Abstract
Recently a commercial real-time reverse transcription polymerase chain reaction (RT-PCR) kit has been marketed for the detection of Bovine respiratory syncytial virus (BRSV). However, diagnostic interpretation of the results of this kit requires its comparison to commonly used methods. Therefore, the objective of this study was to evaluate the performance of this kit in comparison with the conventional direct fluorescent antibody test (FAT). Twenty BRSV strains and 14 heterologous bovine viruses were used to check the kit's sensitivity and specificity. The efficiency and detection limit of the kit were determined by testing dilution series of a BRSV strain. The comparison between real-time RT-PCR kit and FAT was performed with 94 clinical samples from calves with clinical signs of respiratory disease including lung tissues (n = 55), transtracheal aspiration samples (n = 20), and nasal swab samples (n = 19). All of the BRSV strains tested were detected by real-time RT-PCR. No cross-reaction was shown with the 14 heterologous bovine viruses. The real-time RT-PCR was 99.3% efficient with a detection limit of 0.1 TCID50 (50% tissue culture infective dose). The results of real-time RT-PCR and FAT were concordant for 65 of the 94 clinical samples tested. The remaining 29 clinical samples were positive by real-time RT-PCR and negative by FAT, demonstrating the higher sensitivity of real-time RT-PCR. In conclusion, the kit evaluated in this study was sensitive, specific, and had a low threshold of detection. Furthermore, the use of this kit instead of FAT allows an improvement of the sensitivity for the detection of BRSV in clinical samples.
Bovine respiratory syncytial virus (BRSV; order Mono-negavirales, family Paramyxoviridae, subfamily Pneumovirinae, genus Pneumovirus) is one of the major causes of lower respiratory tract diseases in cattle worldwide. 16 Infection with BRSV can cause severe respiratory disease in cattle of all ages but affects primarily young calves. 17 This virus acts either alone or in synergy with other agents. 2 Although clinical signs may raise the suspicion of BRSV infection, laboratory confirmation is needed to make a definitive diagnosis. 1 In diagnostic laboratories, direct BRSV detection in clinical samples, such as nasal swab samples, transtracheal aspiration samples, and lung tissues, is currently carried out by antigen detection (fluorescent antibody test [FAT] or immunohistochemistry) or virus isolation. 1,5,10 These laboratory techniques demonstrate poor sensitivity due to low viral titers shed or lability of the BRSV. 8,13 To overcome these problems, several “in-house” reverse transcription polymerase chain reactions (RT-PCR) targeting glycoprotein, 18 fusion, 7,12,18 or nucleoprotein genes 3,14,19 of BRSV have been developed in research laboratories. However, due to differences in suppliers, thermocyclers, and technician skills among laboratories, these in-house polymerase chain reactions are hard to transfer successfully to diagnostic laboratories. Furthermore, these in-house tests are difficult to standardize because of potential inconsistencies in the materials and reagents used.
Recently, a commercial real-time RT-PCR kit a has been marketed for the detection of BRSV. This commercial kit is an alternative to in-house RT-PCR assays. It can guarantee transferability of the technique to diagnostic laboratories and standard quality of the materials and reagents supplied. However, to interpret the results of this kit in a defined meaning in terms of diagnosis, it is necessary 1) to know its performance 20 (i.e., efficiency, detection limit, sensitivity, specificity, and reproducibility) and 2) to compare its results with those of commonly used techniques 20 in diagnostic laboratories, such as FAT, which is the most commonly used method to detect BRSV in clinical samples. 10 The objective of the current study was, therefore, to evaluate the performance of the commercial real-time RT-PCR kit a in comparison with FAT on clinical samples.
