Abstract
Real-time reverse transcription polymerase chain reaction (real-time RT-PCR) is routinely used for the rapid detection of Avian influenza virus (AIV) in clinical samples, but inhibitory substances present in some clinical specimens can reduce or block PCR amplification. Most commercial RNA extraction kits have limited capacity to remove inhibitors from clinical samples, but using a modified commercial protocol (Ambion® MagMAX™, Applied Biosystems, Foster City, CA) with an added high-salt wash of 2 M NaCl and 2 mM ethylenediamine tetra-acetic acid was shown to improve the ability of the kit to remove inhibitors from cloacal swabs and some tissues. Real-time RT-PCR was carried out in the presence of an internal positive control to detect inhibitors present in the purified RNA. Cloacal swabs from wild birds were analyzed by real-time RT-PCR comparing RNA extracted with the standard (MagMAX-S) and modified (MagMAX-M) protocols. Using the standard protocol on 2,668 samples, 18.4% of the samples had evidence of inhibitor(s) in the samples, but the modified protocol removed inhibitors from all but 21 (4.8%) of the problem samples. The modified protocol was also tested for RNA extraction from tissues using a TRIzol-MagMAX-M hybrid protocol. Tissues from chickens and ducks experimentally infected with high-pathogenicity Asian H5N1 AIV were analyzed by real-time RT-PCR, and the limit of detection of the virus was improved by 0.5–3.0 threshold cycle units with the RNA extracted by the MagMAX-M protocol. The MagMAX-M protocol reported in the present study can be useful in extracting high-quality RNA for accurate detection of AIV from cloacal swabs and tissues by real-time RT-PCR.
Keywords
Introduction
Avian influenza virus (AIV) causes a viral infection in birds that may cause unapparent to severe disease and death depending on the host and virus strain. There are 16 known subtypes, H1-H16, based on the antigenic diversity of the surface glycoprotein hemagglutinin (HA). 16,30 Free-living wild aquatic birds, including waterfowl and shorebirds, are the natural reservoirs of all AIVs in which the viruses cause primarily asymptomatic infections. 31 Upon transmission of virus into gallinaceous birds, such as chickens or turkeys, most AIVs maintain low pathogenicity (LP), but occasionally, they become highly pathogenic (HP) because of mutations, recombination, or genetic exchange (genetic reassortment) between coinfecting AIV subtypes. 12,21 Highly pathogenic AIVs belong to the subtype H5 or H7, but not all H5 and H7 subtypes are highly pathogenic. 6 The HPAIV infections in domestic poultry are extremely contagious, and infected birds often die within a few days. Wild migratory aquatic birds carrying AIVs present an ongoing source of infection to domestic bird populations of North America. Until recently, wild birds were not thought to be a reservoir for HPAIV, but the emergence of Asian H5N1 HPAIV into the wild bird population has demonstrated that wild birds can be a source of virus for poultry. To date, the H5N1 lineage of virus has only been found in Asia, Africa, and Europe, but the potential to spread to North America by migrating wild birds has received consideration.
Low-pathogenic AIV infections are mostly mucosal with the viruses replicating within the respiratory or digestive tracts of the infected birds. 30 Therefore, LPAIV infections can be monitored by analyzing oral (oropharyngeal or tracheal) or fecal (cloacal) samples of infected birds. Highly pathogenic AIV infections, on the other hand, can be both mucosal and systemic, and the infections often spread to multiple organs, tissues, and skeletal muscle. 20,25 Therefore, in addition to oral and cloacal swabs, tissues are often targeted to monitor HPAIV infection in infected birds. Real-time reverse transcription polymerase chain reaction (real-time RT-PCR) is a highly sensitive molecular diagnostic test that can detect AIV infection in clinical and preclinical infected birds. 14 However, real-time RT-PCR diagnostics have been challenged by naturally occurring inhibitory substances present in some clinical samples, which may be coextracted and copurified with the RNA during extractions. 13 There have been large numbers of naturally occurring PCR inhibitors found in a variety of clinical and environmental samples, including feces, blood, soil, tissues, and urine. 19,26,34 Proper processing of the samples to remove inhibitors from these samples during RNA extraction is an important step to ensure high-quality RNA for an accurate diagnosis by real-time RT-PCR. Commercial RNA extraction kits may fail to completely remove RT-PCR inhibitors from the samples. 13
In the current study, the high throughput Ambion® MagMAX™-96 AI/ND Viral RNA Isolation Kit a (MagMAX, hereafter) was tested for RNA extraction from fecal and tissue samples of AIV-infected chickens and wild birds. The standard MagMAX (MagMAX-S) protocol, although previously shown to perform well for RNA extraction from some types of samples, failed to completely remove inhibitors from these samples. The protocol was modified (MagMAX-M) by changing its lysis buffer composition and adding a high-salt-ethylenediamine tetraacetic acid (EDTA) washing step to enhance its capability to remove inhibitors from these samples. The results show that the RNA extracted by the MagMAX-M protocol from the fecal or tissue samples spiked or infected with AIV was detectable by real-time RT-PCR at a higher sensitivity than the RNA extracted by the MagMAX-S protocol.
