Abstract
Porcine TTV isolates Sd-TTV31 and Sd-TTV1p have a relatively high nucleotide sequence identity (69.6%) and are classified as TTV genogroup 1. Sd-TTV31, a prototype of genogroup 1, and Sd-TTV2p, a recently sequenced swine TTV, share a low nucleotide sequence identity (44%) with Sd-TTV31. Sd-TTV2p by itself composes novel TTV genogroup 2.
15
A number of researchers have reported TTV prevalence in various situations.
6,8
Coinfection with different genogroups of TTV in an individual was found to be common.
12
TTV genogroup 2 has been found more frequently in
To date, the only available method for detection of TTV is nucleic acid detection. One method available for porcine TTV, nested polymerase chain reaction (nPCR), is very sensitive but is time consuming, is labor intensive, and creates the potential for significant contamination problems. To date, no tissue culture system has been available for the propagation of the virus, in that immunological detection methods such as enzyme-linked immunosorbent assay (ELISA) and quantitative PCR (qPCR) are not yet available for TTV detection.
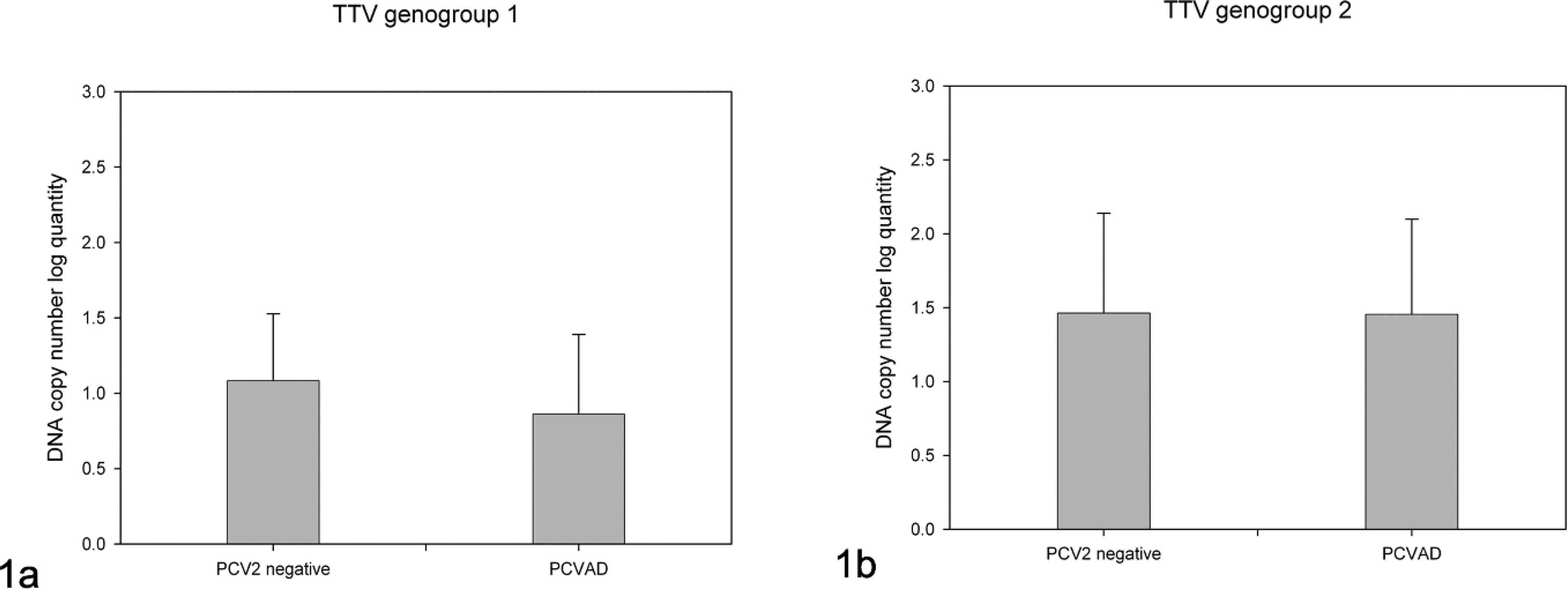
Viral load difference of each
The purpose of the current study was to develop a qPCR assay for quantification and differentiation of TTV geno-groups and investigate a possible link between TTV and PCVAD in pigs. To compare TTV viral load in various situations, blood samples were collected from 11 PCV-2–negative 150-day-old pigs as negative controls and 11 typical PCVAD-affected pigs of the same age and breed. Absence of PCV-2 in control pigs was confirmed by PCV-2–specific PCR in serum, immunohistochemistry of lymphoid tissues, and serological testing (ELISA).