Twenty BRSV strains consisting of 1) 18 BRSV field strains isolated at the Health-Agro-Environment Institute (France) including the BRSV strains B35.1 (subgroup II 15 ) and 90504 (subgroup IV 15 ) and 2) 2 BRSV vaccine strains RB94 b (subgroup II 15 ) and 375 c (subgroup III 15 ) were used to determine the sensitivity of the real-time RT-PCR kit. Fourteen heterologous bovine viruses associated with bovine respiratory diseases were used to check the specificity of the real-time RT-PCR kit. These viruses included Bovine parainfluenza virus 3 (BPIV-3; n = 5), Bovine viral diarrhea virus (BVDV; n = 4), Bovine herpesvirus 1 (BoHV-1; n = 4), and Bovine adenovirus (BAV; n = 1). Six steps of a 10-fold dilution series of the BRSV strain B35.1 15 containing 104 TCID50 (50% tissue culture infective dose)/ml were tested in duplicate to determine the efficiency and detection limit of the real-time RT-PCR kit. The efficiency (E) was calculated from the generated curve, using the formula: E = 10(−1/a) −1, where “a” is the slope value of the curve. Reproducibility of the realtime RT-PCR kit was examined by running 10 times 4 steps of a 10-fold dilution series of the external positive control of the real-time RT-PCR kit (cell culture supernatant infected with a BRSV field strain isolated in France in 2003). Mean threshold cycle (Ct), standard deviation (SD), and coefficient of variation (CV) were calculated for each dilution.
Ninety-four clinical samples, including lung tissues (LT; n = 55), transtracheal aspiration samples (TTA; n = 20), and nasal swab samples (NS; n = 19), were used in the present study to compare the results of the real-time RT-PCR kit with FAT. Clinical samples were collected by bovine practitioners from 94 calves with clinical signs of respiratory disease including fever, nasal discharge, and respiratory distress. The LT were taken from the apical and cardiac lobes of lung directly in the farm, placed in a sterile tube on ice pack and transferred within 2 hr to the diagnostic laboratory. Bovine practitioners were advised to carry out TTAs according to a technique previously described. 6 On average, 5–10 ml of fluid was recovered. Recovered fluid was placed in a sterile tube on ice pack and transferred within 2 hr to the diagnostic laboratory. The NS were collected with sterile plastic applicator rayon-tipped swabs d , immediately agitated in 1.5 ml of NaCl 0.9% solution, placed on an ice pack, and transferred within 2 hr to the diagnostic laboratory. Clinical samples were stored at −80°C until analysis. All the clinical samples were tested for BRSV by real-time RT-PCR and FAT.
Real-time RT-PCR was conducted by using the commercial kit a supplied by the manufacturer. This kit contains specific primers and probes designed to detect BRSV and bovine glyceraldehyde-3-phosphate dehydrogenase (GAPDH[B]), equivalent but slightly different from the human GAPDH, which has been chosen as an internal positive control by the manufacturer. The primers and probe targeting the nucleoprotein gene of BRSV were previously described. 3 Total RNA from all samples was extracted and purified using a commercial kit e according to the manufacturer's instructions. In the negative control, sterile RNase-free water was added instead of the specimen. RNA was amplified by a duplex real-time RT-PCR assay, run in a programmable thermal cycler, f using the following conditions: a single cycle of reverse transcription for 30 min at 48°C, 10 min at 95°C for reverse transcriptase inactivation and DNA Taq polymerase activation followed by 45 amplification cycles of 15 sec at 95°C, and 1 min at 60°C each (annealing extension step). External negative (sterile RNase-free water) and external positive controls (cell culture supernatant infected with a BRSV field strain isolated in France in 2003) were included in each run. Only the samples presenting a typical amplification curve with a Ct below 45 were considered positive.