Material and methods
All AIVs used in the current study were grown and propagated in 9–11-day-old embryonating chicken eggs by incubating at 37°C. Amnioallantoic fluid (AAF) was harvested 30–48 hr postincubation (PI) from eggs inoculated with AIVs (H5 and H7) and 60–72 hr PI from eggs inoculated with LPAIVs and stored in aliquots at −70°C until used. The 50% egg infectious dose (EID50) titers of AIV were determined by the Reed and Muench method by inoculating serial dilutions of the virus in 9–11-day-old embryonating chicken eggs. 27 For detection of the virus in samples by virus isolation (VI), 9–11-day-old embryonating chicken eggs were inoculated in triplicate with 0.2 ml of the soluble fraction of cloacal swabs or tissue homogenate collected from the supernatant after centrifugation of the samples and incubated at 37°C for 2–3 days. The AAFs were collected from inoculated eggs and tested for AIV by HA assays. The HA assays were carried out with 0.5% chicken red blood cells in phosphate buffered saline according to the standard protocol. 8
Bird experiments with live LPAIVs or HPAIVs were carried out in biosafety level (BSL) 2 or 3 agriculture containment facilities at the Southeast Poultry Research Laboratory (SEPRL, Athens, GA). All animal experiments were carried out according to protocols approved by the U.S. Department of Agriculture, and general care was provided as required by the Institutional Animal Care and Use Committee. In bird experiments, viruses were inoculated intranasally into 3-week-old specific-pathogen-free (SPF) White Leghorn chickens (Gallus gallus domesticus), obtained from SEPRL flocks, and into 2-week-old Pekin ducks (Anas platyrhynchos), obtained from a commercial hatchery. All inoculated birds were housed in self-contained isolation units b that were ventilated with high-efficiency particulate-filtered air.
Cloacal swabs used in the current study were obtained from wild bird samples that were collected as part of another project between 2004 and 2006 using a standard protocol as outlined on the official Web site: http://alaska.usgs.gov/science/biology/avian_influenza/monitoring.html. All cloacal swabs were placed into 2 ml of brain heart infusion (BHI) medium in screw-capped collection tubes, and the reconstituted swabs were kept refrigerated before shipment to SEPRL. Upon arrival at SEPRL, the swabs were kept at −70°C until processed for RNA extraction or VI. To extract RNA, cloacal swabs were vortexed vigorously for 15 sec to release the virus from suspended fecal materials and then centrifuged at 4°C at 5,000 × g for 10 min to pellet large particulate matter, and the supernatant was collected for RNA extraction or VI. Wild bird cloacal swabs that tested negative for both AIV by real-time RT-PCR and VI were pooled and used as starting material for the first phase to optimize the RNA extraction protocols described in the present study.
For RNA extraction from tissues infected with HPAIVs, the samples were first inactivated with TRIzol LS c in a BSL-3 facility and then brought to a level 2 biosafety laboratory for RNA extraction. All tissues were sampled and processed according to protocol previously described. 14 Briefly, tissues with approximate weights between 0.5 and 1.0 g were collected in 2-ml collection tubes containing 600 mg of zirconia beads c (0.5 mm in diameter), 0.25 ml of BHI, and 0.75 ml of TRIzol LS and macerated with a commercial tissue disrupter. d Next, 200 μl of chloroform was added to homogenized tissues, and the contents vortexed thoroughly to release RNA from the tissues. The chloroform-TRIzol extracts were centrifuged at 5,000 × g for 10 min, and the upper aqueous phase containing the crude RNA extracts were collected for isolation of RNA as described below.
The following commercial RNA extraction kits were used and compared in the current study: the RNeasy Mini Kit (QR), e the QIAamp Viral RNA Mini Kit (QV), e the EZ1 Virus Mini Kit v2.0 e (EZ1), the TRIzol LS Kit (Trzl), c and the MagMAX AI/ND Viral RNA Isolation Kit (MagMAX). a All RNA extraction methods were performed according to the manufacturer's instructions, except for the QR kit, where an approved standard operating protocol modified from the National Veterinary Services Laboratories was used. 13 The RNA extraction using the MagMAX kit was performed on a semiautomated nucleic acid purification system f according to the manufacturer's instructions. For initial studies, cloacal swabs from wild birds that tested negative for AIV by real-time RT-PCR and VI were randomly selected, pooled, spiked with LPAIV A/chicken/AL/7395/75 (H4N8), and used as a starting material for optimization of RNA extraction protocols. Two batches of pooled cloacal swabs were used, each consisting of 50 different swabs with 500 μl of swab material sampled from each. Fifty microliters of pooled swab material was used directly for RNA extraction with MagMAX, whereas different amounts of BHI medium was added to 50 μl of swab material to increase the volume to 500, 140, 250, and 400 μl for RNA extraction with QR, QV, Trzl, and EZ1, respectively. For tissue samples, the aqueous TRIzol extracts obtained after maceration of tissues in TRIzol LS plus chloroform, as described above, were used as starting material for extraction of RNA using different commercial kits.
