Abstract
Canine distemper virus (CDV) remains a common cause of infectious disease in dogs, particularly in high-density housing situations such as shelters. Vaccination of all dogs against CDV is recommended at the time of admission to animal shelters and many use a modified live virus (MLV) vaccine. From a diagnostic standpoint for dogs with suspected CDV infection, this is problematic because highly sensitive diagnostic real-time reverse transcription polymerase chain reaction (RT-PCR) tests are able to detect MLV virus in clinical samples. Real-time PCR can be used to quantitate amount of virus shedding and can differentiate vaccine strains from wild-type strains when shedding is high. However, differentiation by quantitation is not possible in vaccinated animals during acute infection, when shedding is low and could be mistaken for low level vaccine virus shedding. While there are gel-based RT-PCR assays for differentiation of vaccine strains from field strains based on sequence differences, the sensitivity of these assays is unable to match that of the real-time RT-PCR assay currently used in the authors’ laboratory. Therefore, a real-time RT-PCR assay was developed that detects CDV MLV vaccine strains and distinguishes them from wild-type strains based on nucleotide sequence differences, rather than the amount of viral RNA in the sample. The test is highly sensitive, with detection of as few as 5 virus genomic copies (corresponding to 10−1 TCID50). Sequencing of the DNA real-time products also allows phylogenetic differentiation of the wild-type strains. This test will aid diagnosis during outbreaks of CDV in recently vaccinated animals.
Introduction
Canine distemper virus (CDV; order Mononegavirales, family Paramyxoviridae, subfamily Paramyxovirinae, genus Morbillivirus) remains a common cause of infectious disease in dogs, despite vaccines that produce long-lasting immunity. This is particularly true in intensive housing situations, such as shelters. 12 Current guidelines recommend vaccination of all dogs against CDV at the time of admission to animal shelters, 12 and though a recombinant CDV vaccine is available and effective in these high-risk environments, 11 modified live virus (MLV) vaccines remain more affordable and are still being used by local shelters and clinics.
Several outbreaks have occurred in shelters in eastern Tennessee and also in a local pet store. In each case, the affected dogs had been recently vaccinated for CDV with MLV vaccines. This poses a problem from a diagnostic stand point, given that CDV MLV vaccine can be detected in the blood, urine, and swabs by highly sensitive real-time reverse transcription polymerase chain reaction (RT-PCR) tests, including the one routinely used in the University of Tennessee, College of Veterinary Medicine, Clinical Virology Diagnostic Laboratory (Knoxville, Tennessee). 7 This vaccine interference is recognized to limit the effectiveness of a real-time RT-PCR test for diagnosis of CDV and has prompted a commercial laboratory to develop a test a to deal with this problem. This test provides a quantitative measure of the CDV viral load. Viral load is determined from the threshold cycle (Ct) value obtained, which increases with decreasing amount of viral template in the sample. This quantitative information allows discrimination of vaccine interference from infection with a wild-type strain of distemper virus as the amount of virus present during infection is typically exponentially more than would be detected because of recent vaccination (IDEXX: 2011, Canine distemper virus (CDV) Quant RealPCR™ test differentiates between naturally infected and vaccinated dogs. Available at: http://www.idexx.com/view/xhtml/en_us/corporate/news/press-releases/20111011pr.jsf. Accessed August 21, 2013). While helpful, wild-type virus shedding varies during the course of infection, and this assay is unable to differentiate between vaccine and wild-type strains when virus shedding is low (<1 million particles). This is particularly problematic during acute infection in animals that have been recently vaccinated. Also, there are other variables that may affect Ct values besides the concentration of the target, including artifacts from the reaction mix or instrument that change the fluorescence measurements associated with the Ct calculation and may result in template independent changes to the Ct value (Applied Biosystems: 2011, Real time PCR, understanding Ct. Available at: http://www3.appliedbiosystems.com/cms/groups/mcb_marketing/documents/generaldocuments/cms_053906.pdf. Accessed August 21, 2013).
