Abstract
The detection of an emerging pestivirus species, “HoBi-like virus,” in fetal bovine serum (FBS) labeled as U.S. origin, but packaged in Europe, raised concerns that HoBi-like virus may have entered the United States. In the current study, 90 lots of FBS originating in North America (NA) were screened for pestivirus antigen and antibodies. Lots in group 1 (G1, 72 samples) and group 2 (G2, 9 samples) originated in NA and were packaged in the United States. Group 3 (G3) was composed of 9 lots collected in NA and processed in Europe. Lots in G1 were claimed negative for Bovine viral diarrhea virus (BVDV), while lots in G2 and G3 were claimed positive by the commercial processor. All lots in G1 and G2 tested negative by reverse transcription polymerase chain reaction (RT-PCR) using HoBi-like–specific primers. Two G1 lots tested positive by BVDV RT-PCR. One of these was also positive by virus isolation. All G2 lots were positive by BVDV RT-PCR. In addition, four G2 lots were VI positive while 1 lot was antigen-capture enzyme-linked immunosorbent assay (ELISA) positive. Two G3 lots were positive by HoBi-like–specific RT-PCR tests. All lots were negative for HoBi_D32/00 neutralizing antibodies. Seven lots (4 G1; 1 G2; 2 G3) had antibodies against BVDV by virus neutralization and/or antigen-capture ELISA. While there is no evidence of HoBi-like viruses in NA based on tested samples, further studies are required to validate HoBi-like virus–free status and develop means to prevent the spread of HoBi-like virus into NA.
Introduction
Fetal bovine serum (FBS) is widely used as a growth supplement in cell culture media and as a component of many biologicals used in animal and human health. The use of FBS and its international trade pose a potential risk of introducing extraneous agents, which are contaminants of FBS, into countries. While regulation of FBS, used in the production of veterinary medicinal products, comes under the Code of Federal Regulation Title 9 (CRF9) in the United States and the European Medicines Agency (EMEA), Committee for Medicinal Products for Veterinary EMEA/CVMP/743/00 in Europe, to date there is no body of enforced international standards regarding testing of FBS for extraneous agents and no international certification of product origin.
Similar to Bovine viral diarrhea virus 1 and 2 (BVDV-1 and -2), HoBi-like viruses, an emerging putative species of pestivirus, have been found as contaminants of FBS.11,14,21,25 Reports of pestivirus contamination of animal and human vaccines have been published, and in many cases the source of contamination was the FBS.6,13,22 In addition, the presence of antibodies against bovine viruses in FBS may interfere with virus isolation and detection. 8 In 2011, detection of HoBi-like viruses in FBS lots assembled and packaged in Europe but labeled as originating in North America raised the concern that HoBi-like viruses had been introduced into the United States. 25
The current BVDV diagnostic tests may fail in detecting HoBi-like virus infection and may not differentiate from BVDV infections.1,10 A previous study evaluating reverse transcription polymerase chain reaction (RT-PCR) tests using primers based on BVDV sequences showed that these tests may fail to detect and/or differentiate HoBi-like viruses. 3 A commercial antigen-capture ELISA (ACE) widely used in the United States to screen for persistent BVDV infection was able to detect HoBi-like virus but it was not possible to differentiate between BVDV and HoBi-like virus infections. 1
The detection of HoBi-like virus in FBS lots labeled as U.S. origin opens the possibility that HoBi-like viruses are present in the United States but have not been detected because clinical signs are indistinguishable from infections with typical field BVDV strains 16 and diagnostic tests currently in use do not discriminate between BVDV and HoBi-like virus infections.1,3,10 However, contamination of FBS with HoBi-like viruses during the pooling and processing in a facility, outside the United States, that also processes sera collected from other continents cannot be discounted. The procedures adopted during FBS manufacturing such as pooling and/or processing lots from different sources using the same equipment potentially provide opportunity for cross contamination.
The efficient detection of contaminating agents in FBS is crucial to avoiding the risk of pathogens introduction into nonendemic regions. As the identification of infectious agents in regions claimed to be free may pose barriers for biological products trade. This screen for bovine pestivirus antigen and antibodies in North American FBS represents efforts to look for HoBi-like viruses in United States.