To further improve the capacity of the MagMAX protocol to remove inhibitors from cloacal swabs, 2 modifications were added to the MagMAX-S protocol: the lysis-binding buffer was supplemented with bovine serum albumin (BSA; 15 μg per extraction) and sodium sulfite (870 μg per extraction), and an extra washing step with a low-pH, high-salt solution consisting of 2 M NaCl and 2 mM EDTA (pH 4.0) was added between wash solutions 1 and 2 of the MagMAX-S protocol. The above changes were tested separately at different times to determine their influence on RNA extraction, and all 3 changes were found to improve RNA extraction from cloacal swabs compared with the MagMAX-S protocol (results not shown). The modified MagMAX protocol, now referred to as MagMAX-M, was used in all RNA extractions reported in the current study, except for the tissues, where the RNA extraction with MagMAX-M protocol was performed without any change in the lysis buffer composition.
The detection of AIV by real-time RT-PCR was carried out on a SmartCycler real-time PCR machine g via amplification of the AIV matrix (M) gene using a lyophilized master reaction mix with a 1-step RT-PCR mix e as described previously. 13 The lyophilized master mix also contained an internal positive control (IPC) to monitor and detect RT-PCR inhibitors in the sample. 13 During the assay, the amplification of the M gene was monitored in the FAM (fluorophore 6-carboxyfluorescein) channel with an M-specific probe, and the amplification of the IPC was monitored in the Texas Red channel with an IPC-specific probe as described earlier. 13 During the amplification reaction, any fluorescence over threshold in the FAM channel would indicate the presence of AIV in the sample, whereas fluorescence in the Texas Red channel would indicate no inhibitors were present in the sample. The absence of any fluorescence in either channel would indicate a failed reaction likely because of the presence of 1 or more inhibitors in the sample. 13 A no-template control (NTC; water) was also included in the assay, and the inclusions of the NTC and the IPC ensured the accuracy of the assay, and the assays were repeated in case of false-positive (NTC-positive) results.
Results
Earlier studies show that the wild bird fecal specimens are some of the most difficult samples for RNA extraction because of their extremely high content of RT-PCR inhibitors. 13 In the current study, 5 commonly used commercial RNA extraction kits (Trzl, QR, QV, EZ1, and MagMAX) were compared for RNA extraction from wild bird cloacal swabs. Pooled cloacal swabs from wild birds and BHI medium (reference control) were spiked with LPAIV A/chicken/AL/7395/75 (H4N8) and used as the starting materials. The extracted RNA was analyzed by real-time RT-PCR as described above. Two batches of pooled wild bird cloacal swabs were used: pooled swab 1 and pooled swab 2. Initial evaluations of the RNA kits were based on their ability to extract RNA from serial dilutions of the virus in pooled swab 1 and BHI medium as the starting material. The results (Table 1, BHI data shown within parentheses) show that for the virus in BHI, the Trzl and MagMAX-S methods had the best performance, showing the lowest threshold cycle (Ct) values. For the virus in pooled cloacal swab material, the MagMAX-S and EZ1 had the best performance, showing the lowest Ct values (Table 1). The minimum