Abstract
Introduction
The recent European outbreaks of bluetongue disease in cattle and sheep have demonstrated the economic impact of this disease.
4
In a 1989 estimate, the annual loss to the livestock industry from this disease in the United States and worldwide was significant (Bath GF: 1989, Bluetongue.
An understanding of the molecular genetics of these viruses is needed for the development of rapid and sensitive molecular diagnostic assays. The protein coding assignments for BTV and EHDV are identical, consisting of 10 genome segments that encode 7 structural and 4 nonstructural proteins. 20 Developing molecular diagnostics is complicated by the multiple serotypes of these viruses: 24 for BTV and 8 for EHDV. 19 The outer capsid protein, VP2, is the primary serotype determinant, but the second outer capsid protein VP5 can also influence serotypic epitopes. 21,25,40 The structural genes that are fairly conserved within the virus serogroup encode the inner core proteins VP3 and VP7, 23,42 and this information has been used to develop reverse transcription polymerase chain reaction (RT-PCR) assays. 2 The VP7 protein is commonly used as the antigen for enzyme-linked immunosorbent assays. 24
The nonstructural (NS) proteins are highly conserved among North American orbiviruses. 3,27,28 The conservation of these viral proteins is likely because of their various roles in viral replication. The NS1 protein produces tubules in infected cells 37 and is thought to be important in virus assembly. Binding of single-stranded RNA has been demonstrated with the BTV NS2 protein. 8 Viral egress has been shown to be facilitated by the BTV NS3 protein. 12
A number of real-time RT-PCR assays were recently developed for BTV.
7,14,29,32,36
The genome segments are commonly denoted as L1–3, M4–6, and S7–10, based on sequence size, similar to the
Materials and methods
Prototype viruses
EHDV-1 (New Jersey), EHDV-2 (Alberta), BTV-2 (Ona B), BTV-10 (BT-8), BTV-11 (Texas Station Strain), BTV-13 (67-41B), and BTV-17 (63-66B) were obtained from the Arthropod-Borne Animal Diseases Research Laboratory (ABADRL) virus collection. Prototype strains of BTV-1, 3–9, 12, 14–16, 18, and 19 previously provided by the Onderstepoort Veterinary Institute (OVI; Pretoria, Republic of South Africa) were obtained from the ABADRL virus collection. National Veterinary Services Laboratories (NVSL; Ames, IA) provided prototype strains of BTV-20–24 (originally from OVI). Prototype strains of EHDV-3–8 were also kindly provided by the NVSL. EHDV types 3 and 4 were isolated in Nigeria and provided to NVSL by the Institute for Animal Health (Pirbright, UK). The Australian Animal Health Laboratory originally provided prototype strains of EHDV-5 (CSIRO 157), EHDV-6 (CS753), EHDV-7 (CS775), and EHDV-8 (DPP059) to NVSL. The prototype viruses were propagated in baby hamster kidney (BHK)-21 or African green monkey kidney epithelial (Vero Middle America Research Unit) cell cultures. Cells were infected by using 0.01 multiplicity of infection, and RNA extractions were performed when approximately 80% of the infected cells showed cytopathology.
RNA extraction from infected cells or blood
Total RNA was extracted from cells by using a commercial RNA isolation kit
a
as per the manufacturer's protocol, with the following exceptions: for cell lysis, the initial centrifugation was carried out for 5 min at 15,000 ×
High-throughput RNA extractions from blood were compared by using 2 different magnetic-bead capture kits: MagMAX™-96 for Microarrays Total RNA Isolation c (hereafter, MagMAX-96 kit) and MagMAX™ Viral RNA Isolation c (hereafter, MagMAX kit). Blood samples from sheep were spiked with BTV-17 and EHDV-2 with 10-fold dilutions from 106 to 1 CCID50/ml ([50% cell culture infective dose]/ml). The MagMAX kit was slightly modified. Briefly, 20 μl of bead mix was added to 100 μl of blood and mixed by shaking in a 96-well plate. Lysis-binding solution (400 μl) was then added, and the mixture was shaken for 5 min, after which it was centrifuged for 5 min, and the supernatant was discarded. The original protocol was followed for the wash steps; however, elution was done by using 100 μl of 0.1 mM ethylenediamine tetra-acetic acid (EDTA) at 65°C, instead of the kit elution solution. The MagMAX-96 kit used 200 μl of blood starting material, mixed with 750 μl TRI reagent, c followed by a spin procedure and elution in 100 μl of elution buffer. RNA was quantitated as stated previously.