The FAT was performed on NS and TTA fluid cells, or on cryosections of lung. Nasal swab samples and TTA were centrifuged (380 × g) for 10 min at 5°C. Each pellet (cells) was resuspended in phosphate buffered saline (PBS), spotted on a glass slide, air dried, and fixed for 2 min in a mix of 50% acetone and 50% ethanol. Before each analysis by FAT, a cell count was performed to ensure that a minimum of 100 cells were present on NS and TTA. The LTs were examined on frozen sections (5–7 μm) placed on glass slide. Monoclonal antibody 18 B2 g (directed against the BRSV F protein) conjugated with fluorescein isothio-cyanate was applied on slides. The slides were incubated for 15 min at 37°C, washed 2 times in PBS for 5 min each time, and examined under a fluorescence microscope.
The measure of agreement between real-time RT-PCR and FAT was done using the kappa statistic. 4 The kappa coefficient was calculated using the formula k = (P 0 — P C )/(1 — P C ) where P 0 is the observed proportional agreement and P C is the proportion of agreement expected by chance. The strength of agreement for the kappa coefficient was interpreted using the scale of Landis and Koch 9 : ≤0 = poor, 0.1–0.20 = low, 0.21–0.40 = fair, 0.41–0.60 = moderate, 0.61–0.80 = substantial, and 0.81–1 = almost perfect.
All BRSV field strains and BRSV vaccine strains were amplified successfully by the real-time RT-PCR kit. No cross-reaction was shown with the 14 heterologous bovine viruses associated with bovine respiratory disease. The slope of the curve generated by dilution series of the BRSV strain B35.1 15 (104 TCID50/ml) was −3.34 and a strong linear relationship was demonstrated (R 2 = 0.99; Fig. 1). The efficiency of the real-time RT-PCR calculated from the slope and the formula described previously was 99.3%. This means that, on