detection limit (MDL) of the virus titer (EID50/ml) detected from serial dilutions in BHI was determined to be 101.3 for the RNA extracted by all 5 commercial RNA extraction kits, whereas it was higher and widely varied for serial dilutions in pooled swab 1, depending on the RNA extraction kits being used, and the values of the MDL corresponding to the RNA kits were 107.3 for Trzl, 104.3 for QR and QV, and 102.3 for EZ1 and MagMAX. The above results indicate efficient removal of RT-PCR inhibitors from cloacal swabs by EZ1 and MagMAX compared with other RNA kits used in this study. The poor performance of Trzl to remove RT-PCR inhibitors from cloacal swabs has also been reported earlier. 13
Despite the higher sensitivity of the detection of AIV from cloacal swabs by EZ1 and MagMAX, neither protocol was able to completely remove RT-PCR inhibitors from cloacal swabs because the Ct values were 4–6 units higher with the RNA extracted from cloacal swabs than the RNA extracted from BHI medium, although the same amount of the virus (A/chicken/AL/7395/75) was present in both samples. Between the 2 kits, the RNA extracted by the MagMAX protocol exhibited slightly lower Ct values than the RNA extracted by EZ1 (Table 1). Because the MagMAX protocol had logistical advantages over EZ1, including higher throughput extractions (24 extractions compared to only 6 by EZ1), shorter time of extraction (15 min compared with 35 min with EZ1), lower costs of operation (higher costs of reagents and equipment for EZ1 than MagMAX), and a flexible format that allowed changes in the protocol, the MagMAX protocol was further investigated to improve its ability to remove inhibitors from cloacal swabs. Two modifications of the MagMAX protocol were tested: BSA and sodium sulfite were added to the lysis-binding solution, and an extra washing step, with low-pH, high-salt-EDTA buffer (2 M NaCl and 2 mM EDTA, pH 4.0), was added between the 2 standard washing steps (wash solution 1 and wash solution 2) of the original protocols. To test the effect of these changes on the performance of the MagMAX protocol, pooled swab 2 material spiked with serial dilutions of the LPAIV A/chicken/AL/7395/75 (H4N8) were used as the starting material. The results of real-time RT-PCR (Table 1) show marginal differences in the sensitivity of detection of the AIV RNA from serial dilutions in BHI by both MagMAX-S and MagMAX-M protocols, but the 2 protocols exhibited significant differences in the limit of detection of the virus from serial dilutions in the 2 batches of cloacal swabs used in this study. For pooled swab 1, the MDL of the virus titer was determined to be 102.3 with the RNA extracted by both MagMAX-S and MagMAX-M protocols (data not shown), whereas, for pooled swab 2, the MDL was determined to be 102.3 with the RNA extracted by the MagMAX-M protocol and 106.3 with the RNA extracted by the MagMAX-S protocol. These results suggest that there were higher amounts of RT-PCR inhibitors present in pooled swab 2 than in pooled swab 1 and that the inhibitors were more efficiently removed by the MagMAX-M protocol than the MagMAX-S protocol. The 2 pooled swabs were found to be indistinguishable with respect to their physical appearance, except that the pooled swab 2 was relatively darker (reddish brown) than the pooled swab 1. Therefore, the major differences between the 2 swabs are likely to be their chemical