There have been multiple gel-based PCR protocols designed to distinguish vaccine strains from wild-type strains.2–4,6,15,20,22,24 Such tests are designed to take advantage of sequence differences between the vaccine strains and wild-type strains. Most of the tests amplify regions of the hemagglutinin (H) gene4,6,15,24 or the matrix gene–fusion gene (M-F) intergenic region,3,20 which have been discovered to be variable among CDV strains.13,15 Canine distemper virus strains identified in different geographical settings and from various animal species have revealed that the H gene/glycoprotein is a reliable target to reconstruct and investigate the genetic relationships among CDV strains. The majority of CDV field strains cluster into different genetic lineages, designated America-1 and -2, Asia-1 and -2, European, European wildlife, Arctic, and Africa, based on the geographic regions where the strains have been detected.15,17,23 Many of these gel-based PCR protocols also differentiate among these various strains, but several of them are unable to detect or differentiate the contemporary American strains. 3
Over the past 5 decades, CDV isolates from the America-1 lineage were applied in vaccine production and used as conventional distemper vaccines. These include the Snyder Hill strain isolated in Ithaca, New York, in the 1950s from the brain of a dog and the Onderstepoort strain, used worldwide as an attenuated live vaccine, which dates back to a disease outbreak among North American ranched foxes in the 1930s. 15 America-1 CDV strains have not been detected in domestic dogs over the past 5 decades, and it is thought that they are no longer circulating in the field. However, these strains have been reported occasionally in wildlife animals in the United States, suggesting vaccine-induced disease, eventually followed by adaptation of vaccine viruses in some animal populations, or persistence of America-1 CDV strains in certain ecological settings. 14
All the currently used MLV vaccines except one 5 contain an America-1 strain and the existing gel-based protocols differentiate between this America-1 lineage and all the other lineages. The one exception is the MLV vaccine that contains a Rockborn-like strain, which more closely resembles America-2 strains and therefore makes differentiation of this vaccine strain impossible with most of the published methods. 5 One protocol has been developed to differentiate this Rockborn-like or “contemporary vaccine strain” from the Asia-1 lineage, and this particular protocol may be able to differentiate the Rockborn-like strain from currently circulating American strains; however, the proposed method, which requires restriction enzyme analysis, was not tested. 3 Many of these differentiating assays are RT-PCR reactions, combined with restriction fragment length polymorphism.2,4,6,22 While effective, this is time consuming and gel-based PCR assays are not as sensitive as real-time RT-PCR assays. Gel-based nested PCR tests can be as sensitive as real-time PCR tests, and some of the published protocols are nested reactions.6,15,20 However, these tests are inherently prone to contamination and are also time consuming. 24 Test sensitivity is usually not an issue for most diagnostic samples, which contain large amounts of CDV RNA, 15 but recently, during CDV outbreaks in area shelters and a pet store, CDV RNA was detected in recently vaccinated animals, and it was not possible to determine if the RNA was from the MLV vaccine strain or the wild-type strain using the most sensitive differentiation assay that has been described in the literature. 20 Therefore, a highly sensitive real-time RT-PCR assay was developed for differentiation of vaccine strains from wild type strains based on sequence differences, not just differences in sample viral load.
Materials and methods
Samples
RNA previously extracted and stored at −80°C, from clinical samples (including conjunctival swabs and urine) submitted to the University of Tennessee, College of Veterinary Medicine, Clinical Virology Laboratory from 2010 to 2013, that previously tested positive for CDV by detection of genomic RNA with a real-time RT-PCR assay that detects a conserved region of the genome, 7 was evaluated with this new assay. Assay validation was done with 10 samples with Ct values ≤25. 7 Three of the commonly used MLV vaccines,b -d Snyder Hill and Onderstepoort strains were also evaluated. Functionality of the test was determined with 17 clinical samples (2 from 2010, 3 from 2011, 3 from 2012, and 9 from 2013) that previously tested positive with Ct values ≥ 31 (approximately 5,000 genomic copies). This included 5 samples from independent outbreaks at two area shelters and a local pet store in eastern Tennessee from 2012 to 2013.