average, the number of DNA copies increased by 99.3% at every cycle. Amplification curves with increasing Ct values were achieved for 5 steps of the 10-fold dilution series of the BRSV strain B35.1 15 (104 TCID50/ml; Fig. 1). Consequently, the obtained detection limit was 0.1 TCID50. The reproducibility of the kit was high with CVs ranged between 2.61% and 4.92% (Table 1).
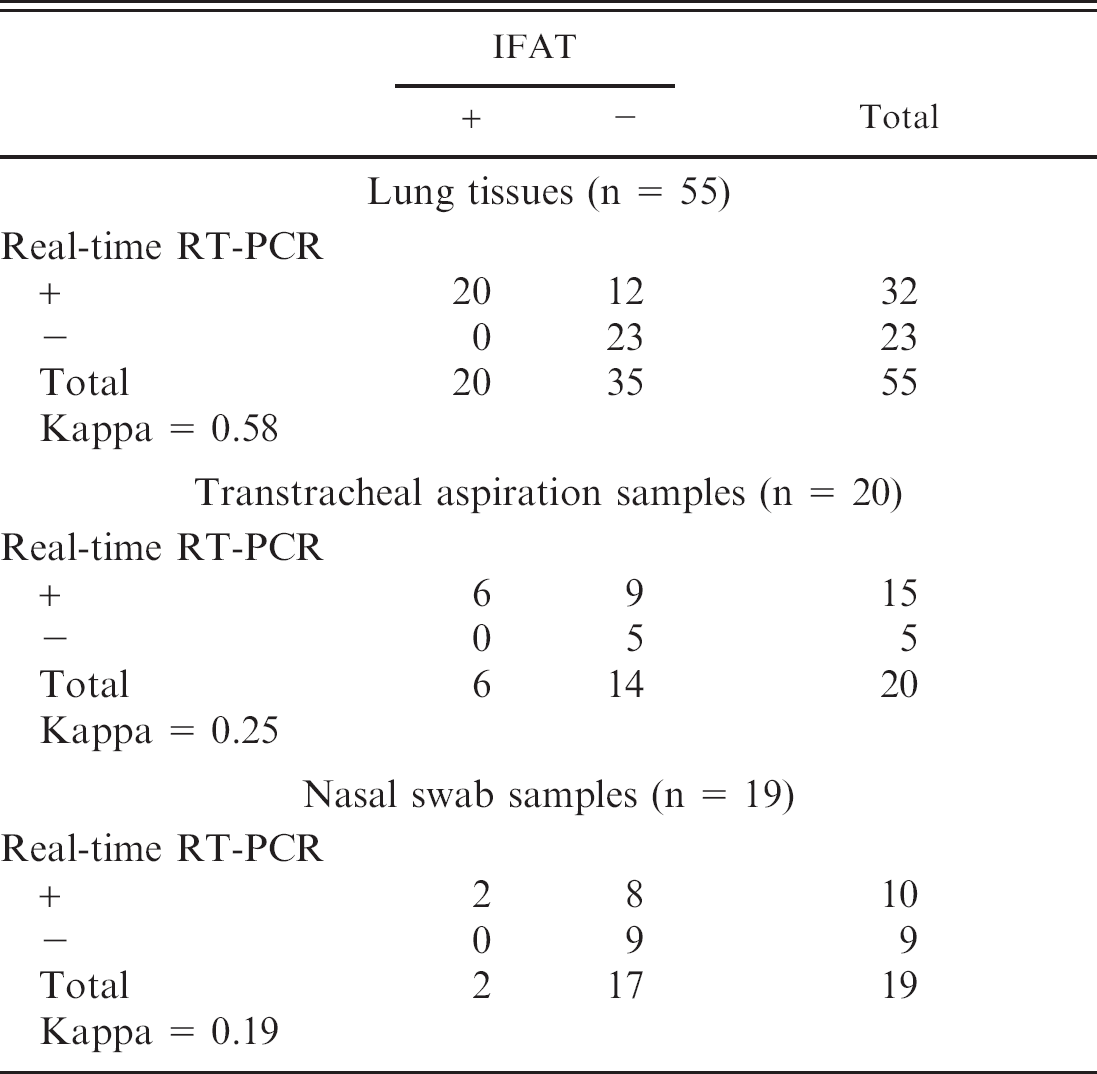
Of the 55 LT, 20 TTA, and 19 NS tested 32, 15, and 10, respectively, were positive for BRSV by real-time RT-PCR (Ct range: 19–43), whereas 20, 6, and 2, respectively, were positive for BRSV by FAT (Table 2). Internal positive control was amplified in all real-time RT-PCR assays performed confirming that negative results were not due to failure of RNA isolation or RT-PCR inhibition. Negative control included in each run gave no amplification signal suggesting that cross-contamination was avoided. The results of real-time RT-PCR and FAT were concordant for 65 of the 94 clinical samples (Table 2). Of the 29 test results that differed between the 2 techniques, all were realtime RT-PCR positive and FAT negative. Most of the samples positive by real-time RT-PCR and negative by FAT were found in the higher Ct range, which correlates with lower viral loads, demonstrating the higher sensitivity of the real-time RT-PCR (Fig. 2). A sequencing of RT-PCR products of 2 samples positive by real-time RT-PCR and negative by FAT confirmed that the RT-PCR products were part of the BRSV nucleoprotein gene (data not shown). The level of agreement between real-time RT-PCR and FAT depends on the type of clinical samples performed. The level of agreement between real-time RT-PCR and FAT on LT was moderate (kappa = 0.58), whereas it was fair (kappa = 0.25) and low (kappa = 0.19) on TTA and NS, respectively (Table 2).