composition, including the content of the RT-PCR inhibitors. To further verify how well the MagMAX-M protocol performed on the extraction of RNA from other AIV subtypes, serial dilutions of AIV subtypes H2, H5, and H7 from pooled swab 2 were used as the starting materials. With the RNA extracted by the MagMAX-S protocol, the MDLs of the virus titers were determined to be 103, 103.1, and 103 for the AIV subtypes H2, H5 and H7, respectively, whereas they were 102, 101.1, and 101, respectively (data not shown), with the RNA extracted by the MagMAX-M protocol. These results indicate consistency and wide applicability of the MagMAX-M protocol for extraction of RNA from major subtypes of AIV from cloacal swabs.
Comparison of commercial or modified commercial RNA kits for extraction of Avian influenza virus RNA from pooled cloacal swab samples or brain heart infusion (BHI).*
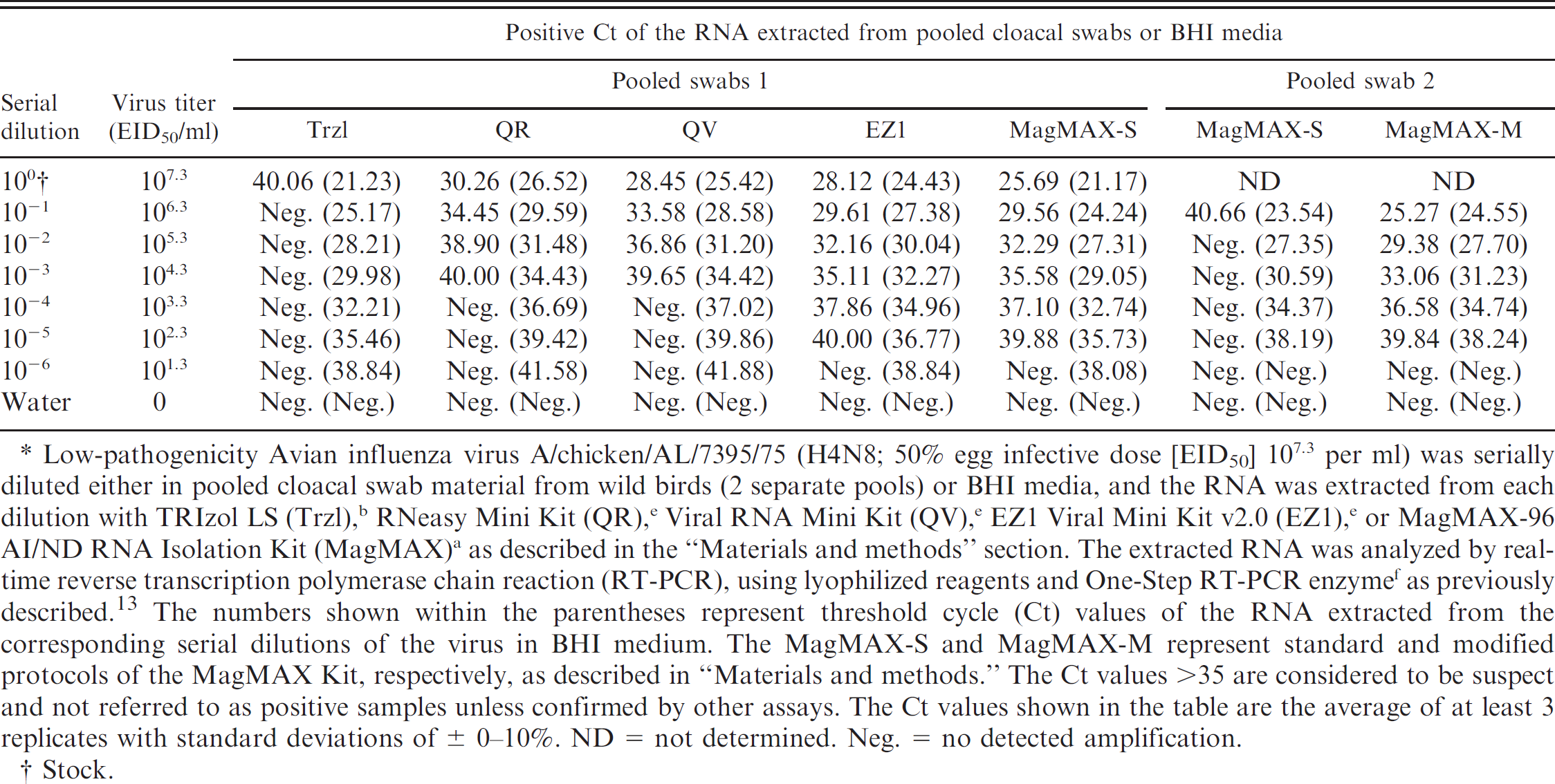
Low-pathogenicity Avian influenza virus A/chicken/AL/7395/75 (H4N8; 50% egg infective dose [EID50] 107.3 per ml) was serially diluted either in pooled cloacal swab material from wild birds (2 separate pools) or BHI media, and the RNA was extracted from each dilution with TRIzol LS (Trzl), b RNeasy Mini Kit (QR), e Viral RNA Mini Kit (QV), e EZ1 Viral Mini Kit v2.0 (EZ1), e or MagMAX-96 AI/ND RNA Isolation Kit (MagMAX) a as described in the “Materials and methods” section. The extracted RNA was analyzed by real-time reverse transcription polymerase chain reaction (RT-PCR), using lyophilized reagents and One-Step RT-PCR enzyme f as previously described. 13 The numbers shown within the parentheses represent threshold cycle (Ct) values of the RNA extracted from the corresponding serial dilutions of the virus in BHI medium. The MagMAX-S and MagMAX-M represent standard and modified protocols of the MagMAX Kit, respectively, as described in “Materials and methods.” The Ct values >35 are considered to be suspect and not referred to as positive samples unless confirmed by other assays. The Ct values shown in the table are the average of at least 3 replicates with standard deviations of ± 0–10%. ND = not determined. Neg. = no detected amplification.
Stock.
As the MagMAX RNA extraction protocols performed better than the other commercial RNA extraction protocols described in the present study, the MagMAX protocol was used in a surveillance study to compare RNA extraction methods from wild bird cloacal swabs. At first, all RNA extractions were performed with the MagMAX-S protocol, and the extracted RNA were analyzed by real-time RT-PCR as described above. Out of 2,668 cloacal swabs used in the current study for RNA extraction with the MagMAX-S protocol, 403 samples tested positive for AIV (15.1%), and 1,774 samples tested negative for AIV (66.5%; Table 2). However, 491 samples tested negative for both AIV and IPC (18.4%; potential false negatives), indicating the presence of inhibitors in these samples. Out of the 491 potentially false-negative samples, 433 samples (88.2%) were re-extracted with the MagMAX-M protocol, and the extracted RNA was analyzed by real-time RT-PCR. The results showed 46 out of 433 samples tested positive for AIV (10.6%), 366 tested negative for AIV (84.5%, positive for IPC), and only 21 tested negative for both AIV and IPC (4.9%), indicating efficient removal of RT-PCR inhibitors from cloacal swabs by the MagMAX-M protocol.
