the FeLV-U3-F(2) and FeLV-G-R(2) primers or the P27 IF 744 and P27 IR 744 primers were used. Five microliters of the products from the second PCR were analyzed in a 2% agarose gel. For sequencing, the products from the second PCR of the LTR and p27 genes were purified, cloned into a commercial vector,
f
and transformed into
In vitro transcription of FeLV plasmid
The viral RNA was produced using a commercial kit
h
according to the manufacturer’s instructions. Two micrograms of p-TEasy-p27 plasmids were linearized with the
Amplification refractory mutation system RT-PCR and ARMS qRT-PCR combined with high-resolution melt analysis
The cDNA was amplified using a PCR program similar to that described above with the P27 OF 744 and P27 OR 744 primers and then subjected to ARMS RT-PCR in a total reaction volume of 25 μl containing 0.1 μM P27 OF, P27 OR, P27 vaccine-F, and P27 wild-R primers; 5 μl of the PCR products; 2.5 μl of 10× buffer; 10 μM of each dNTP; 1.5 mM MgCl2; and 0.5 μl of Taq polymerase e (5 units/μl). The PCR program was 95°C for 5 min, 35 cycles of denaturation at 95°C for 30 sec, annealing at 58°C for 30 sec, and extension at 72°C for 1 min, and a final extension at 72°C for 10 min. For ARMS qRT-PCR, the PCR mixture contained 0.1 μM P27 OF, P27 OR, P27 vaccine-F, and P27 wild-R primers; 12.5 μl of PCR master mix j ; and 5 μl of the PCR products. The PCR program was 95°C for 5 min and 35 cycles of denaturation at 95°C for 30 sec, annealing at 59.5°C for 30 sec, and extension at 72°C for 1 min. After qRT-PCR, HRMA was immediately performed, and the PCR products were melted from 69°C to 95°C at a rate of 0.05°C/sec. The melting profiles were analyzed using a commercial software k for the generation of a high-resolution melt (HRM) algorithm. The highest and lowest fluorescence values were set to 100 and 0, respectively, to draw a best-fit line to normalize the raw data. The analysis was performed using normalization regions between 78°C and 88°C. The wild-type FeLV (TW1) was assigned as a “genotype,” and the HRM genotype confidence percentage (confidence %), which was defined as the similarity between the strain of interest and the representative genotype, was calculated. If necessary, the Tm for each sample was demonstrated.
Sensitivity and specificity of ARMS RT-PCR and ARMS qRT-PCR
To determine the sensitivity, a series of 106, 105, 104, 103, 102, 10, and 1 copies of transcribed FeLV, which represent the copy number for each strain in the mixed templates (the FeLV P vaccine, FeLV SP vaccine, and FeLV TW strains) per 0.5 ml of sample, were examined by ARMS RT-PCR. Using the ARMS qRT-PCR protocol, 104, 5 × 102, 50, 5, and 1 copies of transcribed FeLV, which represent the copy number for each strain in the mixed templates (the FeLV P vaccine, FeLV SP vaccine, and FeLV TW strains) per 0.5 ml of sample, were also tested. A no-template control (NTC) was included. To determine the specificity, the cDNA from the mixture of P vaccine, SP vaccine, and Taiwan strains of FeLV,
Phylogenetic analysis
The background information of the cats that tested positive for FeLV is shown in Table 2. For the phylogenetic analysis, the LTR and p27 genes from different FeLV strains available in GenBank were included. For the LTR gene analysis, the GenBank accession numbers of the FeLVs included were AB18756, AY364318, AY364319, AY745878, DQ821502, EU48346, EU48348, EU48351, EU48352, EU048356, EU048362, EU048365, EU090946, and M18247. These sequences were compared with the FeLV TW1–TW12, P vaccine, and SP vaccine strains. For the p27 gene, the FeLV TW1–TW12, P vaccine, and SP vaccine strains were compared with the following FeLV GenBank accessionnumbers: AB052723, AB060732, AB187565, AB672612, ABAB673426, AB673427, AB673431, AY364318, AY364319, FCVF6A, JF957361, JF957363, K01643, K01803, M18247, and NC001940. The nucleotide sequences were aligned and edited using the CLUSTALW algorithm of a commercial software. l For each strain pair, the identity and divergence percentages of both the FeLV and p27 genes were calculated. The phylogeny of the aligned nucleotides was analyzed using both the neighbor-joining and the Bayesian methods of a commercial software. l A resampling of the original dataset with a thousand replications was conducted to obtain the bootstrap value. The 50% majority rule consensus tree was constructed through a Bayesian analysis based on the clade credibility value. The mortality rates for cats in clade I and clade II in the current study period were analyzed using the χ2 test.