Real-time RT-PCR
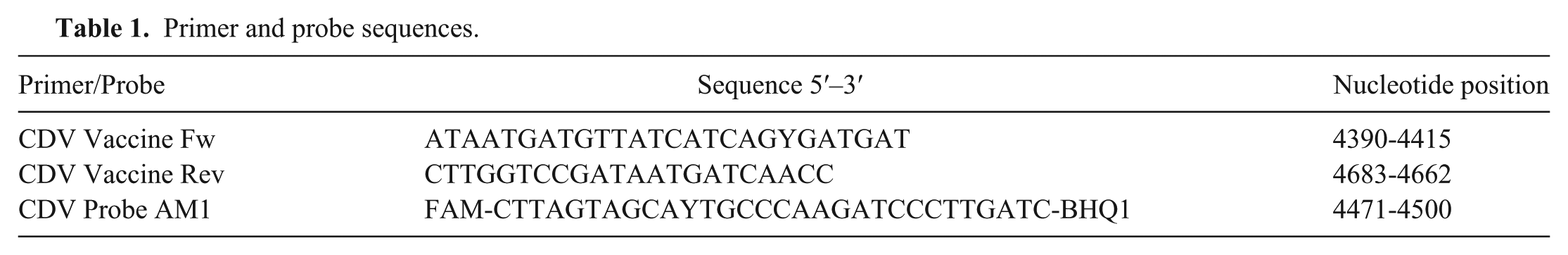
RNA was extracted e according to the manufacturer’s instructions. Primers were designed 18 to amplify a 294-bp portion of the CDV M gene and the M-F intergenic region (Table 1). Conserved areas were used for primer design to allow detection of all CDV isolates, including wild-type and vaccine strains (America-1 lineage and the contemporary vaccine). A probe was developed 18 to detect vaccine strains of the America-1 lineage,b,c based on an area between the primers that is variable between the America-1 vaccines and the wild-type strains (Table 1). Five-microliter RNA samples were run in 25-µl total volume reactions using a commercially available master mix f for real time RT-PCR. The optimized mix contained 200 µM deoxyribonucleotide triphosphates, 3 mM MgCl2, 200nM of the FAM-labeled probe, and 300 nM of each primer. One unit of RNase inhibitor g was also added to the mix. Samples were amplified h with a RT step at 50°C for 30 min, activation step for the hot start Taq polymerase at 95°C for 10 sec, followed by 45 cycles of denaturation at 95°C for 5 sec, annealing at 48°C for 1 min, and elongation at 60°C for 1 min. Various reaction conditions were evaluated, and these were found to be optimal. Water was used as a negative control for each run and standard RNA (see below) was used as a positive control. The real-time PCR products were electrophoresed on a 1.5% Tris–borate–EDTA (TBE) agarose gel stained with ethidium bromide, and visualized by ultraviolet light with a CCD camera system. i The PCR products were purified j and directly sequenced k in both the forward and reverse directions.
Primer and probe sequences.
Sensitivity, specificity, and reproducibility
Test sensitivity was determined by preparing standard RNA by cloning the PCR products l and performing in vitro transcription m from purified plasmids. n The standard RNA was purified, o and the RNA concentration was determined with a spectrophotometer. p The number of RNA copies in the sample was estimated based on the molecular weight of the transcription product and the RNA concentration. Ten-fold serial dilutions were prepared, and aliquots of each dilution were subjected to the real-time RT-PCR assay to determine the lowest number of RNA copies the assay can detect.
Sensitivity was also evaluated by testing dilutions of vaccine virus with a known titer. Vaccine virus b was replicated in Vero (African green monkey kidney epithelial) cells and harvested from the cell culture by freeze-thaw cycles; cellular debris was removed by centrifugation (1,000 × g) and the virus in the supernatant was titrated by the Reed and Muench method as previously described. 1 Ten-fold serial dilutions of the virus were prepared, and RNA was extracted from each dilution. e The RNA was then tested by the real-time RT-PCR assay.
Test specificity was evaluated by 1) testing sequenced wild-type strains and vaccine strains, and by 2) testing DNA or RNA from other pathogens known to produce “kennel cough” in dogs (Canine adenovirus A, Canid herpesvirus 1, Influenza A virus, Parainfluenza virus 5, Respiratory coronavirus, and Bordetella bronchiseptica).
Test reproducibility was determined by calculating intra- and interassay coefficient of variation (CV), using at least 3 replicates each of the 10-fold serial dilutions of the standard RNA, and a standard curve was produced. The efficiency of the assay was determined with the following calculation: Efficiency = 10(–1/slope) – 1.
Phylogenetic analysis
A portion of the hemagglutinin gene was amplified using a previously described protocol. 9 The PCR product was electrophoresed on a 1.5% TBE agarose gel stained with ethidium bromide, and visualized by ultraviolet light with a CCD camera system. i The PCR products were purified q and sequenced k using the both the forward and reverse primers for phylogenetic evaluation and aligned with H gene CDV sequences in GenBank from other geographic regions. This data was used to validate the assay’s ability to distinguish wild-type strains from vaccine strains. The sequences from the real-time PCR products from the M gene/M-F intergenic region were also compared with similar sequences in GenBank. Chromatograms were manually evaluated and all the sequences, including those used from GenBank, were trimmed to approximately equal length. Multiple sequence alignments were performed online using M-COFFEE, a meta-alignment server 16 that uses a variety of alignment algorithms to compute alignments, and visualized and edited using a bioinformatics software platform. r Consensus unrooted phylogenetic trees were generated within the software using the Tamura–Nei model 21 with unweighted pair group method with arithmetic mean clustering and 1,000 bootstrap replicates.