Materials and methods
Study design
The current study included samples from 90 lots of FBS. Samples in G1 and G2 were from filtered and non–gamma-irradiated lots of FBS that originated in North America (including the United States, Canada, and Mexico; personal communication with processor, 2012). Of these samples, 81 (1–81) were processed in the United States. Group 1 was composed of 72 samples (1–72) claimed negative for BVDV by the commercial processor. Group 2 was composed of 9 samples (73–81) from FBS lots that were claimed positive for BVDV. Group 3 was composed of samples from 9 filtered and gamma-irradiated FBS lots (A–I) that previously tested positive for BVDV by the processor. The FBS lots in G3 was collected in North America, but pooled and processed in Europe. Included in G3 were samples of 2 FBS lots that had previously been described as positive for HoBi-like virus. 25
Pestivirus screening included RT-PCR using 2 different sets of primers that amplify a portion of the viral 5′-untranslated region (5′-UTR). These primer sets have been widely used in the detection and genotyping of pestiviruses (HCV90-368 and 324-326 primers).15,24 Samples were also tested using HoBi-like–specific real-time RT-PCR 12 and a HoBi-like–specific RT-PCR described below. Samples were additionally tested for the presence of pestivirus antigen using a commercial ACE a and the presence of live virus by virus isolation (VI) as described below. Detection of pestivirus antibodies was carried out using a commercial antibody detection (Ab)-ELISA b and a comparative virus neutralization test (VNT). 2
Viruses and cells
Pestivirus isolates used as controls included both cytopathic (cp) and noncytopathic (ncp) BVDV strains from both the BVDV-1 (ncp BVDV1b-NY-1; cp BVDV1a- C24V; cp BVDV1a-NADL) and BVDV-2 species (ncp BVDV2a-296nc; ncp BVDV2a-1373a; cp BVDV2a-296c), noncytopathic Border disease virus (BDV) strains (BDV-BD31; BDV-WABD), a noncytopathic isolate from the putative pestivirus species Pronghorn (Pronghorn), and noncytopathic strains of HoBi-like viruses (HoBi_D32/00 and Italy-1/10-1). The BVDV isolates and HoBi-like virus were propagated and titered in bovine turbinate (BT) cells. Pronghorn virus and BDV isolates were amplified and titered in ovine fetal turbinate (OFT) cells. Cells were grown in minimal essential medium (MEM), supplemented with L-glutamine (final concentration: 1.4 mM), gentamicin (final concentration: 50 mg/l), and 10% FBS. The cells tested free for pestivirus antigen and antibodies by PCR and VNT, respectively. Cells were tested by PCR and found free of both BVDV and HoBi-like viruses. 1
RNA samples
An aliquot of 140 µl of each FBS lot was used for RNA extraction using a robotic workstation c for automated RNA purification by spin-column system d according to the manufacturer’s recommendations. The positive virus controls were diluted in FBS to a final titer of 102 tissue culture infective dose (TCID)/ml. Aliquots of FBS, MEM, and supernatant of mock infected cells (BT and OFT) were used as negative controls.
Pestivirus detection and sequencing from FBS contaminated with known concentration of HoBi-like virus and BVDV-1
The following study was performed to determine if samples contaminated with more than one pestivirus would interfere with RT-PCR testing and sequencing. Fetal bovine serum was spiked with known concentrations (TCID) of BVDV-1 (C24V) and/or HoBi_D32/00 to a final concentration as presented in Table 1. The RNA was extracted as described above and samples were submitted to RT-PCR and sequencing using primers 324-326, HCV90-368, and N2-R5, using reagents and conditions as described below.