Real-time reverse transcription polymerase chain reaction of wild bird cloacal swabs with the RNA extracted by standard and modified MagMAX viral RNA isolation protocols.*
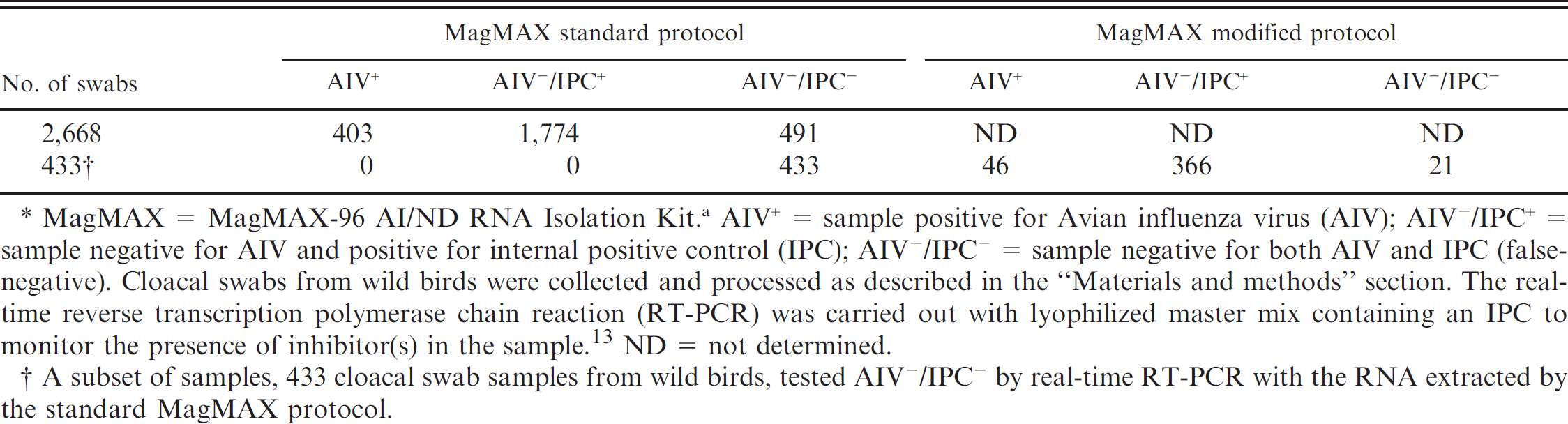
MagMAX = MagMAX-96 AI/ND RNA Isolation Kit. a AIV+ = sample positive for Avian influenza virus (AIV); AIV−/IPC+ = sample negative for AIV and positive for internal positive control (IPC); AIV−/IPC− = sample negative for both AIV and IPC (false-negative). Cloacal swabs from wild birds were collected and processed as described in the “Materials and methods” section. The real-time reverse transcription polymerase chain reaction (RT-PCR) was carried out with lyophilized master mix containing an IPC to monitor the presence of inhibitor(s) in the sample. 13 ND = not determined.
A subset of samples, 433 cloacal swab samples from wild birds, tested AIV−/IPC− by real-time RT-PCR with the RNA extracted by the standard MagMAX protocol.
To extend its applicability to other types of sample matrices, MagMAX protocols were tested for RNA extraction from tissues infected with AIV. The MagMAX kits are not designed for direct extraction of RNA from tissues. However, the authors recently used a hybrid protocol consisting of TRIzol LS and MagMAX-96 AI/ND viral isolation kits for RNA extraction from tissues, which helped to detect AIV in chickens infected with HPAIV H5N1 during the preclinical and clinical stage of infection. 14 Although this hybrid protocol was found to be effective in detecting AIV from infected tissues, some evidence of PCR inhibition was observed with the RNA extracted from these tissues. Because an additional washing step with low-pH, high-salt-EDTA buffer was shown to be effective in the removal of RT-PCR inhibitors from cloacal swabs, this step was added to the TRIzol-MagMAX hybrid protocol for RNA extraction from tissues. Multiple tissue types from chickens and ducks infected with HP H5 and H7 AIV subtypes were extracted with this modified TRIzol-MagMAX hybrid protocol, and the extracted RNA was analyzed by real-time RT-PCR. The results (Table 3) show improved RNA extraction from tissues with the modified TRIzol-MagMAX hybrid protocol over the standard TRIzol-MagMAX hybrid protocol. The average Ct values corresponding to the RNA extracted by the modified TRIzol-MagMAX hybrid protocol were found to be marginally improved in breast (-0.46), brain (-0.03), lung (-0.69), and kidney (-0.53) and significantly improved in thigh (-2.33) and heart (-3.71) from infected chickens compared with the RNA extracted by the standard TRIzol-MagMAX hybrid protocol. Improved sensitivity in the detection of the virus was also observed in brain (-1.03), lung (-0.83), spleen (-0.73), heart (-1.54), and muscle (-1.09) from infected ducks with the RNA extracted by the modified TRIzol-MagMAX hybrid protocol compared with that extracted by the standard TRIzol-MagMAX hybrid protocol.