Results
Real-time RT-PCR
The MLV America-1 vaccine strains and the Snyder Hill and Onderstepoort strains were all positive by real-time RT-PCR. The probe did not detect the RNA from the 10 clinical samples with low Ct values, as expected, because the binding region of the probe shows specificity for vaccine strains versus wild-type strains, with the exception of the “contemporary vaccine strain” d (Fig. 1). However, bands of the appropriate size from these strains were visible by gel electrophoresis. The probe did not detect 14 out of 17 of the clinical samples with Ct values ≥31, but 13 out of 14 had a band of appropriate size observable by gel electrophoresis (Fig. 2). The sample without a detectable band or fluorescence was from 2011 (Ct 39.4). Of the 3 samples from which virus were detected with the vaccinal probe, 2 were from 2010 and 1 was from 2013.
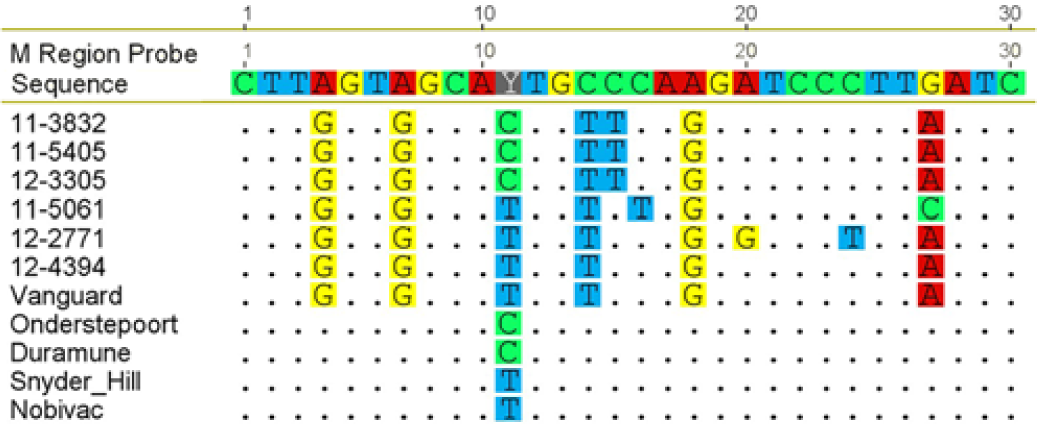
Alignment of real-time reverse transcription polymerase chain reaction probe with sequences from clinical samples (depicted as laboratory accession numbers, which include 2-digit year of collection) and with sequences from Canine distemper virus vaccine strains, including America-1 type strains (Onderstepoort, Duramune, Snyder Hill, and Nobivac) and a Rockborn-like “contemporary” vaccine (Vanguard). Alignments were produced and viewed a bioinformatics software program. r The probe is specific for detection of America-1 vaccine strains, based on sequences differences between these strains and the wild-type strains and Vanguard vaccine in this region.
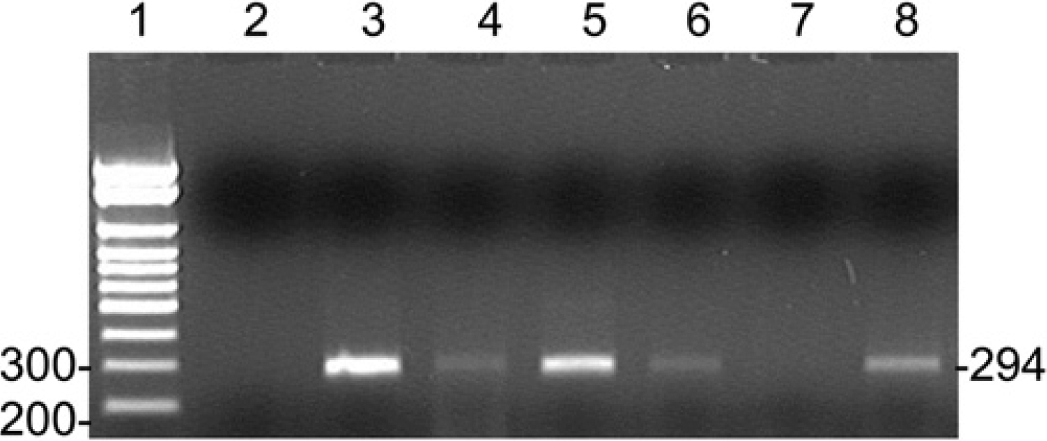
Detection of real-time reverse transcription polymerase chain reaction (RT-PCR) products by gel electrophoresis. A 294-bp product was produced by the real-time RT-PCR assay from clinical samples with a low level of virus present, representative gel showing 6 of the samples tested. Lane 1: ladder; lane 2: water; lane 3: 12-3526, Ct 33.81; lane 4: 13-1670, Ct 40; lane 5: 12-6031, Ct 38.51; lane 6: 11-3826, Ct 34.31; lane 7: 11-4381, Ct 39.4; lane 8: 13-3628, Ct 37.92.