Background information on cats that tested positive for
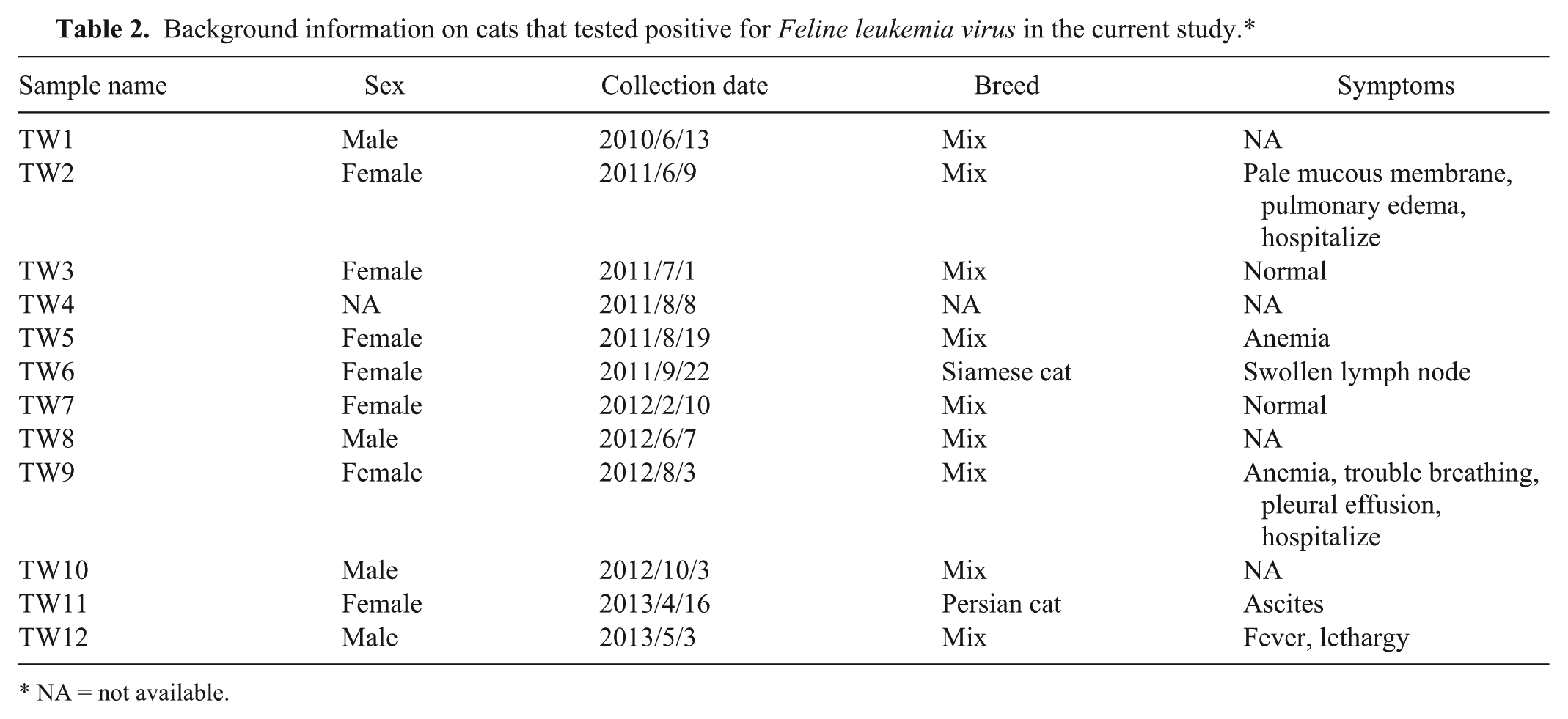
NA = not available.
Results
Samples detected by PCR and p27 immunochromatography and sequence analysis for the design of ARMS RT-PCR primers
Among 55 samples, 12 samples were positive by both the p27 immunochromatography assay and PCR. The partial LTR sequence of exogenous FeLV (nucleotides 1473–2244) was amplified from the above positive samples, named as TW1–TW12 strains, and from the P and SP vaccine strains. All of the LTR sequences were submitted to GenBank, and the accession numbers for TW1–TW12, and the P and SP vaccine strains were KF736909–KF736920 and KF792076, and KF792075, respectively. The GenBank accession numbers for the p27 genes of TW1–TW12, and the P and SP vaccine strains were KF736921–KF736932 and KF792078, and KF792077, respectively. After the sequences were aligned, a T at position 321 was found in all of the TW strains, whereas a G and C were observed in the SP vaccine and P vaccine strains, respectively. Therefore, 2 pairs of ARMS RT-PCR primers were designed for allelic discrimination. The most abundant 3′-termini of the inner primers for the vaccine and TW strains were S and T, respectively. An additional mismatch included 2 bases from the 3′-termini to destabilize the primers (Fig. 1).