Testing of fetal bovine serum spiked with known concentrations of Bovine viral diarrhea virus 1 (BVDV-1) and/or HoBi-like virus.*
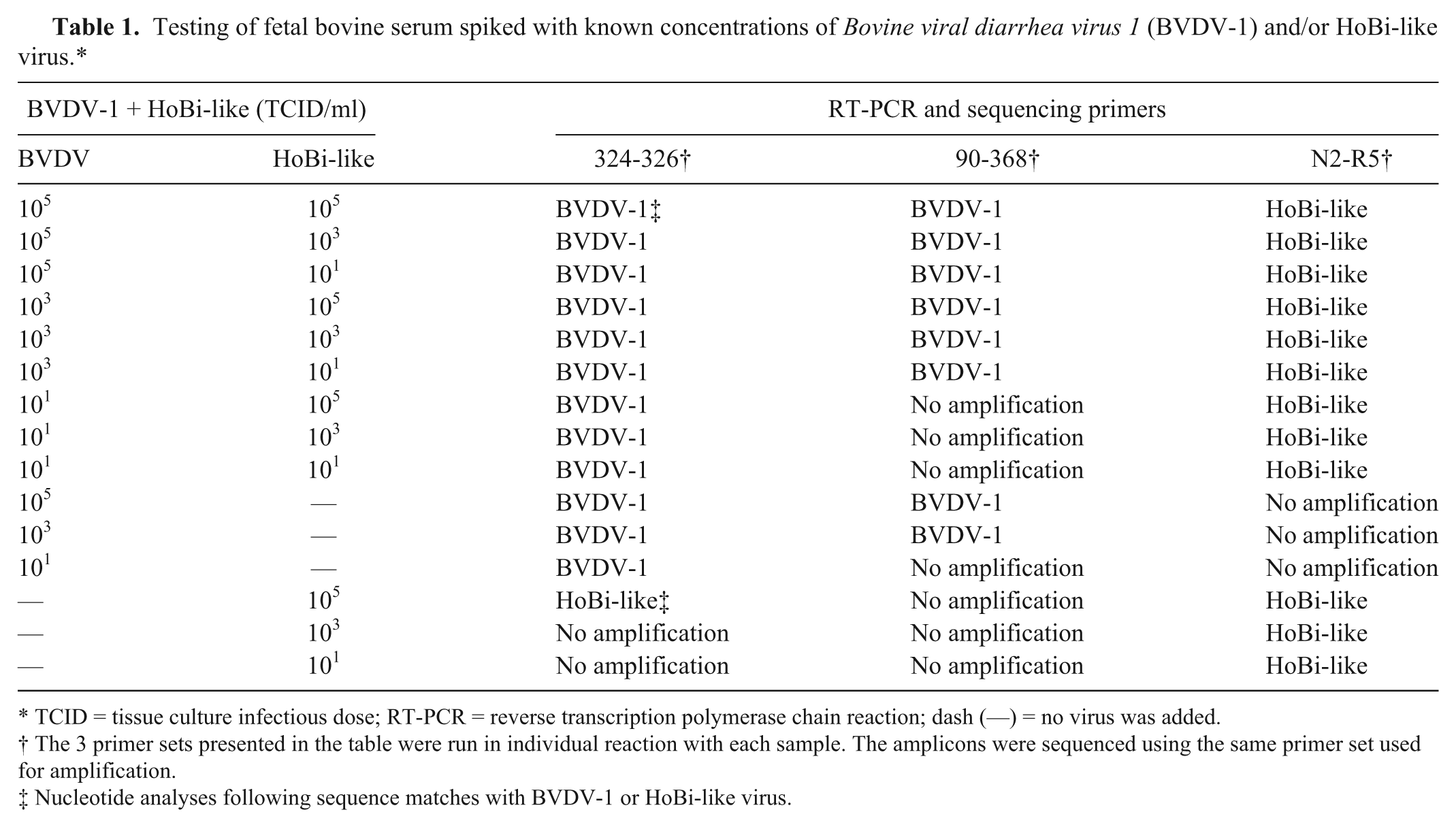
TCID = tissue culture infectious dose; RT-PCR = reverse transcription polymerase chain reaction; dash (—) = no virus was added.
The 3 primer sets presented in the table were run in individual reaction with each sample. The amplicons were sequenced using the same primer set used for amplification.
Nucleotide analyses following sequence matches with BVDV-1 or HoBi-like virus.