Sensitivity, specificity, and reproducibility
There was no amplification from non-CDV agents, including Canine adenovirus A, Canid herpesvirus 1, Influenza A virus, Parainfluenza virus 5, Respiratory coronavirus, and B. bronchiseptica. The assay was highly specific, with detection of approximately 5 copies of each vaccine virus, which compared to 0.1 TCID50 of titrated vaccine virus stock. The standard curve produced with dilutions of the standard RNA was linear over a wide range of dilutions (R = 0.996; Fig. 3). The assay was reproducible, with intra-assay CVs ranging from 0.005 to 0.01 and interassay CVs of 0.005–0.027. The efficiency of the assay was 82%.
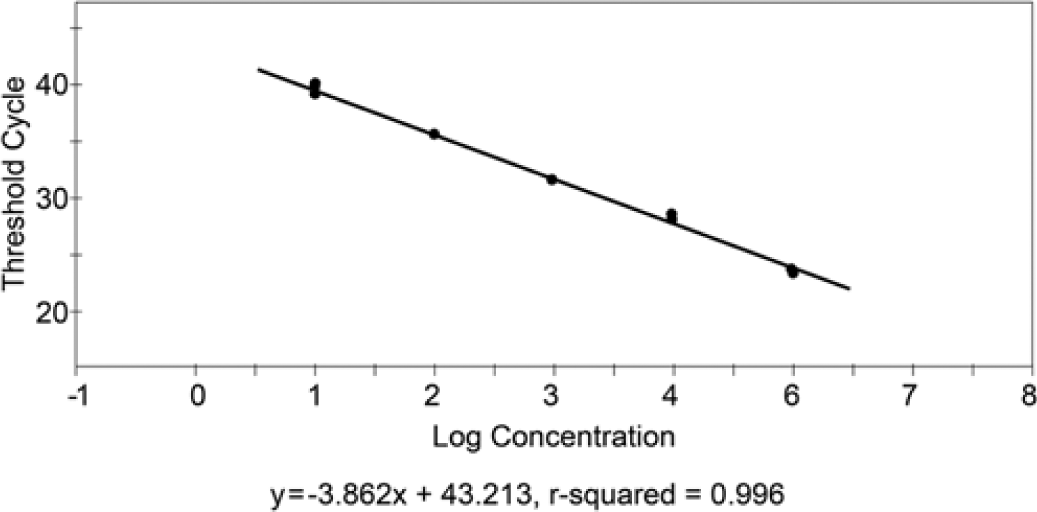
Standard curve produced from 10-fold serial dilutions of standard Canine distemper virus RNA. The curve is linear over five 10-fold serial dilutions, and the assay detection limit is approximately 5 virus genomic copies.
Phylogenetic analysis
The phylogenetic tree generated from the H gene sequences confirmed the samples used for assay validation were wild-type strains, and as previously demonstrated in the literature, the wild-type strains associated with disease are divergent from the America-1 lineage vaccine strains (Fig. 4).