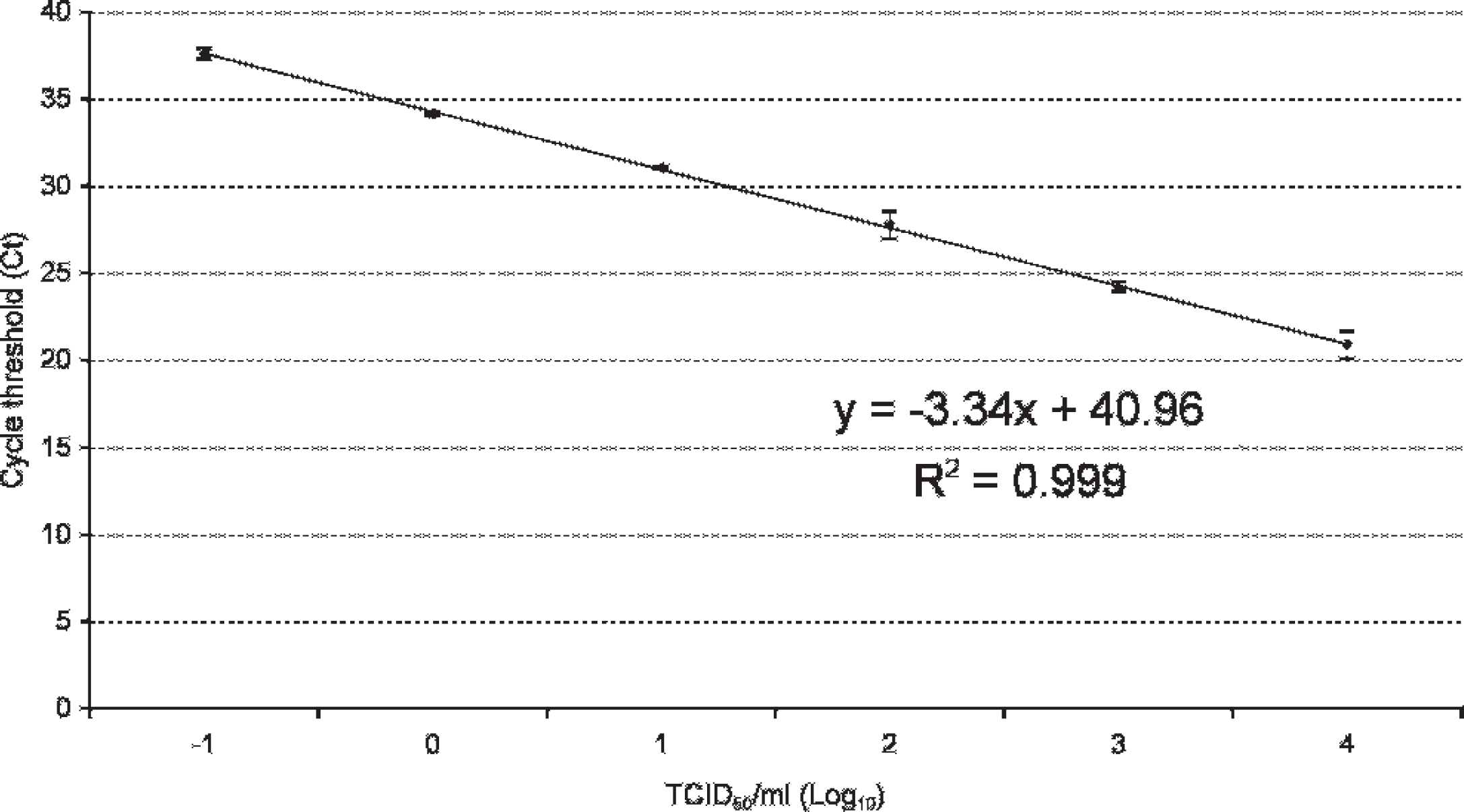
Real-time reverse transcription polymerase chain reaction standard curve on 10-fold dilutions of a titrated Bovine respiratory syncytial virus strain B35.1 15 containing 104 TCID50 (50% tissue culture infective dose)/ml. In the figure, the mean values with standard deviations are shown, as well as equations (y) and correlation coefficients (R 2 ).