Bovine viral diarrhea virus RT-PCR
The primer pairs HCV90-368 and 324-326 were used as previous described15,24 and hereafter referred to as BVDV primer pairs. The PCR products were not cloned but sequenced directly in both directions, and all samples were done in duplicate. Sequencing templates were labeled according to manufacturer’s recommendations using commercial available chemistries and sequenced e using a genetic analyzer. f Phylogenetic analyses 23 were performed using software.g,h
HoBi-like virus–specific detection
HoBi-like viruses–specific quantitative RT-PCR was performed as previously described. 12 Samples were run in a thermocycler i in quadruplicate (2 runs with 2 duplicates per run). Also, an in-house HoBi-like–specific one-step RT-PCR using the primers forward N2 (TCGACGCATCAAGGAATGCCT) and reverse R5 (TAGCAGGTCTCTGCAACACCCTAT) was run in parallel. The reaction amplifies an approximately 150 nucleotide fragment from the 5′-UTR. A 3-µl aliquot of total RNA was amplified using a commercial kit j following manufacturer’s recommendations. Reaction conditions included reverse transcription at 55°C for 25 min, followed by 2 min at 94°C, 35 cycles of 94°C for 30 sec, 55°C for 30 sec, 68°C for 25 sec, with a final extension at 68°C for 5 min. The PCR products were detected by electrophoresis in a 1.0% agarose gel, stained, k and visualized under ultraviolet light. The PCR products were sequenced and phylogenetic analyses performed as described previously. 17
Virus isolation
Virus isolation was carried out on BT cells grown in 24-well plates using protocols described previously for the isolation of BVDV. 18 Three passages of 4 days each were performed using 70% confluent, 24 hr fresh cell monolayer. Plates were kept at 37°C with 5% CO2 and monitored for cytopathic effect during the period. Following the last passage, supernatant was removed and cells were fixed. Immunoperoxidase test (IMPT) was used to detect antigens of noncytopathic pestiviruses in fixed cells, using the monoclonal antibody (mAb) N-2 as previously described. 1 The N-2 mAb reacts with a wide range of pestivirus species, including BVDV, BDV, and HoBi-like virus isolates.1,20 RNA was extracted from the supernatant from the last passage and subjected to RT-PCR as described above.
Antigen-capture ELISA and BVDV antibody detection ELISA
Lots of FBS were individually tested with ACE a and Ab-ELISA, b following the manufacturer’s recommendations. Samples were run in quadruplicate (duplicates in 2 separate runs).
Virus neutralization test
A comparative VNT was performed using the following cytopathic strains BVDV1-NADL and BVDV2-296c, as well as the noncytopathic strain HoBi_D32/00 as previously described, 2 with minor modifications. Briefly, serum was diluted in MEM from 1:2 to 1:256 in 96-well plates, and 200 TCID50 of each virus were added individually in accordance with the World Organization for Animal Health (OIE) standards (30–421 TCID/ml). The BT cells were included after incubation of 90 min at 37°C with 5% CO2. After 4 days of incubation, dilution endpoints were obtained for strains BVDV1-NADL and BVDV2-296c by observing cytopathic effect in cell monolayer and for HoBi_D32/00 by IMPT as described. 1 Sera, collected from convalescent animals infected under controlled conditions with BVDV or HoBi-like viruses, with known neutralizing antibody titer, were used as positive controls. Samples with titer ≥1:4 were considered positive.
Results
Pestivirus detection in FBS contaminated with known concentration of HoBi-like virus and/or BVDV-1
The 90-368 primer set did not amplify the HoBi-like virus strain at any of the concentrations tested while the 324-326 primer set amplified the HoBi-like strain at a TCID of 105. When these BVDV primers were used to amplify RNA extracted from FBS spiked with both a HoBi-like virus and a BVDV virus, sequences from the BVDV virus were preferentially amplified. This was true even when the TCID of HoBi-like virus was 10,000 times higher than that of the BVDV-1 (Table 1). Conversely, the primer pair N2-R5 (HoBi-like specific) amplified the HoBi_D32/00 sequence even when the concentration of BVDV-1 was 10,000 higher than that of the HoBi-like virus. Thus, it did not appear that the presence of multiple pestivirus species affected detection of HoBi-like viruses using HoBi-like–specific primer set N2-R5.