M5 and S10 cloning and sequence analysis
Amplification of the M5 and S10 genes was performed as described previously, with primer modifications for the M5 gene. 3,13 The amplification primers used for the S10 gene were the following: BTV S10+1D (GTTAAAAAGTG TCGCTGYCA) and BTV S10-822D (GTAAGTGTRTA GYRYCGC) or BTV S10-822ND (GTAAGTGTG TAGGGCCGC). The amplification primers for the M5 gene were the following: BTV10NS1-N1–5 (atccgatccGT TAAAAAAGTTCTCT) and BTV10NS1-N1–3 (atccggatccGTAAGTTGAAAAGTT; lower case bases are not complementary to the gene). The amplicons were then cloned into the TOPO (topoisomerase I) 2.1 vector d and transformed into TOP10 F' chemically competent cells. d Sequencing was performed by using standard automated protocols e on at least 3 individual clones, and on PCR products to resolve any discrepancies in consensus sequences. SeqMan software f was used for sequence assembly. Viral gene sequences, minus the primer sequences (except for sequences in the coding region of S10), were aligned by using either ClustalX 34 or MUSCLE 6 with default settings. Phylogenetic trees were calculated from these multiple sequence alignments by using MrBayes, 31 with the standard nucleotide substitution model (4 × 4) with GTR (nst = 6). Substitution rates were set to invgamma (gamma-shaped rate variation with a proportion of invariable sites); default values were used for all other settings. A total of 750,000 generations were calculated by sampling every 100th tree, and a consensus tree was calculated after a burn-in of 2,500 trees by using the allcompat setting. Phylogenetic trees were also generated by using ClustalW alignment based on absolute number of differences with gaps distributed proportionally, and Unweighted Pair Group Method with Arithmetic mean (UPGMA; bootstrap [1,000 reps]; tie breaking = systematic). g
BTV primer-probe sets or signatures
A panel of signatures (set of primers and probe with specificity for a distinct target sequence) was generated by using bioinformatics as previously described 16 on published sequence data and data generated in the current study for gene segments L1, M5, S8 (NS2), and S10. These signatures were prescreened in silico and further screened to demonstrate no cross-reactivity with more than 2,500 potential background samples, as described previously. 16 The remaining signatures were then further evaluated for their ability to detect BTV strains but not the closely related EHDV RNA.
Analysis of new BTV signatures for real-time RT-PCR
All 24 BTV serotypes were run against all signatures individually by using a commercial real-time PCR detection system. h The real-time RT-PCR procedure was performed as described previously. 16 Cycle threshold (Ct) values were recorded for each signature. Of the signatures that amplified all 24 BTV serotypes, a limit of detection (LOD) was then determined for each signature on each serotype. For LOD, samples were run in triplicate with 10-fold dilutions from 2 pg to 2 fg of dsRNA, and Ct values were recorded and averaged. The signatures that detected all serotypes were also tested on prototype EHDV 1–8 serotypes to validate lack of cross-reactivity. The selected signatures were then used to detect RNA (200-pg run in duplicate) from 45 random BTV strains and 60 BTV- and EHDV-suspect clinical samples from the authors' inventory.