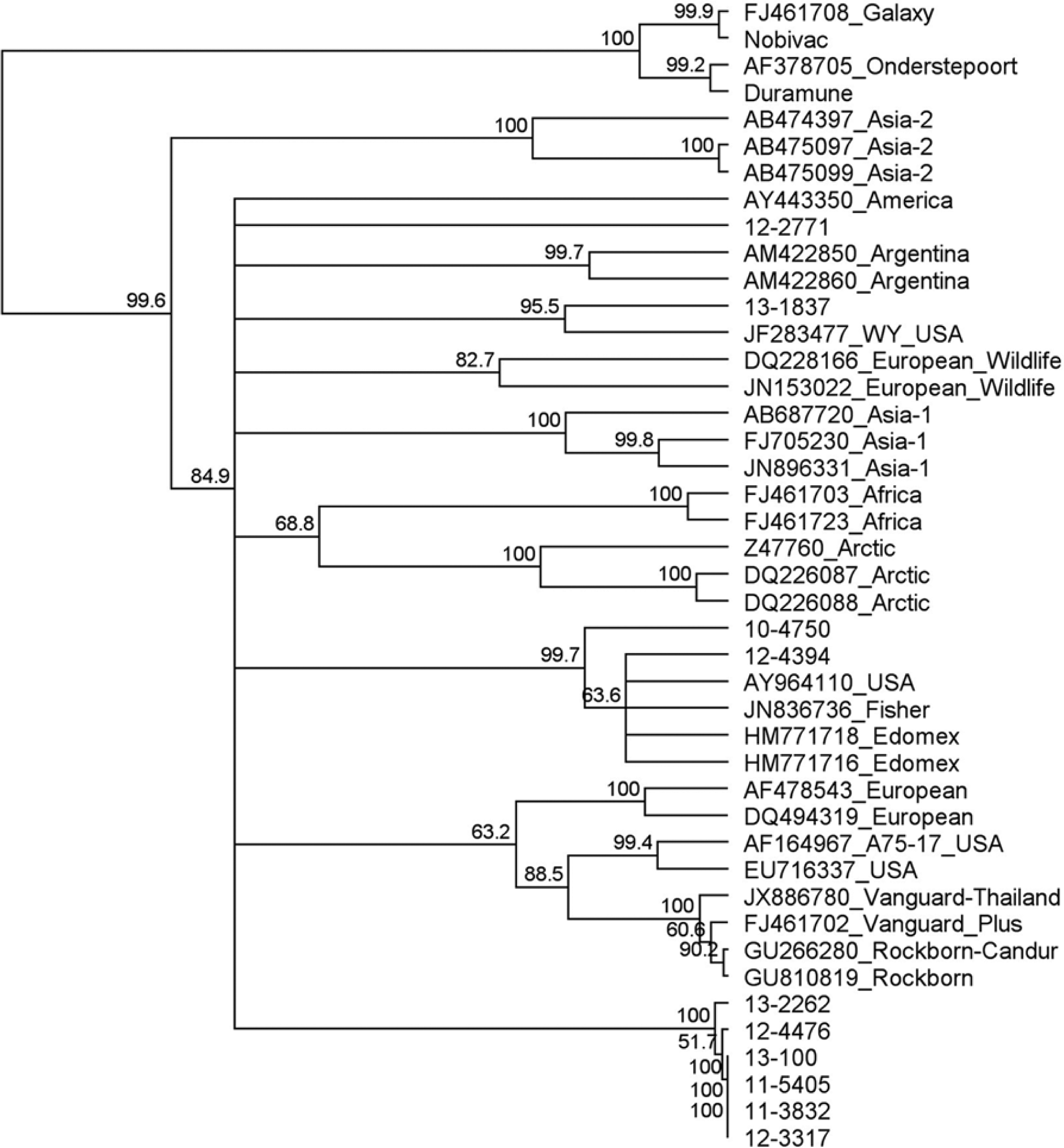
Phylogenetic tree of a portion of the Canine distemper virus (CDV) H gene including representative sequences from GenBank. GenBank accession numbers and geographic region are provided. Clinical samples are represented by the University of Tennessee, College of Veterinary Medicine, Clinical Virology Laboratory accession numbers, which begin with a 2-number designation for the year collected. The America-1 vaccine sequences group separately from wild-type CDV strains. Several strains from this study group closely but separately from the other wild-type strains, representing a distinct clade.
Sequences from the end portion of the M gene/M-F intergenic region produced a tree resembling that produced when using the H gene (Fig. 5). This tree also contained the sequences obtained from the 13 samples that originally had high Ct values, and the phylogenetic analysis identified 1 sample was a Rockborn-like vaccine (100% nucleotide identity with the vaccine strain d ) and the other 12 were wild-type strains.