Discussion
The sensitivity and accuracy of the real-time RT-PCR diagnostics can be compromised by several factors, including RNA extraction, primer and probe design, RT-PCR enzymes, thermocycling conditions, and PCR inhibitors. Each of the above parameters can potentially influence the amplification of the target amplicon by PCR resulting in different levels of sensitivity in tests. False-negative results caused by the inhibition of PCR amplification by naturally occurring inhibitory substances present in the sample are a common problem in molecular diagnostics of viral diseases by PCR-based methods when testing certain types of samples. In the current study, different commercial RNA extraction kits were tested and analyzed for their ability to remove inhibitors from cloacal swabs and tissues. Cloacal swabs and tissues are more difficult samples from which to extract RNA because of their complex chemical composition and high content of organic materials, which are difficult to remove by commonly used commercial RNA extraction kits. Three different chemistries for RNA extraction were tested in this study, including organic solvent extraction (Trzl), column-based silica extraction (QR and QV), and magnetic bead-based extraction (MagMAX and EZ1). With the same amount of virus used to spike the BHI controls and cloacal swabs, the Ct values corresponding to the viral RNA extracted by Trzl from the BHI controls were much lower than the Ct values of the viral RNA extracted from the cloacal swabs, indicating the presence of RT-PCR inhibitors in the cloacal swabs that were copurified with the RNA, resulting in inhibition and in false-negative results. On the other hand, the automated magnetic bead protocols (MagMAX and EZ1) were shown to perform well and yield relatively cleaner RNA (less inhibition), which minimized the occurrence of false-negative results. However, both protocols (MagMAX and EZ1) were far from perfect because they failed to completely remove RT-PCR inhibitors from the cloacal swab samples. Because the MagMAX protocol had a more flexible format than the EZ1 system, it was further investigated to improve its ability to remove inhibitors from the samples. The modifications that improved the capacity of the MagMAX kit to remove inhibitors from cloacal swabs and tissues included addition of BSA and sodium sulfite in the lysis-binding buffer and an additional washing step with a high-salt-EDTA buffer between the 2 standard washing steps (wash solution 1 and wash solution 2) of the original protocol. Both BSA and sodium sulfite were reported to reduce the inhibitory effects of naturally occurring PCR inhibitors found in blood, meat, and fecal samples. 3,7,11,18,19,22,28 In addition, sodium sulfite was also reported to improve the yield and stability of the nucleic acids (both RNA and DNA). 7,28 The higher stability of nucleic acids by sodium sulfite could be linked to its role as an effective reducing agent, thereby preventing oxidation of nucleic acids as described earlier. 11,28,29 The low-pH, high-salt wash could facilitate the removal of RT-PCR inhibitors, whereas the EDTA inhibits RNase activity and removes metal ions by forming complexes with the latter. 10,24,33
Comparison between the standard and modified MagMAX viral RNA isolation protocols for extraction of RNA from various tissues of chickens and ducks.*
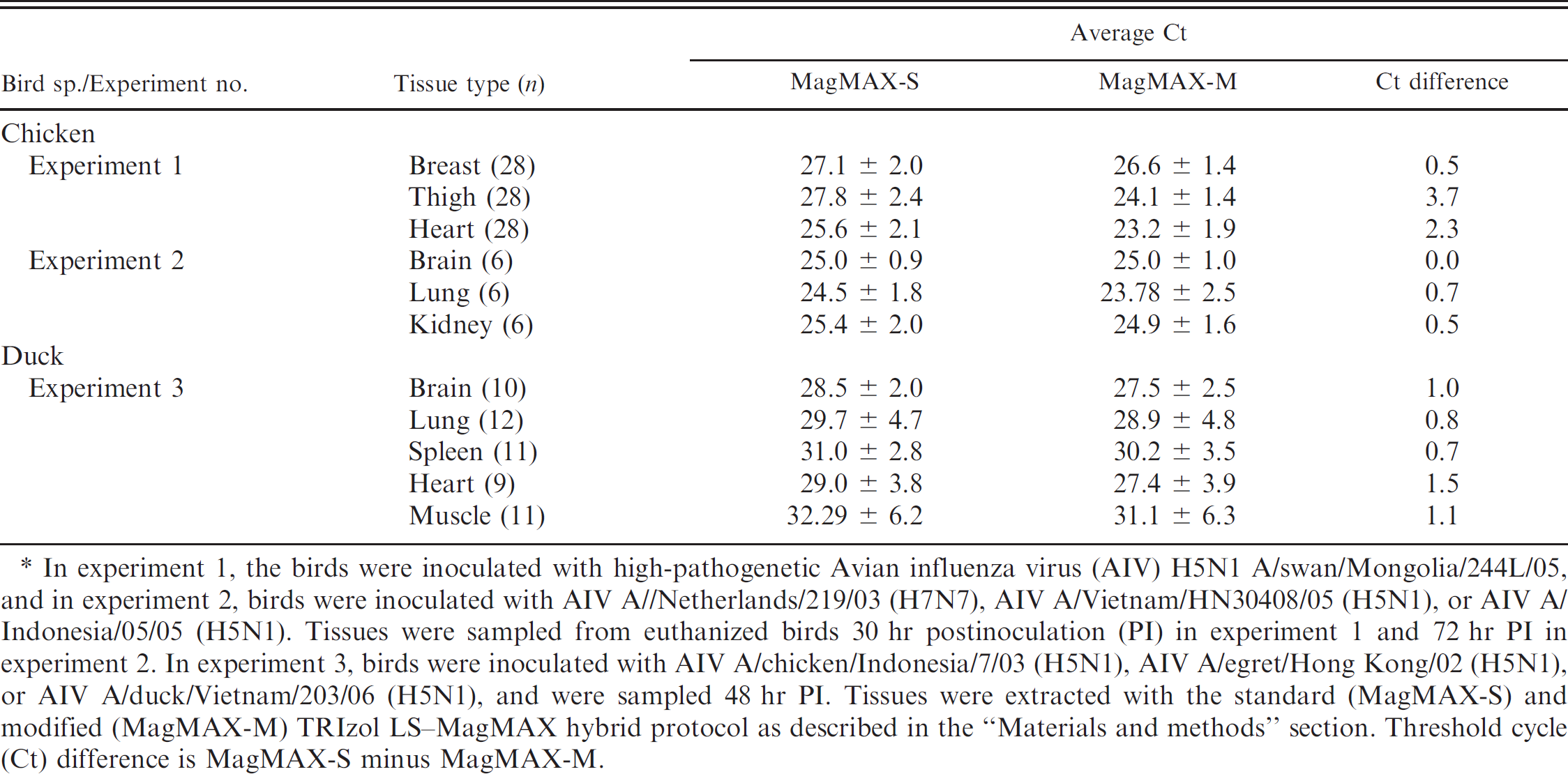