Real-time RT-PCR
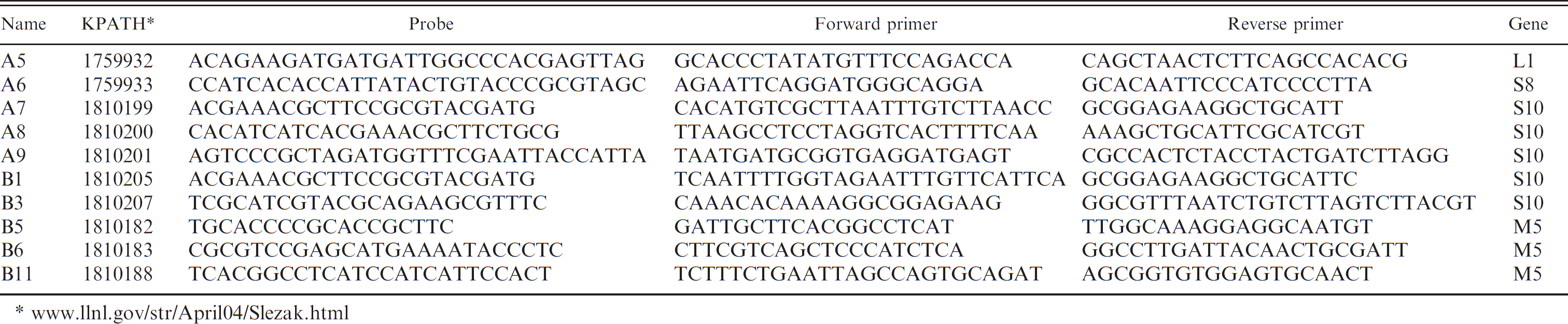
Initial experiments were conducted with only the BTV signatures generated in the current study. Final analysis was done after combining the BTV signatures with the previously published EHDV signatures 43 (Table 1) by using a multiplex format. Two different quencher and/or reporters were used to differentiate between BTV and EHDV. The quencher and/or reporter choices varied depending on the filter sets of the instrument available at the time of the experiment. Initial experiments on RNA from cell cultures were conducted with 3 different thermocyclers e,h,i by using the TaqMan EZ RT-PCR core reagents e and the cycling parameters based on the authors' previous EHDV real-time PCR assay 43 of 55°C for 25 min, 95°C for 2 min, followed by 40 cycles of 95°C for 10 sec and 55°C for 1 min. The BTV probes used with the Bio-Rad iCycler h was dual labeled with fluorophore 6-carboxyfluorescein (FAM) and Black Hole Quencher (BHQ)-1 j during the initial primer evaluation studies. In the multiplex assay development, when the Applied Biosystems SDS7000 was used, the probes were dual labeled with 5′ CAL Fluor Orange 560 and BHQ-1. j The BTV probes for use with the Cepheid SmartCycler II i was dual labeled with 5′ CAL Fluor Gold 540 and BHQ-1. j The EHDV probes were dual labeled with FAM and BHQ-1. j Two real-time RT-PCR protocols were compared by using 2 different kits (AgPath-ID kit c and TaqMan EZ e ) and 2 different cycling parameters on the Applied Biosystem SDS7000 real-time PCR instrument. e For both kits, 8 μl of template was used from 100 μl of the blood extraction procedure. The template was denatured at 95°C for 8 min, and 2 cycling parameters were compared. In some cases noted in the text, the template was heated to 95°C for 3 min with 10% dimethyl sulfoxide, as described previously. 36 The first set of parameters was Applied Biosystems's thermal profile parameters (manufacturer recommendation): 1 cycle of 48°C for 10 min and 95°C for 10 min, followed by 40 cycles of 95°C for 15 sec and 60°C for 1 min. The second set of parameters was based on the authors' previous EHDV real-time PCR assay 43 as stated above.
Results
Sequence and phylogenetic analysis
The M5 (NS1) and S10 (NS3) genes from BTV prototype strains, where sequence information was not available, were amplified by using terminal primers and the sequences were determined. The primer sequences, except for the first 5 bases on the 5′ end that included the ATG start codon for the S10, were removed for data analysis. The 40 new sequences generated were reported to GenBank (accession nos. FJ713316–FJ713355). Unfortunately, the origins of these prototype viruses were not able to be confirmed. The BTV M5 gene sequences had a percent identity that ranged from 78.7% to 99.7%. The percent identity range between BTV and EHDV M5 gene sequences was 56.3–58.4%. The BTV S10 gene sequences had a percent identity range from 81.5% to 99.9%. The percent identity range between BTV and EHDV S10 gene sequences was 58.3–61.8%. The BTV M5 sequences separated into 2 major lineages, with a very high degree of confidence. Phylogenetic analyses of M5 and S10 sequences were performed by using several algorithms, and all showed essentially the same profiles (Figs. 1, 2).