Many commercial kits are used in the field without assessment of their performance. However, without such assessment, the results obtained cannot be validly interpreted in terms of diagnosis. 20 The commercial real-time RT-PCR kit used in the current study exhibited a high sensitivity and specificity, a low threshold of detection, as well as reproducible results. All of the BRSV strains tested, including strains from different subgroups of BRSV (II, III, and IV), were detected by the kit. This large spectrum of detection is explained by the high conservation of the nucleoprotein gene, 15 which is targeted by the primers and probe of the kit. Because the real-time RT-PCR product is detected with a sequence-specific probe, there is confirmation that the correct target was amplified, reducing the chances for false positives. Indeed, none of the 14 heterologous bovine viruses associated with bovine respiratory disease were detected, attesting to the high specificity of this assay. The detection limit obtained in the present study was 0.1 TCID50, which is comparable to the detection limit of previously reported real-time RT-PCR using the same primers and probe, 3 but 10 and 100 times more sensitive than real-time RT-PCR targeting the fusion gene 7 and conventional RT-PCR. 11 The reproducibility of the assay estimated on 10 runs was high with CVs lower than 5%.
Reproducibility of the real-time reverse transcription polymerase chain reaction kit examined by running 10 times 4 steps of a 10-fold dilution series of the external positive control. *
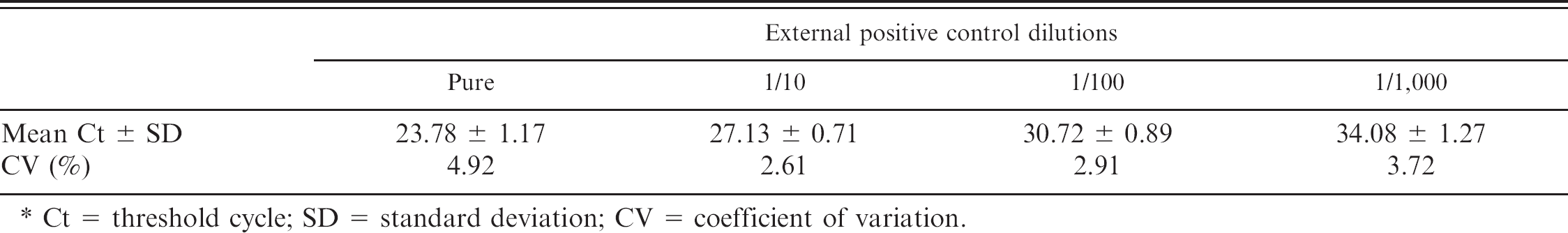
Ct = threshold cycle; SD = standard deviation; CV = coefficient of variation.
Agreement between real-time reverse transcription polymerase chain reaction (RT-PCR) and indirect fluorescent antibody test (IFAT) for the detection of Bovine respiratory syncytial virus on lung tissues, transtracheal aspiration samples, and nasal swab samples.
The higher sensitivity of the RT-PCR assay compared with FAT for the detection of BRSV in clinical sample is in agreement with the results of previous studies. 14,18 Polymer-ase chain reaction requires only minimal amounts of virus compared with antigen detection and provides, therefore, a more sensitive diagnostic approach. 11 In this study, the difference of sensitivity between the real-time RT-PCR and FAT depended on the type of clinical samples. This difference was more pronounced on NS and TTA than on LT. This discrepancy in sensitivity could be explained by an insufficient quantity of infected cells on NS and TTA samples for BRSV detection by FAT. However, cell counts performed on NS and TTA samples ensured that the number of cells was sufficient for an analysis by FAT. Another explanation could be the differences of viral loads between the different types of clinical samples obtained in this study: LT having lower Ct values (high viral titers) than NS and TTA (data not shown).
In conclusion, based on these results, the commercial realtime RT-PCR kit evaluated in the current study can be reliably used to detect BRSV in clinical samples. This kit displayed analytical features necessary in routine laboratory diagnosis: a high sensitivity and specificity, a low threshold of detection, as well as reproducible results. Furthermore, the use of this kit instead of FAT allows an improvement of the sensitivity for the detection of BRSV in clinical samples.