In experiment 1, the birds were inoculated with high-pathogenetic Avian influenza virus (AIV) H5N1 A/swan/Mongolia/244L/05, and in experiment 2, birds were inoculated with AIV A//Netherlands/219/03 (H7N7), AIV A/Vietnam/HN30408/05 (H5N1), or AIV A/Indonesia/05/05 (H5N1). Tissues were sampled from euthanized birds 30 hr postinoculation (PI) in experiment 1 and 72 hr PI in experiment 2. In experiment 3, birds were inoculated with AIV A/chicken/Indonesia/7/03 (H5N1), AIV A/egret/Hong Kong/02 (H5N1), or AIV A/duck/Vietnam/203/06 (H5N1), and were sampled 48 hr PI. Tissues were extracted with the standard (MagMAX-S) and modified (MagMAX-M) TRIzol LS-MagMAX hybrid protocol as described in the “Materials and methods” section. Threshold cycle (Ct) difference is MagMAX-S minus MagMAX-M.
It is unclear how the inhibitors interfere with real-time RT-PCR assays. It has been suggested that the inhibition could be due to any or all of the following reasons: inactivation of DNA polymerase, degradation or capture of nucleic acids, and poor yield of nucleic acids because of incomplete cell lysis. 26,34 Inactivation of DNA polymerase by naturally occurring inhibitors is by far the most common cause for PCR failure and false-negative results. Some of the naturally occurring components identified as PCR inhibitors include bile salts and complex polysaccharides in fecal samples, 19,23 heme and immunoglobulin in blood, 1,4,5 melanin and myoglobin in tissues, 9,15 humic acid in soil, 32 and urea in urine. 17 Because of the complexity in the composition of fecal material, it is virtually impossible to identify the naturally occurring PCR inhibitors present in the fecal samples from wild birds used in this study. Based on the results shown in Table 1 and as described in the text, the PCR inhibitors of wild bird cloacal swabs appear to be mostly water-soluble compounds, and as such, they are hard to remove by conventional RNA extraction protocols. 13 Furthermore, the level of PCR inhibitors in the wild bird cloacal swabs could vary depending on the location (geographic origin), ecosystem, dietary differences, and the habitats of the birds.
The MagMAX-M protocol and the modified TRIzol-MagMAX hybrid protocol described in the present study were shown to remove PCR inhibitors from both cloacal swabs and tissues. Although efficient removal of PCR inhibitors from the test sample is the best solution to resolving the inhibition problem, it is also important to test the efficacy of the DNA polymerases against these inhibitors because the DNA polymerases were shown to have different levels of sensitivity against the inhibitors in biological specimens. 2,3,26 Therefore, an alternative approach to reduce false-negatives by PCR inhibitors is to optimize the enzymes and conditions that best suit the sample type. The MagMAX-M protocol described in the current study provides a high-quality RNA extraction from both cloacal swabs and tissues that can be processed in a timely and cost effective manner. Although not tested in the present study, the potential exists for this application to be extended to extraction of nucleic acids from viruses other than AIV, which are present in similar matrices, and to other sample types, such as blood and urine.
Acknowledgements
The authors wish to thank Scott Lee, Suzanne DeBlois, and Diane Smith. This project was supported by Agriculture Research Current Research Information System project 6612-32000-048.
Footnotes
a.
Applied Biosystems, Foster City, CA.
b.
Invitrogen Corp., Carlsbad, CA.
c.
BioSpec Products Inc., Bartlesville, OK.
d.
FastPrep® FP120, Savant Instruments Inc., Holbrook, NY.
e.
Qiagen Inc., Valencia, CA.
f.
KingFisher® 24, Thermo Fisher Scientific Inc., Waltham, MA.
g.
SmartCycler, Cepheid Corp, Sunnyvale, CA.