BTV primer and/or probe (PCR signatures) design
The sequence data from the current study and previous publications were used for BTV signature generation based on 27 sequences for the M5 gene and 52 sequences for the S10 gene (unpublished data). An in silico analysis, by using a previously described alogorithm, 16 was used to identify multiple candidate signatures of highly conserved genome regions. This analysis provided 11 PCR signatures for M5 and 16 PCR signatures for S10 that were discriminatory by the in silico analysis. These signatures were tested against 50 soil, 60 prokaryotes, 21 eukaryotes, 2,000 aerosol samples, and 4 U.S. strains of EHDV. The specificity testing by gel RT-PCR screening, followed by real-time RT-PCR assay screening against these potential background samples, resulted in 8 potential segment M5 signatures and 10 potential segment S10 signatures. Preliminary target detection with 5 domestic serotypes by using known amounts of BTV RNA eliminated 4 of the signatures. Further screening on the remaining seven M5 and seven S10 signatures resulted in three M5 and five S10 signatures that were further screened against all 24 BTV serotypes. The specificity of these signatures, along with previously designed L1 and S8 signatures, is shown in Table 1.
The 4 signatures that detected all 24 BTV serotypes, at 200 pg dsRNA, were A7, A8, B1, and B3, which all targeted the S10 gene (Table 2). The LOD was determined for each of these 4 signatures, and Ct values were recorded and sensitivity analyzed (Table 3). Although the linearity between Ct and dilution is not always consistent, the general trend was and the results were consistent with the experimental error of real-time RT-PCR assays around the LOD (data not shown). In some cases, there was no amplification of specific targets that were amplified in the initial screening. This was likely because of experimental error around the LOD for the target. The signatures A7, B1, and B3 were selected for future inclusion in the multiplex vesicular disease panel. 16 The intent was to use separate signatures in the real-time RT-PCR and the multiplex vesicular disease panel, 10 so that the real-time RT-PCR assay could be a confirmatory test based on an independent target sequence. Therefore, the remaining signature, A8 was used for the real-time RT-PCR BTV assay. After further analysis of inventoried strains, some of the strains were not detected when using A8, notably, BTV-2 Ona A. The other signatures that did not detect all serotypes were tested again to determine if they were able to detect the BTV-2 Ona A strain. One of the 6 remaining signatures (B5 from M5) was able to detect BTV-2 Ona A. The BTV real-time RT-PCR assay thus consists of 2 signatures (A8 and B5), or 2 probes and 4 primers, and detected all 24 BTV prototype serotype strains. In addition, it detected 45 random BTV strains and 4 BTV virus isolation positive clinical samples from our inventory.