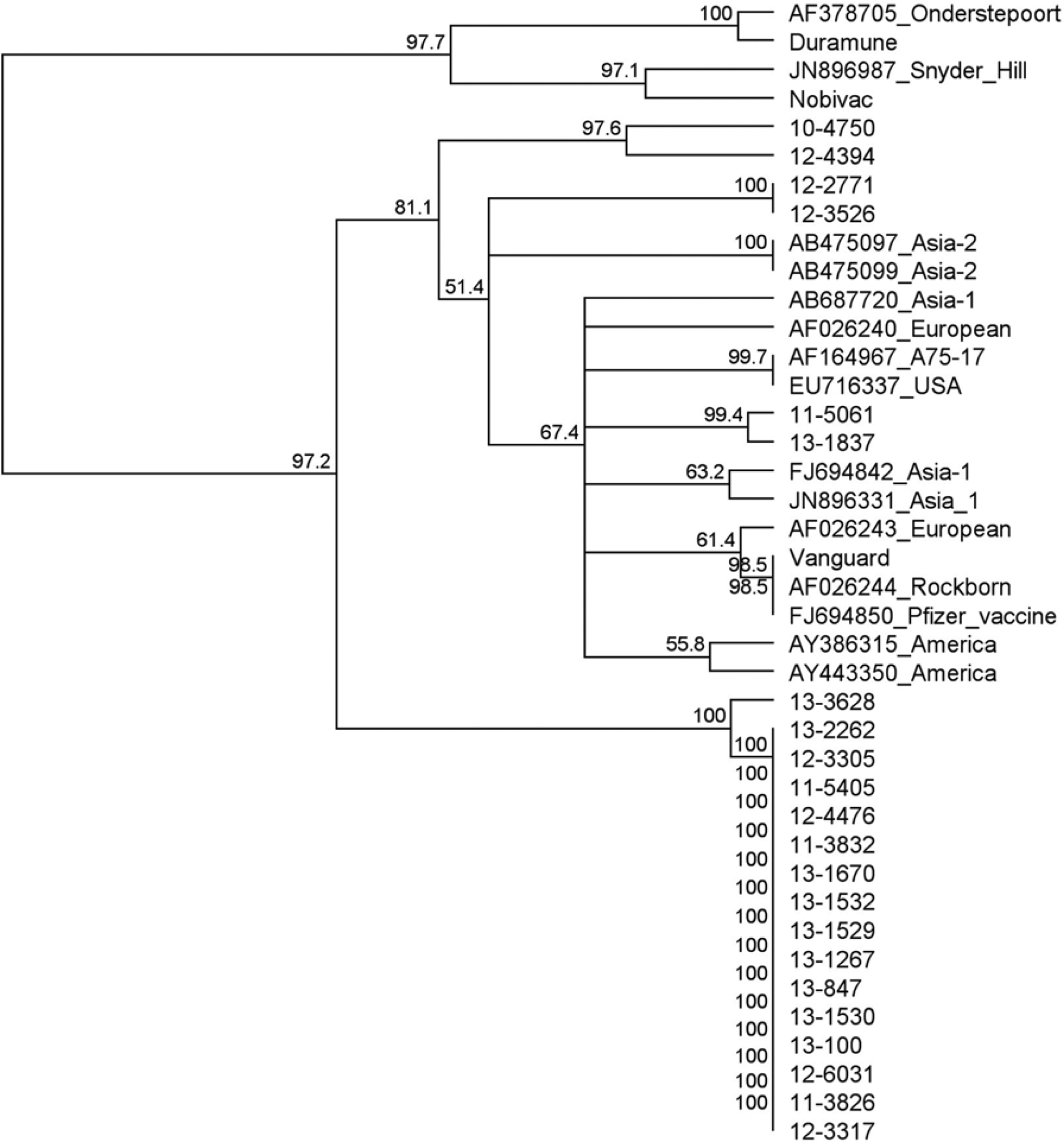
Phylogenetic tree of sequences from the real-time reverse transcription polymerase chain reaction products M gene/M-F intergenic region, including available representative Canine distemper virus sequences from other geographic regions. GenBank accession numbers and geographic region are provided. Clinical samples are provided as laboratory accession numbers, which each begin with the 2-number designation for the year collected. This region is noncoding and more divergent, but the phylogeny is similar to the H gene. It is interesting to note that the sequences in the unique clade from the current study are almost clonal.
Discussion
This assay is useful for detection of MLV vaccine shedding (America-1 strains) in clinical samples to allow rapid and sensitive differentiation from acute infection with wild-type CDV strains. While this test cannot stand alone as a real-time RT-PCR test to detect the wild-type strains (no probe specific for these strains), detection of these strains can be done with gel electrophoresis of the real-time PCR product and differentiation done with sequencing of the product. Attempts were made to design probes to detect the Rockborn-like vaccine strain d and wild-type strains. The variability among the wild-type strains detected, even in this geographic region, made use of a probe that was specific for the Rockborn-like strain difficult, and use of a probe that would detect all of the different geographic strains impractical due to the reduced sensitivity of the assay. This test also has poor efficiency, likely due to the size of the product and the use of a polymorphic site in the forward primer and AM1 probe. While this is not ideal for a real-time PCR assay and the assay certainly cannot be used accurately for quantitative analysis, it does provide the high sensitivity of the real-time assay combined with amplification of products large enough for sequence analysis. The usefulness of this M-F intergenic region for phylogenetic differentiation has been previously described 3 ; however, sequence data for this region of the genome from all of the various geographic strains is lacking.