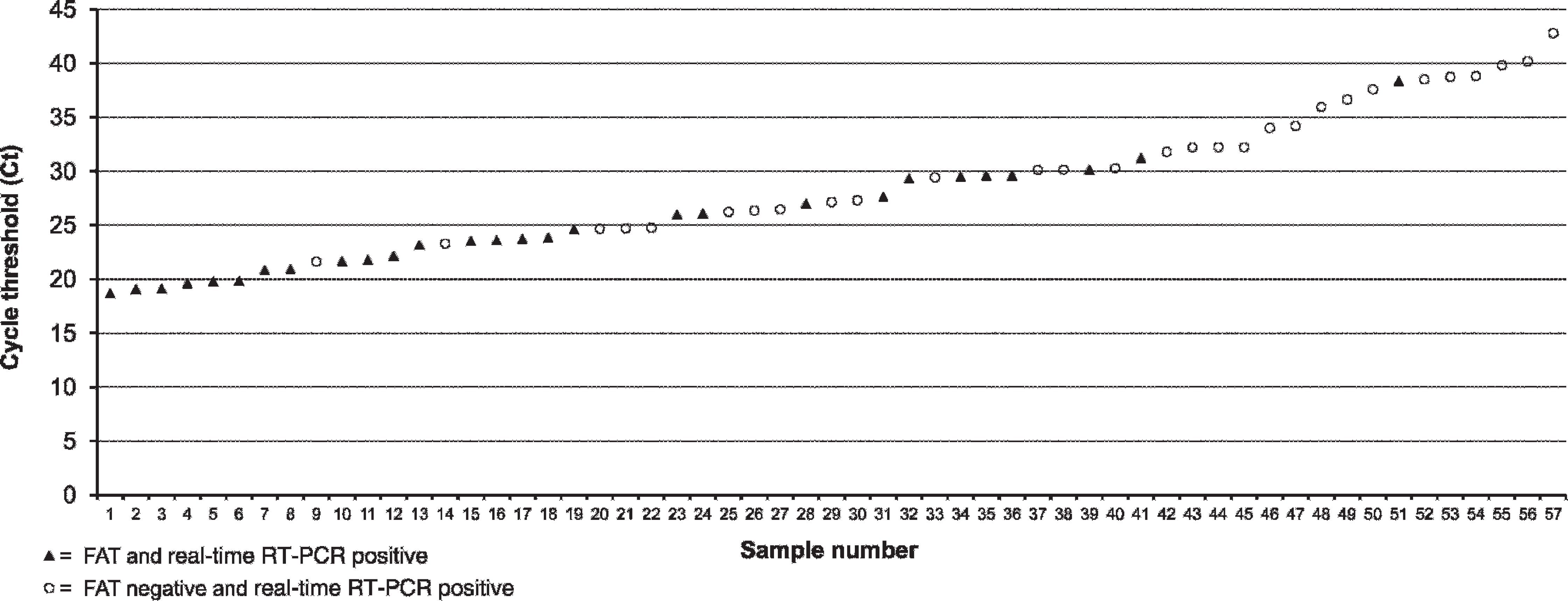
Fluorescent antibody test (FAT) results for Bovine respiratory syncytial virus (BRSV) antigen detection for all 57 clinical samples positive for BRSV by real-time reverse transcription polymerase chain reaction (RT-PCR) arranged in order of increasing threshold cycle.
Footnotes
a.
Kit Taqvet® BRSV, L.S.I., Lissieu, France.
b.
Rispoval RS®, Pfizer, Paris, France.
c.
Rispoval RS + Pi3 IntraNasal®, Pfizer, Paris, France.
d.
Copan Italia, Bresca, Italy.
e.
QIAamp® Viral RNA kit, Qiagen GmbH, Hilden, Germany.
f.
ABI PRISM 7000 Sequence Detection System Real-Time apparatus®, Applied Biosystems, Foster City, CA.
g.
Argene Biosoft, Varilhes, France.