DNA was extracted from serum samples with DNazol
a
according to the manufacturer's instructions and eluted with 30 μl of diethylpyrocarbonate (DEPC)-treated distilled water and stored at −70°C until use. The 3' untranslated region (UTR) of nucleotide sequences of TTV genogroup 1 representative strain TTV 1p (GenBank accession AY823990.1) and TTV genogroup 2 representative strain TTV 2p (GenBank accession AY823991) were selected as target regions of the qPCR, and primers and probes were designed with the Beacon designer.
b
TTV genogroup 1–specific probe (5'-/56FAM/CCCGACCAATCAGATTTGACTCCTCCT/3BHQ_1/-3'), sense primer (5'-CGGGCAAAATGGCGGACAG-3'), and antisense primer (5'-ACACCCTATGCAAATGAGGACTTG-3') were diluted a concentration of 10 pmol and used for qPCR. TTV genogroup 2–specific probe (5'-/5HEX/TCTACCACCAGCGGCATAAACTCAGC/3BHQ_1/-3'), sense primer (5'-AAAAGAACCAATCAGAAGAACTTCCTC-3'), and antisense primer (5'-AGGCGGTTAGACACTCAGCTC-3') were diluted and also used for qPCR. Amplicon sizes of TTV genogroups 1 and 2 were 140 and 116 bp in length, respectively. The sense primer (5'-AGCACTCTTGGTCTGGACATCG-3') and antisense primer (5'-AGCAGGCACAGAAGCAGTAGTC-3') were designed for PCV-2 SYBR Green–based qPCR on the basis of the nucleotide sequence of the open reading frame 2 (ORF2) gene retrieved from GenBank (accession AF027217). Quantitative PCR was performed with a real-time PCR detection system
c
and a commercial mix
c
containing 2x reaction buffer with each deoxyribonucleotide triphosphate (dNTP), Taq DNA poly-merase, 6 mM MgCl2, and stabilizers, as well as a SYBR Green mix
c
containing 100 mM KCl, 40 mM Tris–HCl, pH 8.4, 0.4 mM each dNTP (dATP, dCTP, dGTP, and dTTP), Taq DNA polymerase (50 U/ml), 6 mM MgCl2, SYBR Green I 20 nM fluorescein, and stabilizers for SYBR Green–based qPCR. The PCR conditions were predenaturation at 95°C for 10 min followed by 45 cycles of 95°C for 15 sec and 60°C for 40 sec. For SYBR Green–based qPCR, melting curve analysis was done to check specificity of PCR reaction. After PCR cycling, the sample temperature was dropped to 30°C for 1 min then heated from 70°C to 90°C at a linear transition rate of 0.1°C/sec. A decrease in fluorescence was detected, and melting temperature (Tm) values were analyzed by commercial analysis software.
c
The PCR products were analyzed by electrophoresis in 3% agarose gel with 30 min running on Tris acetate–ethylene-diamine tetra-acetic acid buffer. To check the detection limit of qPCR and compare the results, previously published nPCR methods for amplification of 3'UTR of each TTV genogroup were used.
9
Nested PCR for TTV and PCR for PCV-2 were carried out with a commercial PCR kit.
d
Briefly, 1 μl of DNA, 1 μl of each primer of 10 pmol concentration, and 17 μl of DEPC-treated distilled water were mixed into a premix tube containing 2.5 U of

Viral load related to clinical signs.
The duplex qPCR assay and SYBR Green–based qPCR developed in the present study detected TTV DNA in the sample efficiently. The detection limit of qPCR assay was 10 DNA copies/μl of suspension buffer (Fig. 1). The Tm value of the SYBR Green–based qPCR assay was 83.3°C (±0.2°C). The PCVAD-affected pigs had PCV-2 viremia, and as previously reported, pigs with severe clinical signs showed a higher PCV-2 viral load (Fig. 2A). 19 The result of qPCR compared with nPCR is summarized in Table 1.