Fetal bovine serum pestivirus detection
None of the lots in G1 tested positive by PCR using HoBi-like–specific primers, while 2 lots (G1-#24 and G1-#48) tested positive by PCR using both sets of BVDV primers. Lot G1-#48 was also positive by VI (Table 2). Phylogenetic analysis of amplicons demonstrated that the amplified viral sequences were derived from BVDV-1b isolates.
Pestivirus detection in fetal bovine serum lots using Bovine viral diarrhea virus 1 and 2 (BVDV-1 and -2) and HoBi-like virus–specific diagnostic assays. Samples with negative results in all performed tests are not displayed.*
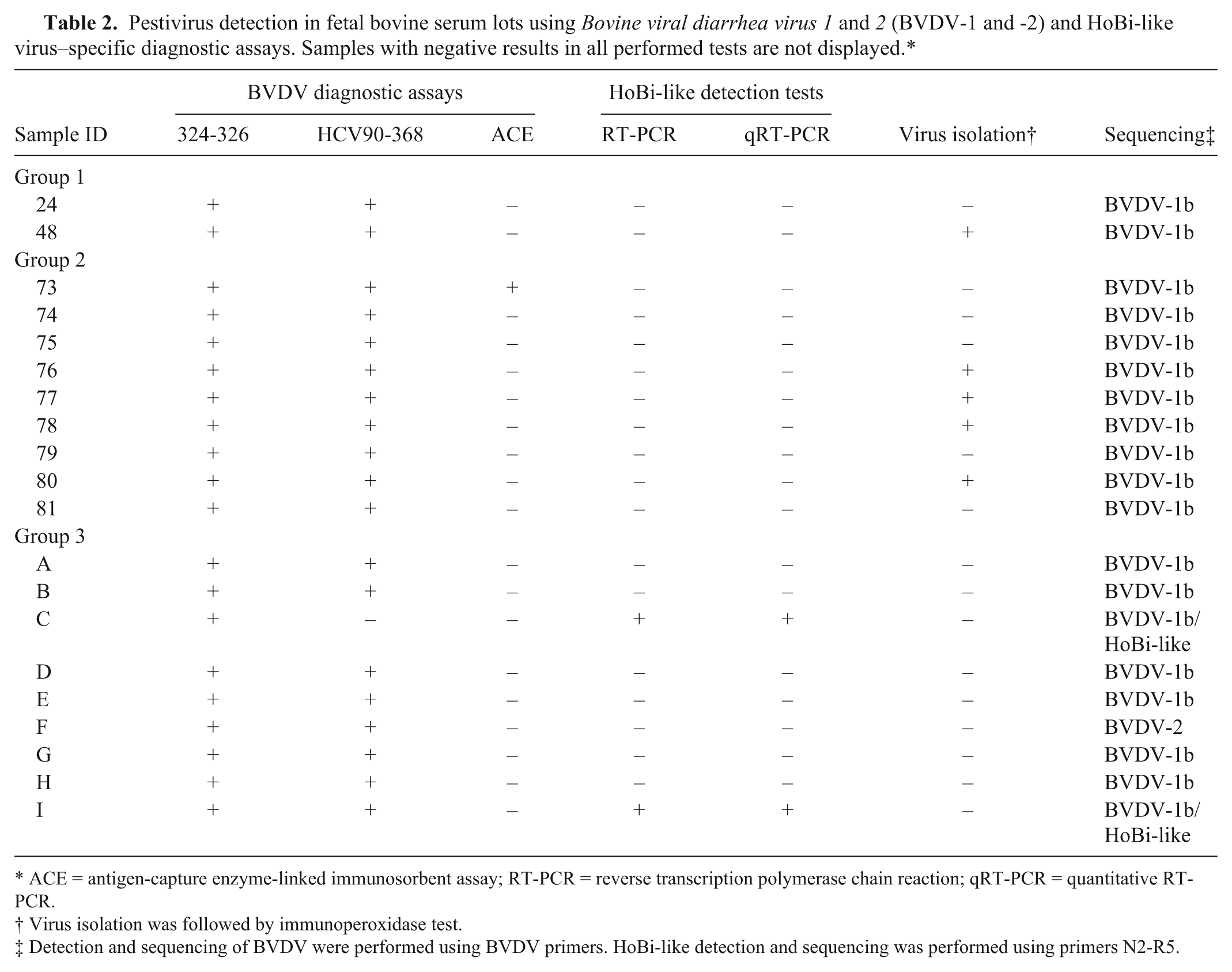
ACE = antigen-capture enzyme-linked immunosorbent assay; RT-PCR = reverse transcription polymerase chain reaction; qRT-PCR = quantitative RT-PCR.