Sensitivity of ARMS RT-PCR and ARMS qRT-PCR
The detection limit of the ARMS RT-PCR assay was 102 copies of transcribed FeLV per 0.5 ml of sample. No amplicons were observed in the negative control (Fig. 2A). In the ARMS qRT-PCR, sigmoid curves composed of both an exponential rise and a plateau characterized productive amplifications. The threshold cycle (Ct) values for 104, 5 × 102, 50, 5, and 1 copies of transcribed FeLV per 0.5 ml of sample were 29.45, 32.77, 33.75, 34.58, and 36.26. No amplification of the NTC was found. The findings suggest that the detection limit of ARMS qRT-PCR was 1 copy of transcribed FeLV per 0.5 ml of sample (Fig. 2B). Taken together, the results demonstrate that the sensitivity of the ARMS qRT-PCR assay is 100-fold higher than that of ARMS RT-PCR.
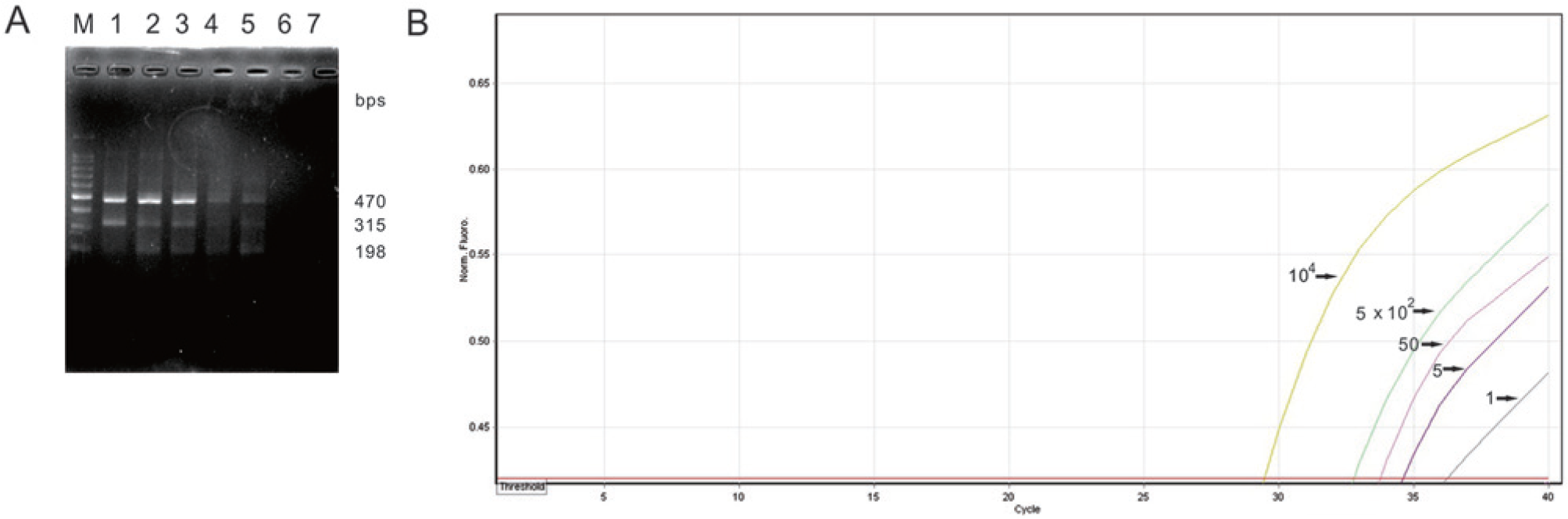
Specificity of ARMS RT-PCR and ARMS qRT-PCR
The amplification of a mixed sample of the P vaccine, SP vaccine, and TW strains using ARMS RT-PCR resulted in 3 fragments of 470, 315, and 198 bp. This finding suggests that the P vaccine, SP vaccine, and TW strains were successfully amplified. No amplification of RNA from feline pathogens and cells, including FIV, FIPV, CRFK, FVR, FCV, FPV, and