In the case of TTV genogroup 1, the nPCR assay showed an approximately 63% positive ratio in the PCV-2–negative group, whereas a positive ratio of 27% was shown in the PCVAD-affected group. Quantitative PCR assay resulted in 27% positive in both groups. TTV genogroup 2 was more prevalent in both groups. Both nPCR and qPCR resulted in a 90% positive ratio. TTV genogroup 2 was more prevalent in the PCVAD-affected group and in PCV-2–negative pigs than in TTV genogroup 1. No remarkable differences in prevalence of TTV genogroups between PCVAD- and PCV-2–negative groups were determined in the present study. Dissimilarity of TTV viral load in the PCV-2–negative group compared with the PCVAD-affected group is shown in Figure 2. Viral load of TTV genogroup 1 was generally low in both groups. The mean value of DNA copies showed possible divergence between groups, but it was not significant by statistical analysis (
In the current study, prevalence of TTV genogroup 2 in serum was much higher than TTV genogroup 1 in both PCV-2–negative and PCVAD-affected pigs, and the viral load of each group was not significantly different. The TTV viral load of each genogroup was not altered by clinical severity related with manifestation of PCV-2–associated disease (Fig. 2). This result suggests that TTV genogroup 2 could readily cause viremia even in the PCV-2–negative pigs. In the case of TTV genogroup 2, there were no significant differences in prevalence between groups of pigs with or without PCV-2. Recent research on porcine TTV was mainly focused on the relationship with PCV-2, but high prevalence and a relatively less remarkable viral load divergence of TTV has raised questions that TTV coinfection could be a risk factor for PCVAD pathology. However, on the basis of the current study, there was no evidence that supported TTV as a trigger for PCVAD. Further study of TTV pathology is required to answer these questions, and diverse technologies are required for defining the role of TTV in clinics. Although results of nPCR were more sensitive compared with qPCR, the duplex qPCR assay introduced in the current study could reduce time and labor for further TTV research.
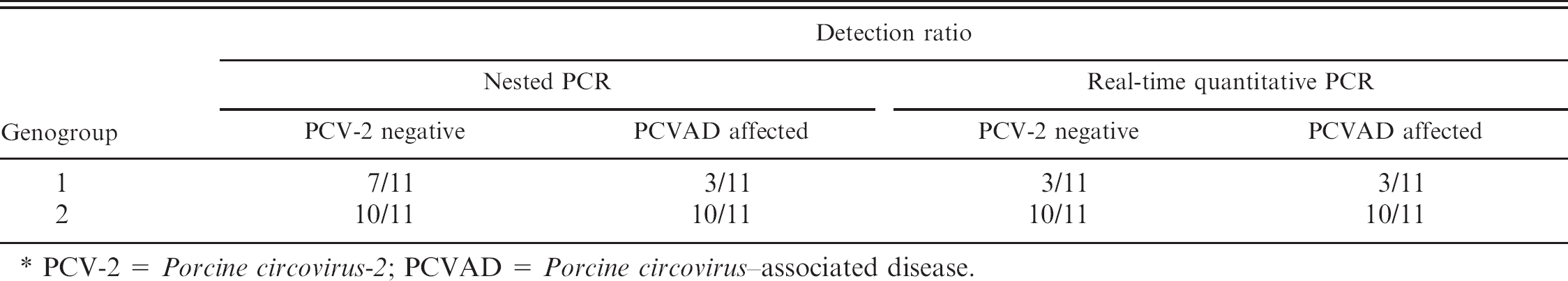

PCV-2 =
Footnotes
a.
Invitrogen Corp., Carlsbad, CA.
b.
Premier Biosoftware, Palo Alto, CA.
c.
Chromo4™ System, iQ™ Supermix, iQ™ SYBR® Green Supermix, Opticon Monitor™; Bio-Rad Laboratories, Hercules, CA.
d.
Maxime™ PCR preMix Kit, MEGAquick spin™; INtRON Biotechnology Inc., Seongnam, Korea.
e.
Promega Corp., Madison, WI.