Virus isolation was followed by immunoperoxidase test.
Detection and sequencing of BVDV were performed using BVDV primers. HoBi-like detection and sequencing was performed using primers N2-R5.
None of FBS lot samples in G2 were detected as positive for HoBi-like viruses based on PCR. In contrast, all 9 lots tested positive using the BVDV primers. Phylogenetic analyses revealed that the contaminating viruses belonged to the BVDV-1b subgenotype. Lots G2-#76, G2-#77, G2-#78, and G2-#80 were also positive by VI. The sequence of the isolated viruses matched the sequence directly amplified from these lots. Only one of the samples (G2-#73) was detected as positive by ACE. a
The G3 group included samples from 2 lots (G3-C and G3-I) previously identified as HoBi-like virus positive. 25 Eight of the 9 samples in G3 tested positive for BVDV with both BVDV reactions. Sample G3-C was detected as positive using the primers 324-326 but not primers HCV90-368. Phylogenetic analyses of amplified sequences demonstrated that 8 samples were contaminated with BVDV-1 (G3-G) and 1 with BVDV-2. G3-C and G3-I tested positive using both HoBi-like virus–specific tests. The amplicons were confirmed as HoBi-like viruses by sequencing followed by phylogenetic analysis. All 9 samples were negative by ACE a and VI.
Antibody detection
None of the tested lots contained antibodies that neutralized HoBi_D32/00 while 7 lots had antibodies against BVDV by VNT and/or Ab-ELISA b (Table 3). Four were from G1 (#41, #42, #50, #51), 1 from G2 (#80), and 2 from G3 (F and H). Three positive samples in G1 (#41, #50, #51) were detected with Ab-ELISA b and displayed higher antibody titer to BVDV-1 than to BVDV-2. Sample #42 was negative using Ab-ELISA. b By VNT, this sample had a low titer against BVDV-1 (1:4) but no detectable titers against BVDV-2 or HoBi-like. The single antibody positive sample in G2 (#80) was detected using Ab-ELISA, b while no antibodies to BVDV-1, BVDV-2, or HoBi_D32/00 were detected by VNT. Two samples in G3 (F and H) were detected as antibody positive using the Ab-ELISA b and VNT. The neutralizing antibodies titers of both samples were similar against BVDV-1 and BVDV-2 but had no detectable titers against the HoBi-like strain.