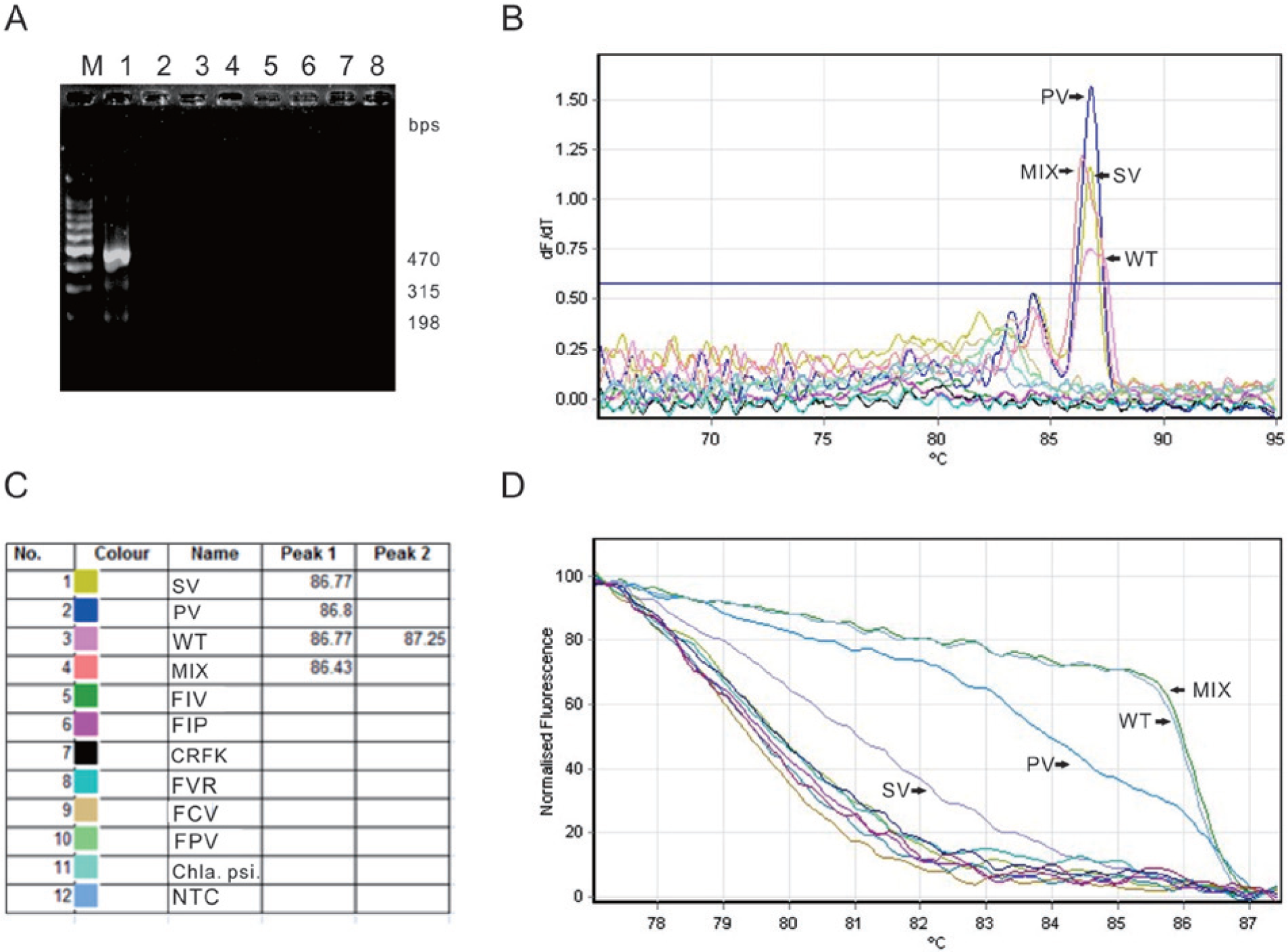
Clinical samples detected by ARMS RT-PCR and ARMS qRT-PCR
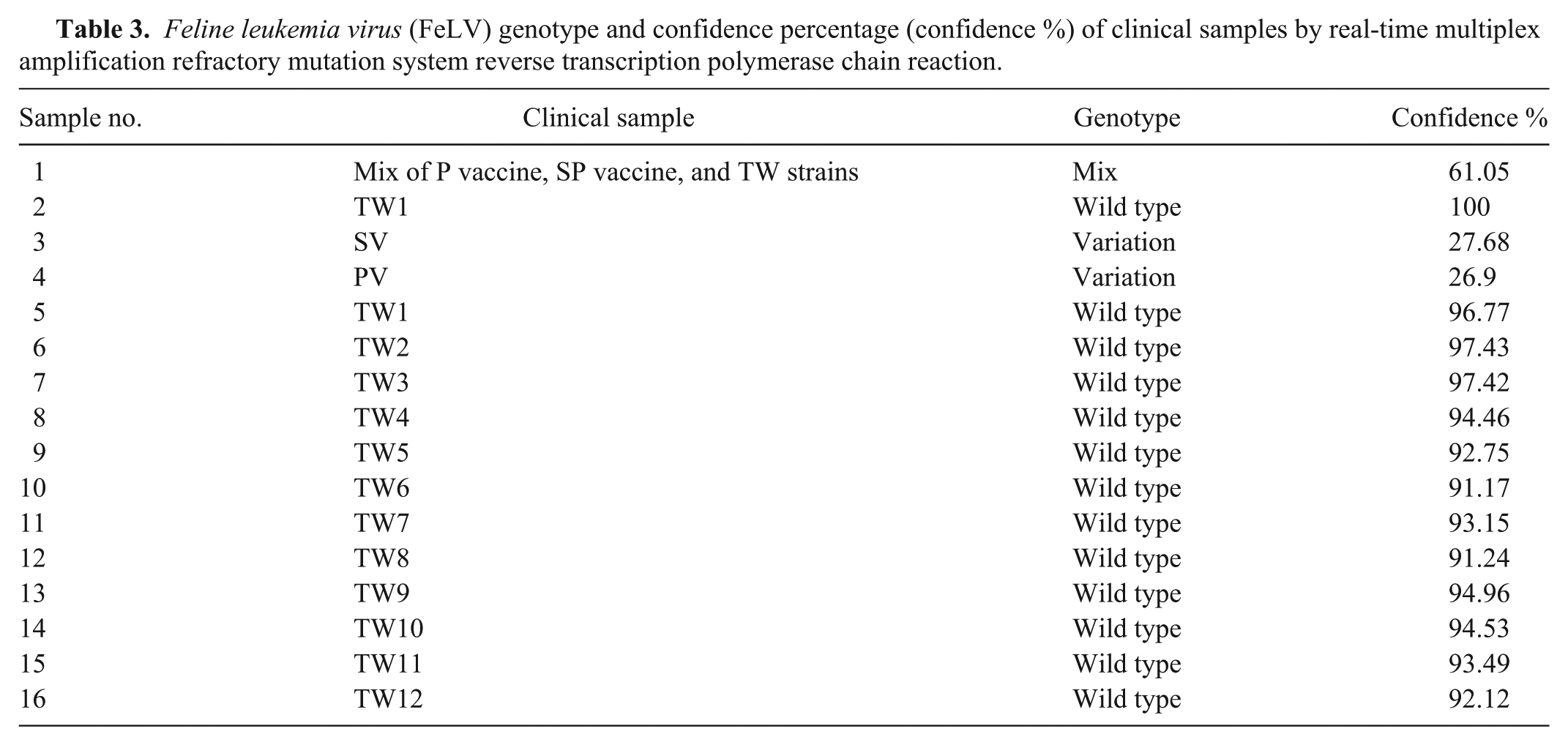
Positive results were only obtained by ARMS RT-PCR for cases TW1–TW12 among the 55 samples tested. Two fragments of 470 and 198 bp were obtained, which suggests that these samples are FeLV wild-type strains (Fig. 4). The confidence percentage values of TW1–TW12 ranged from 91.17% to 97.43%, as determined using ARMS qRT-PCR combined with HRMA (Table 3). This finding further validated that all of the samples were FeLV wild-type strains.
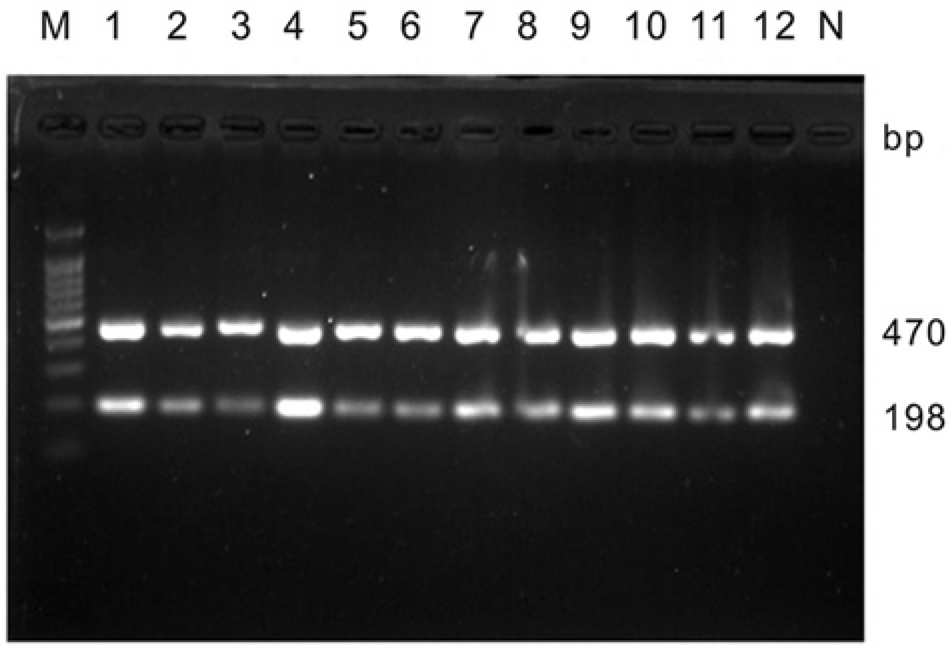
Results of the clinical samples including
Phylogenetic analysis of FeLV vaccine and FeLV TW strains
The sequence identities among the LTR genes from the different TW strains ranged from 96.7% to 100%, but the identity between the LTR gene of the P vaccine strain and the LTR gene of the SP vaccine strain was 95.5% (data not shown). The comparison of the LTR genes from the TW strains with that of the P vaccine strain or that of the SP vaccine strain revealed identity ranges of 97.1–99.7% and 94.8–98.8%, respectively (data not shown). The LTR gene of the P vaccine strain instead of the SP vaccine strain was more closely related to the TW strain. They are clustered in the same group with a bootstrap value of 81. The nucleotide sequences of the LTR genes of the TW strains, vaccine strains, and strains from different countries were analyzed. The neighbor-joining tree indicated that all of the strains were distinct from endogenous FeLVs. Consistent with the above-mentioned sequence identity, the SP vaccine strain had a closer phylogenetic relationship with the FeLV Brazil strains (GenBank accession nos. EU048356 and EU048362; Fig. 5A). The topology of the Bayesian tree of the FeLV LTR phylogeny was highly similar with that of the tree obtained using the neighbor-joining method. The TW strains and the P vaccine strain were clustered in the same group with a clade credibility value of 0.95. A more detailed grouping of the TW strains and the P vaccine strain was observed with the Bayesian analysis. The TW2, TW4, TW7, and TW8 strains, P vaccine strain, and other TW strains were paraphyletic with a clade credibility value of 0.84. The P vaccine strain was closely related to the TW4 and TW7 strains, which are represented as a monophyletic group with a clade credibility value of 0.99 (Supplementary Fig. 1A; available online at http://vdi.sagepub.com/supplemental).