Based on BLAST comparisons with a portion of the H gene, many of the wild-type strains had sequences similar to sequences already in GenBank, including sequences from fishers that died of CDV in California, in 2004 (JN836736 and JN836737). 10 These included 2 samples, 1 from Alberta, Canada received in 2010 (10-4750) and 1 from an animal shelter in Tennessee (12-4394), which also had 99% nucleotide (nt) identity to a sample from Missouri from 2004, 17 and to “Edomex” strains that were detected in Mexico from 2007 to 2010. 8 Strains in this cluster with high nucleotide identity were actually spread across three countries in North America, and were detectable over at least an 8-year period. A sample from 2013 (13-1837), received from Virginia, had a 98% nt identity to and groups with a strain detected from a CDV outbreak in a pet store in Wyoming in 2010 (JF283477) that was linked to a high volume breeding property. 19 One of the sequences from a sample collected in 2012 from Virginia (12-2771) grouped independently and the closest match in GenBank was the Rockborn-Candur strain, which is a vaccine strain of the Rockborn group. This group has been implicated in vaccine associated disease. 14 Another interesting observation is that many of these sequences from 2011 to 2013 are similar and group independently from the other strains from the United States, as well as other geographic regions. Evaluation of the M gene sequences displays this same grouping, and 9 of the 12 sequences from the samples that had high Ct values also group with this clade. This appears to be a new clade and has been associated with the multiple outbreaks in shelters and in a pet store in eastern Tennessee. Ongoing phylogenetic analysis of new cases of CDV is warranted, particularly as new strains increase in divergence from currently available vaccine strains, potentially leading to significant antigenic changes and vaccine escape. These relationships are demonstrated by both regions sequenced (Figs. 4, 5).
Strains from geographic regions other than America, with the exclusion of the “Edomex” group, were not available for evaluation with this real-time RT-PCR method; however, given the sequence divergence between the probe used here and the strains from the other geographical lineages, this method should work for differentiation of these strains from the America-1 vaccine strains. There was one clinical sample from 2011 that failed to be amplified by the primers used in this assay. While there could be sequence differences in the primer regions with this particular strain, it is more likely the reaction failed due to very little RNA in the sample (previous Ct 39.4) and the age of the sample.
Clinical samples from 2010 that tested positive for vaccine strain had previous Ct values between 30 and 35. These samples were previously reported as positive. It is important for clinicians to adequately interpret PCR results in light of recent vaccination with a MLV vaccine. While this particular assay may only be useful in specific situations, it could mean the difference between life and death for an animal involved in a shelter or pet store outbreak, when management decisions have to be made.
Footnotes
Acknowledgements
The authors would like to thank Ms. Anik Vasington for assistance with figures and Dr. David Bemis for editorial assistance. Disclaimer: The findings and opinions expressed herein belong to the authors and do not necessarily reflect the official views of the U.S. Army or the Department of Defense.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
This work was supported by the Department of Biomedical and Diagnostic Sciences, College of Veterinary Medicine, The University of Tennessee.
a.
The CDV Quant RealPCR Test, IDEXX Laboratories Inc., Westbrook, ME.
b.
ULTRA Duramune, Boehringer Ingelheim Vetmedica Inc., St. Joseph, MO.
c.
Nobivac, Merck Animal Health, Summit, NJ.
d.
Vanguard Plus 5, Zoetis, Florham Park, NJ.
e.
Viral RNA Mini Kit, Qiagen Inc., Valencia, CA.
f.
SuperScript III Platinum One-Step qRT-PCR kit, Invitrogen, Life Technologies, Grand Island, NY.
g.
RNase Out, Invitrogen, Life Technologies, Grand Island, NY.
h.
Smart Cycler II, Cepheid, Sunnyvale, CA.
i.
UVP Inc., Upland, CA.
j.
ExoSAP IT, Affymetrix, Santa Clara, CA.
k.
UT Molecular Biology Core Facility, The University of Tennessee, Knoxville, TN.
l.
TA cloning kit, Invitrogen, Life Technologies, Grand Island, NY.
m.
MEGAscript T7, Ambion, Life Technologies, Grand Island, NY.
n.
Wizard Plus SV Minipreps, Promega Corp., Madison, WI.
o.
Turbo DNA free, Ambion, Life Technologies, Grand Island, NY.
p.
NanoDrop, Thermo Scientific, Wilmington, DE.
q.
QIAquick gel extraction kit, Qiagen Inc., Valencia, CA.