Detection of antibodies against pestiviruses in fetal bovine serum lots. Samples with negative results in all performed tests are not displayed.*
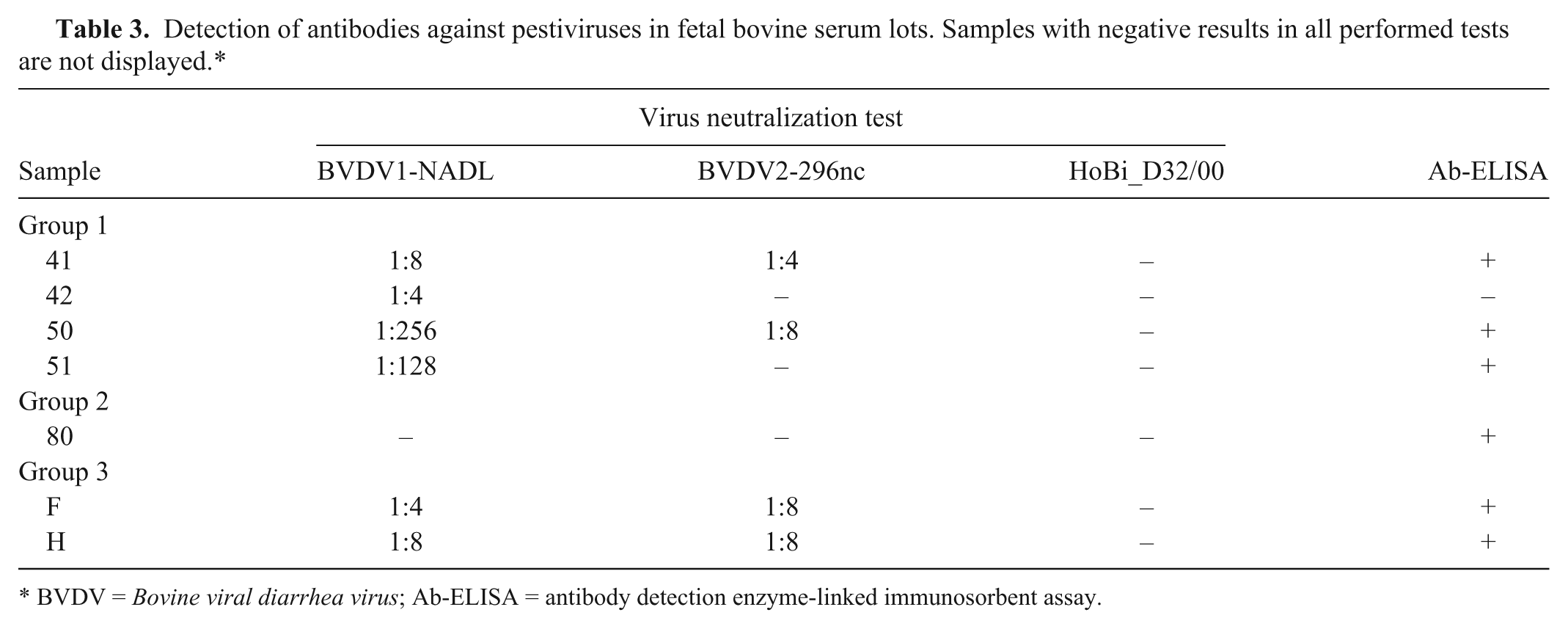

BVDV = Bovine viral diarrhea virus; Ab-ELISA = antibody detection enzyme-linked immunosorbent assay.
Discussion
While no HoBi-like virus persistently infected (PI) animals have been detected while monitoring for BVDV PI animals in the United States, the presence of HoBi-like viruses in the country cannot be ruled out. The report of HoBi-like viruses in FBS processed in Europe and labeled as originating in the United States 25 raised a great deal of concern. The current study confirmed the presence of HoBi-like viruses in 2 samples previously described as positive for these viruses. Nucleotide sequencing demonstrated that these samples correspond to the sequences available at GenBank (available at http://www.ncbi.nlm.nih.gov/pubmed/, identification JN967714.1 and JN967748 for samples C and I, respectively). Both samples were also contaminated with BVDV-1.
It was observed both in the FBS spiked study and in the testing of samples from lots C and I that only BVDV sequences were amplified using the 2 sets of primers frequently used for BVDV surveillance, 90-368 and 324-326, and that HoBi-like sequences were only detected when the HoBi-like–specific primers were used. This is probably related to the nucleotide mismatches between viral and primer sequences (Fig. 1), as noted in other studies. 4 Thus, the detection and identification of HoBi-like viruses required the application of specific HoBi-like primers. Although the genetic diversity of HoBi-like viruses is not fully known, it is probable that, like the other members of the pestivirus genus, they will be genetically diverse. Thus, effective surveillance should not rely on a single test. It is reported that a TaqMan real-time RT-PCR 12 and a nested RT-PCR 4 can discriminate between BVDV species and HoBi-like viruses. However, further investigation is needed to determine specificity and sensitivity using a variety of pestiviruses species in different combinations and concentrations.

Alignment of pestivirus species with the Bovine viral diarrhea virus (BVDV) primer pairs (324-326 and HCV90-368) and the HoBi-like–specific primer pair (N2-R5).