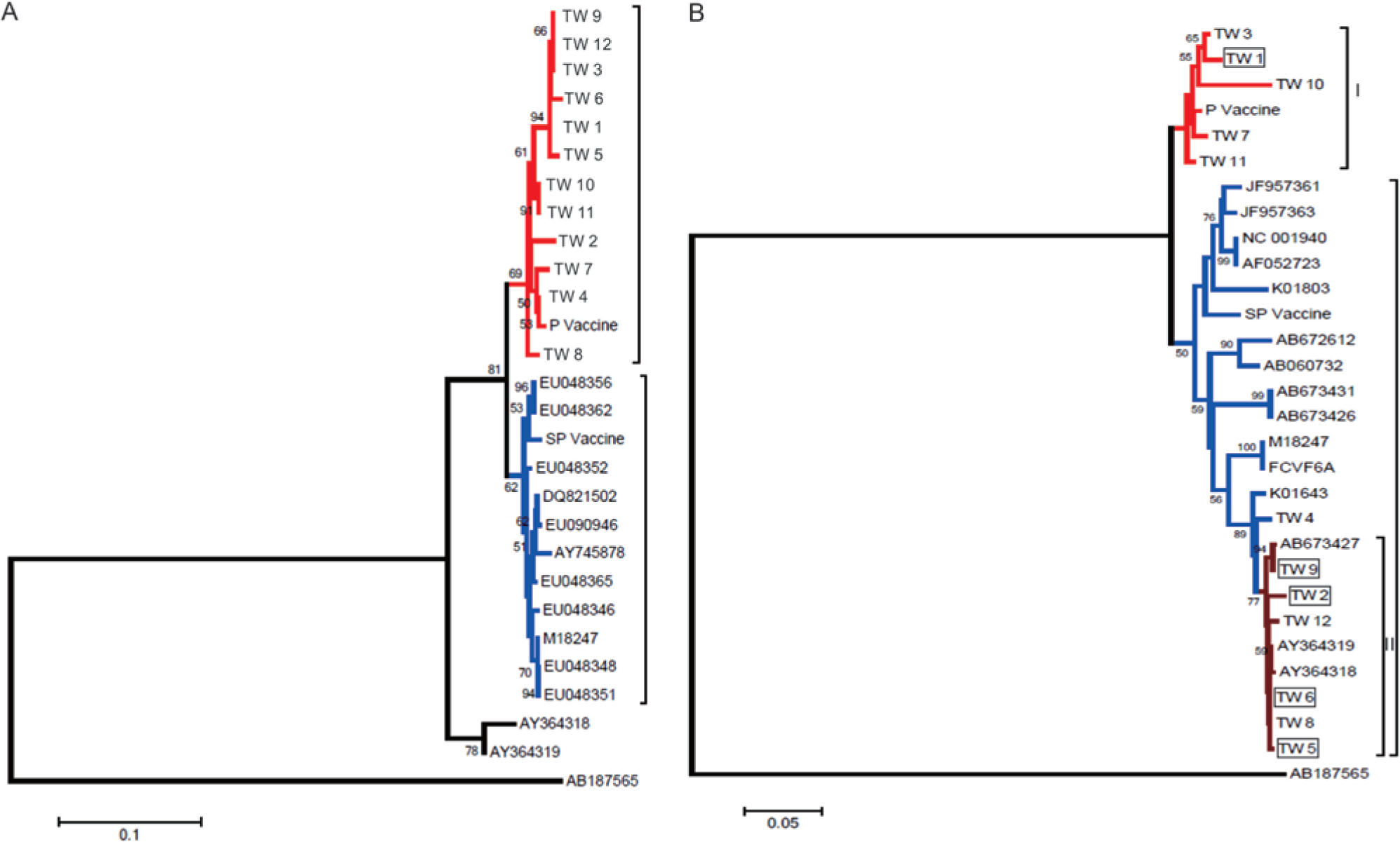

The sequence identities between the p27 genes from different TW strains ranged from 88.7% to 99.3%, and the identity between the p27 gene from the P vaccine strain and that of the SP vaccine strain was 95.2% (data not shown). A wider range of identities (88.7–99.3%) was found between the p27 genes of the TW strains and the vaccine strains (data not shown). Compared with the SP vaccine strain, the sequence identities between the p27 gene of the P vaccine and that of the FeLV Taiwan strains were higher. With respect to the p27 gene, the TW strains were clustered in 2 groups based on the neighbor-joining method. The P vaccine strain was more closely related with the TW7 strain, and these were clustered with the TW1, TW3, TW10, and TW11 strains. The SP vaccine strain formed an independent cluster with the type A FeLV strains from other countries. Even though some TW strains were placed in the clade that included both endogenous FeLVs (GenBank accession nos. EU048356 and EU048362), the TW4 strain was further separated from the remaining TW strains with a bootstrap value of 94 (Fig. 5B). The TW1, TW3, TW7, TW10, and TW11 strains and the P vaccine strain were separated from the other TW strains as monophyletic groups with a clade credibility value of 0.99, and both the TW7 strain and the P vaccine strain belonged to a subclade, as determined through Bayesian analysis. The TW4 strain was paraphyletic with other TW strains and both endogenous FeLVs but separate with a clade credibility value of 0.99. However, the endogenous FeLVs were further grouped as a subclade with a clade credibility value of 0.99 (Supplementary Fig. 1B; available online at http://vdi.sagepub.com/supplemental). No significant difference (
Discussion
The positive rate of the current study, which was based on 55 sick cats collected from 2010 to 2013 and which used proviral PCR, was 22%. This ratio is close to the lower value of the average prevalence of 30–40% previously reported. 17 The positive rate of FeLV is largely determined by geographical location, lifestyle, and density of domestic and wild cats. 10 In the island of Taiwan, a strict quarantine of imported cats from overseas, a higher rate of vaccination, and improved education campaigns may all lead to a reduced prevalence of the disease. 10 In addition, it is not possible to exclude a bias in sampling because the samples tested were obtained from cats with suspected symptoms submitted by veterinarians that were usually sick due to secondary and opportunistic infections. 14