Detection of BVDV-contaminated samples in all 3 groups, including G1 (claimed negative for pestiviruses by commercial processor) was not surprising based on previous studies. 8 Phylogenetic analyses revealed that, from a total of 20 positive samples for BVDV, 19 belong to BVDV-1 and 1 to BVDV-2. This finding likely reflects the higher distribution and prevalence of BVDV-1b in North America. 19 It should be noted that the recovery of viable viruses was successful only from non–gamma-irradiated samples: lots G1-#48, G2-#76, G2-#77, G2-#78, and G2-#80. Thus, it appears gamma irradiation is effective at some level. However, the present study was not designed to test the effectiveness of irradiation; therefore, no assumption about the safety of sterilizing process can be made.
The ACE a test is highly indicated to detect BVDV PI animals, and the effectiveness has been demonstrated in detecting HoBi-like in supernatants of infected cells. 1 The high number of undetected samples using ACE is probably due to pooling of samples to compose a FBS batch, leading to excessive antigen dilution.
Using the Ab-ELISA b and VNT, 7 samples (7.8%) were detected by Ab-ELISA b and/or VNT. Lot G2-#80 was positive by Ab-ELISA, b but negative by VNT. One possible reason for this discrepancy is that viable virus (BVDV-1) present in G2-#80 might have interfered with the VNT. In contrast, sample G1-#42 was negative by Ab-ELISA b but positive by VNT when tested against BVDV-1 (titer 1:4). The low VNT titer may explain the lack of recognition using Ab-ELISA. b Virus neutralization using BVDV-1 and Ab-ELISA b demonstrated equivalent sensitivity (85.7%) to detect antibodies in FBS, although both assays have drawbacks. An optimum diagnostic may require a combination of tests; in the current study, combined sensitivity of tests used reached 98%.
It is possible that the 2 lots of FBS serum, labeled as U.S. origin, were contaminated with HoBi-like viruses during processing in Europe. One of the FBS lots, labeled as U.S. origin, was found to be contaminated with a BVDV-1d strain. While the circulation of isolates from the BVDV-1d subgenotype have been reported in Europe, 9 to date no isolates from this subgenotype have been found in the United States.5,7,17,19 Thus, it is probable that the contamination of this FBS lot with BVDV-1d occurred during processing in a European plant that handled FBS originating from several different continents using the same equipment. Contamination with HoBi-like virus could have occurred through a similar route.
The current findings reinforce the need for a constant update in pestivirus diagnostics and the use of multiple tests. It also points to the need of further regulations for FBS trade, including the development and validation of internationally accepted tests and protocols. The documentation of chain of custody, country of processing, as well as country of origin must be included in labeling. Pooling of FBS lots originated from different parts of the world should be avoided. The use of the equipment to manufacture FBS with different origins may also need to be reviewed, in order to avoid contamination among lots. The data herein presented found no evidence of HoBi-like virus in FBS originated in North America and manufactured in the United States, although further efforts and constant monitoring are required to validate the status and avoid introduction and/or dissemination of these agents in the country.
Footnotes
Acknowledgements
The authors thank Matthew Carpenter for consulting and thank Kathryn McMullen, Patricia Federico, and Renae Lesan for technical assistance.
a.
HerdChek BVD Antigen Test Kit, IDEXX Laboratories, Westbrook, ME.
b.
PrioCHECK BVDV Ab, Prionics AG, Schlieren-Zurich, Switzerland.
c.
Qiacube, Qiagen GmbH, Hilden, Germany.
d.
QIAamp Viral RNA Mini Kit, Qiagen GmbH, Hilden, Germany.
e.
Terminator BigDye v3.1, Invitrogen Corp., Carlsbad, CA.
f.
3130xl Genetic Analyzer, Invitrogen Corp., Carlsbad, CA.
g.
BioEdit v7.2.0, Tom Hall Ibis Biosciences, Carlsbad, CA.
h.
Mega5 v.5.2.2, Center for Evolutionary Medicine and Informatics, Tempe, AZ.
i.
DNA Engine Opticon2 cycler, MJ Research, Bio-Rad, Hercules, CA.
j.
SuperScript III one-step RT-PCR system with Platinum Taq high fidelity, Invitrogen Corp., Carlsbad, CA.
k.
GelRed, Biotium, Hayward, CA.
Authors’ note
Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) received no financial support for the research, authorship, and/or publication of this article.