To identify FeLV-infected cats, proviral PCR or RT-PCR is used depending on the objective. The PCR detection of the exogenous proviral DNA targets the FeLV sequences archived inside the cell genome. 33 These sequences may represent active viruses or silenced circulating viruses. The detection of proviral DNA facilitates an identification of asymptotic cats and also serves as the standard method for the sequence analysis of FeLV isolates to obtain an understanding of their phylogenetic relationship. 5
In the present study, a RT-PCR was developed for the detection of the actively replicating virus, which may originate from viremia or the potential transcriptional reactivation of the integrated proviral DNA. The sensitivities of the RT-PCR and qRT-PCR assays were 100 and 1 viral copies per 0.5 ml of sample, respectively. It is noteworthy that an advantage of ARMS RT-PCR is its nested amplification, which can rapidly increase the template amounts for allelic-specific PCR through a simultaneous amplification of the target DNA by the outer primers. 21 In addition, oligo-dT primers that specifically target the poly-A tail of FeLV are used to enhance the efficiency of the reverse transcriptions. The developed RT-PCR assay can also be used to detect FeLV in primary viremia at the early phase of infection before its integration into the genomic DNA and, most important, a local infection characterized by FeLV that only exists in local tissues instead of the bone marrow under strong immune pressure. Moreover, a previous study also indicated that, as long as a small amount of FeLV is integrated into the cells and then actively transcripted, the qRT-PCR assay can detect FeLV RNA even without the presence of an antigen, whereas ELISA would provide negative results. 31 In the current study, 12 cats were tested positive by proviral PCR, ARMS RT-PCR, ARMS qRT-PCR, and p27 immunochromatography assay. Based on the progress of FeLV, these cats were likely exposed to FeLV more than 8 weeks prior to the study, which implies that the cats exhibit viremia, the presence of circulating FeLV, and proviral DNA in peripheral white blood cells. 29 It is noteworthy that, of the 12 cats, 4 cats (TW2, TW5, TW6, and TW9) were dead with clinical symptoms of FeLV-associated diseases.
Viral replication cannot be totally inhibited by contemporary vaccines, but these vaccines are able to protect cats from the development of FeLV-associated diseases. 15 When vaccinated cats are exposed to FeLV, transient viremia may still occur (Sparkes AH: 2003, Feline leukemia virus and vaccination. Proceedings of the ESFE Feline Congress, Stockholm, September 2002. J Feline Med Surg 5:97–100). The most common vaccines are based on inactivated virus and are manufactured by different vendors with various adjuvants. Regardless of the content of the FeLV vaccines, type A FeLV is likely the background virus. Even though there is no available information on the spread of type B and type C FeLVs via a horizontal route, the productive infection of type B or type C FeLVs always occurs after recombination with type A FeLVs. 3 However, a different subgroup of FeLV may still be used. This fact may explain why a closer relationship is only observed between the SP vaccine and type A FeLVs (GenBank accession no. M18247) in the LTR phylogeny. All of the FeLV TW strains and the FeLV P vaccine strain belong to a separate clade in the LTR phylogeny. Unlike vaccines for other viruses, safety control for assurance of an effective inactivation of retroviruses, such as FeLV, is a significant issue. To date, no differentiation of the FeLV vaccine and wild-type strains can be made using serology-based methods; however, the method developed in the current study, which is based on a genetic test, appears to be suitable for differentiation of vaccine and wild-type infections.
Even though all of the purine and pyrimidine mismatches on the 3′-terminal nucleotides of the ARMS RT-PCR primers are more refractory to extension, the primer/template S/T mismatches in the present study are less refractory to extension; thus, an additional mismatch was included 2 bases from the 3′-termini.
25
For destabilization of the ARMS RT-PCR primers and improvement of the specificity of the assay, the annealing temperature was increased to 58°C. In terms of sensitivity, specificity, and diagnostic capability, this assay is acceptable. To obtain a rapid process, a simple procedure, and a more accurate analysis with a quantitative standard, the ARMS qRT-PCR was developed. For quantitative purposes, different amounts of FeLV RNA, produced by in vitro transcription of FeLV plasmids, were amplified with samples in parallel. This single-step and closed-tube system is especially useful for SNP analysis. The EvaGreen dye, instead of the SYBR green dye, was used for HRM because the fluorescence reduction of the former in the beginning of the melting analysis is prone to be quenched.
32
The discrimination of Tm based on the G + C content (%) can only be achieved between the vaccine strains and the TW strains but not between the SP vaccine strain (86.77°C) and the P vaccine strain (87°C). Furthermore, a model-free technique was applied to differentiate between different strains based on the shape of the melting curve. By including controls of all of the strains in each analyzed batch, the reproducibility of the results was confirmed. Based on the confidence percentages, the TW strains were successfully distinguished from the vaccine strains (at least 91.17% vs. 26.9%). This finding suggests an acceptable reliability as a differential diagnosis for the TW and vaccine strains. This type of ARMS qPCR assay has been used for the detection of SNPs of the
Based on their LTR gene sequences, all of the FeLV TW strains and other exogenous FeLVs are clustered as a single clade and are monophyletic with endogenous FeLV. The LTR gene was selected as the target for phylogenetic analysis because the sequence variation in the LTR gene largely determines the tumorigenicity and outcome of the disease. 2 Based on the LTR phylogeny, the TW strains are clustered as an isolated group, and this finding was strongly supported by a clade credibility value of 0.95 from the Bayesian analysis. The result suggests that the TW strains exhibit a geographic characteristic. The p27 gene of FeLV not only encodes a major core protein but is a target for the detection of viremia. This gene may have a higher variation than the LTR gene under the pressure of an immune response. Two major clades were found for the TW strains. The strains in the upper clade including the P vaccine strain are paraphyletic. The strains are also paraphyletic with the p27 gene of endogenous FeLVs (GenBank accession nos. AY364318 and AY364319). Interestingly, 4 cats (TW2, TW5, TW6, and TW9) associated with clade II died, while only 1 cat (TW1) associated with clade I died prior to the end of the study. This possibly suggests a difference in pathogenicity between the clades; however, the number of positive samples is statistically insufficient for confident conclusions. In conclusion, the phylogenetic relationship of the FeLV Taiwan strains with other strains was reported. The developed ARMS RT-PCR assays were determined to be useful for the differentiation of the FeLV Taiwan strains and the FeLV vaccine strains.
Footnotes
Acknowledgements
The authors thank Dr. ML Wong for the guidance provided during this study.
a.
Snap combo feline leukaemia virus antigen/feline immunodeficiency virus antibody test kit, IDEXX Laboratories Inc., Westbrook, ME.
b.
EPICENTRE Biotechnologies, Madison, WI.
c.
MD Co., Taipei, Taiwan.
d.
Life Technologies, Grand Island, NY.
e.
Protech, Taipei, Taiwan.
f.
pGEM-T vector, Promega, Madison, WI.
g.
Promega Corp., Madison, WI.
h.
Riboprobe
i.
Promega Corp., Madison, WI.
j.
Type-it HRM PCR kit, Qiagen GmbH, Hilden, Germany.
k.
Rotor-gene Q series software, Qiagen GmbH, Hilden, Germany.
l.
DNASTAR Inc., Madison, WI.
Declaration of conflicting interests
The author(s) declare no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