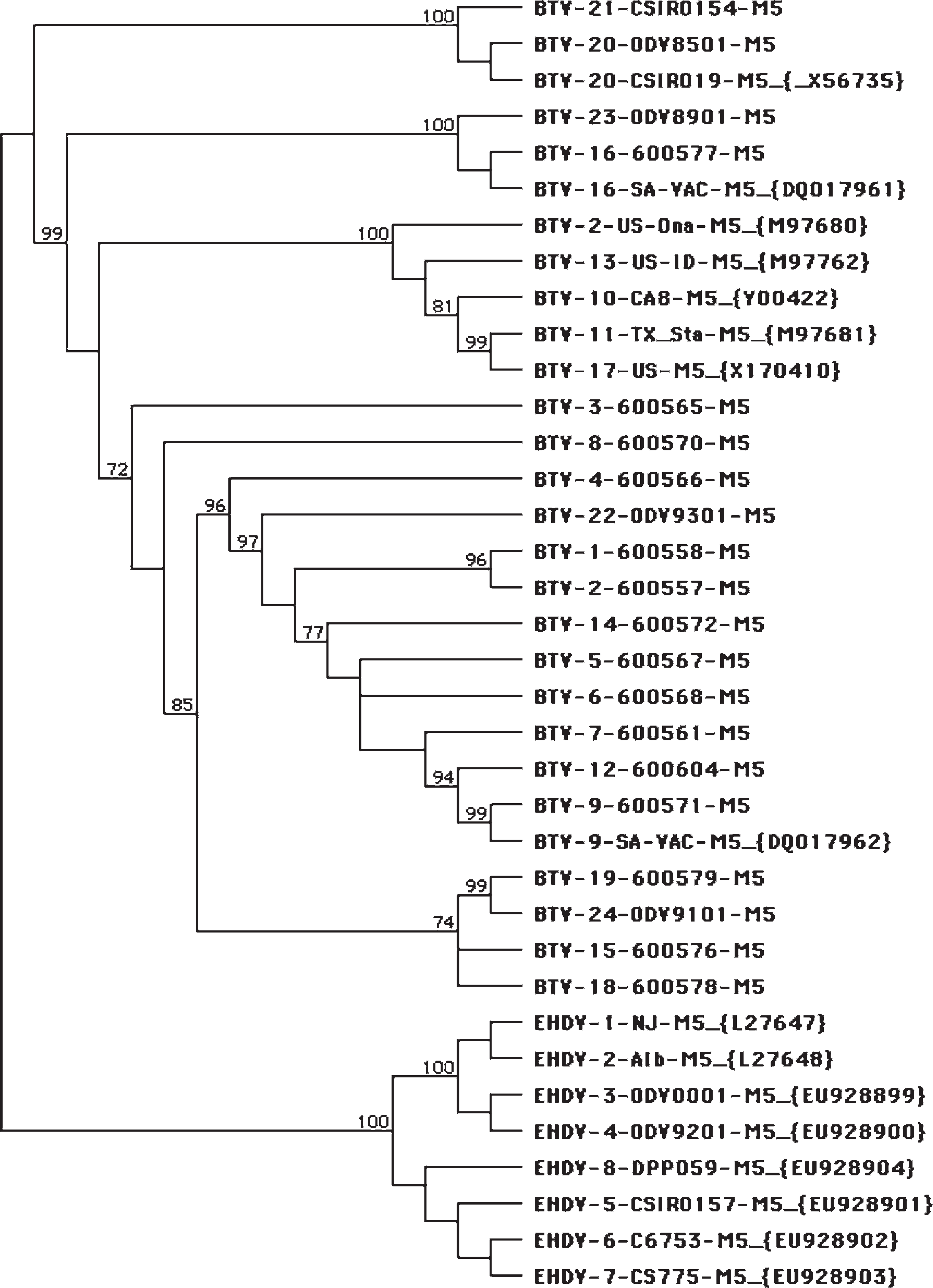
Example of phylogenetic analysis of
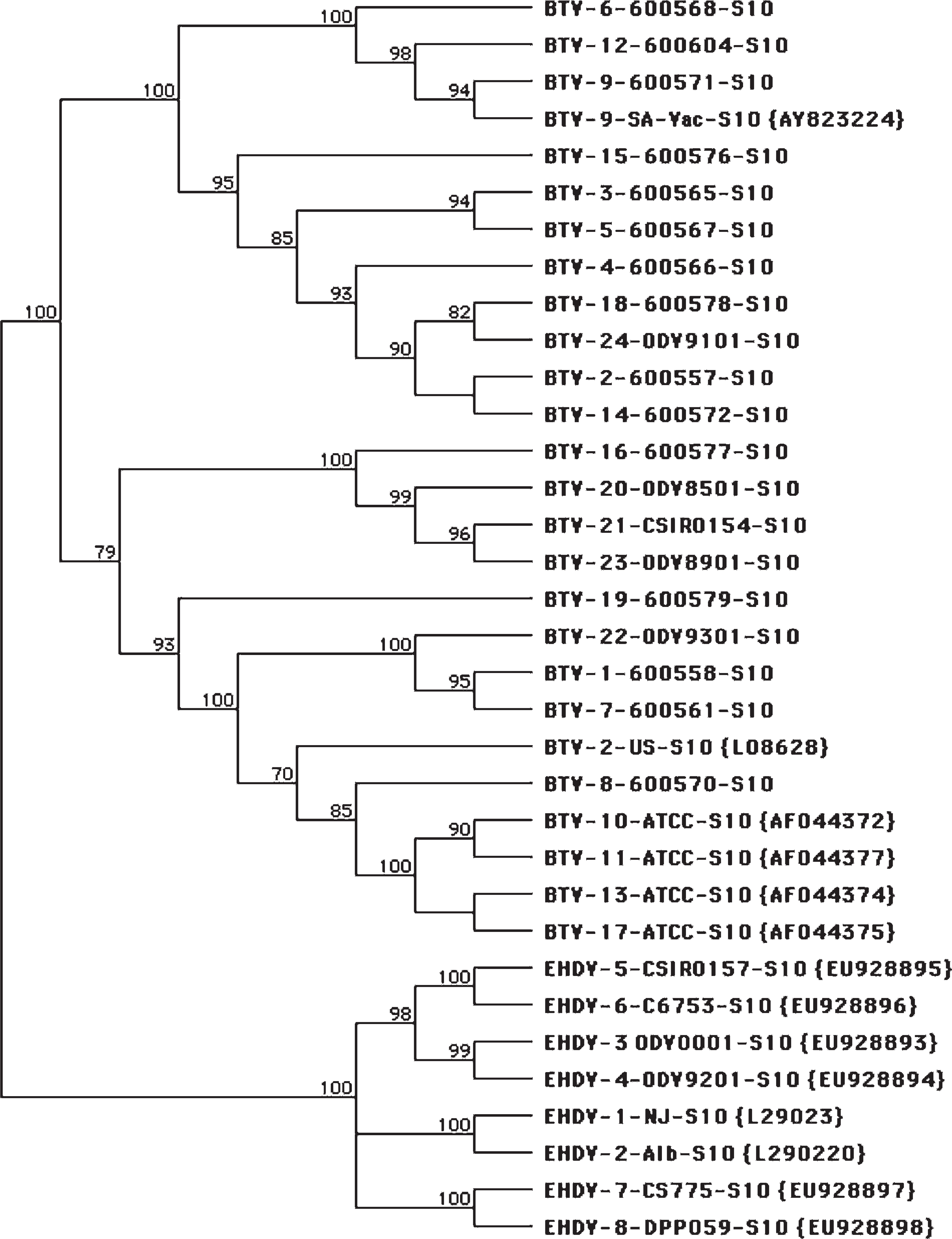

Example of phylogenetic analysis of
Threshold cycle values from 10 signatures tested against 24
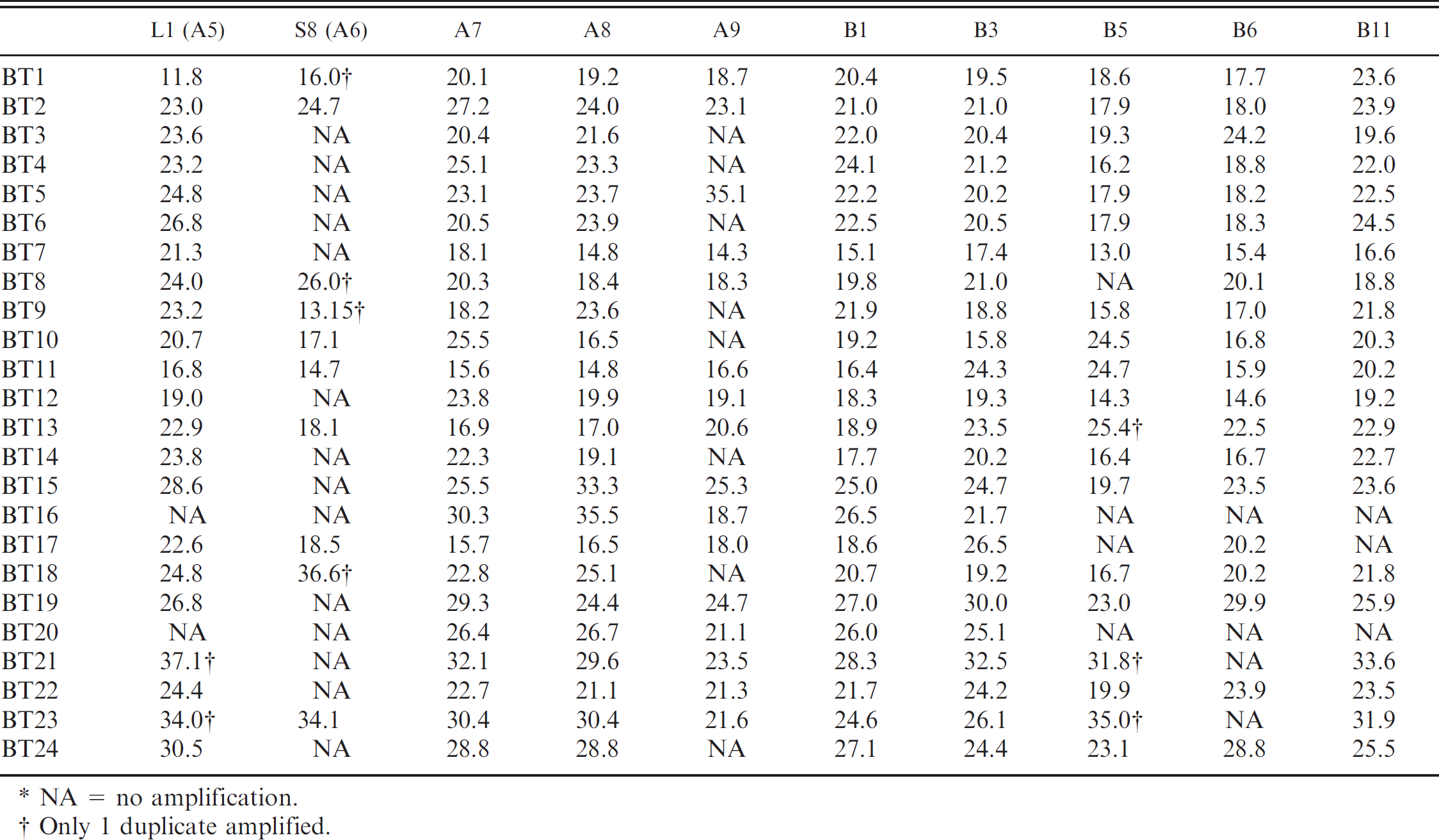
NA = no amplification.
Only 1 duplicate amplified.
BTV-EHDV multiplex real-time RT-PCR
In the final design, the BTV A8 and B5 primers and respective probes were combined with the previously published EHDV real-time RT-PCR assay. 43 The probes for BTV and EHDV contained different quencher and reporter dyes. The BTV probes contained CAL Fluor Orange 560 j on the 5′-end and BHQ-1 j on the 3′ end. The EHDV probes had FAM on the 5′ end and BHQ-1 on the 3′ end. Initial tests with this design detected and differentiated cell culture source BTV and EHDV RNA; however, the assay lacked sensitivity with clinical samples. To address the sensitivity, heat or heat plus dimethyl sulfoxide denaturation of the dsRNA target was evaluated. The assay was able to detect 1,000 copies of BTV-10, BTV-17, EHDV-1, and EHDV-2, with Ct values in the mid 20 range. At 100 copies of EHDV-1 or -2, a slightly, but not significantly lower, Ct value was found when denaturing the template alone before reagent addition. The BTV-10 and BTV-17 RNA were not detected at 100 copies. To further increase the sensitivity, various commercial RNA extraction and real-time RT-PCR reagents were evaluated. This evaluation was done by using sheep blood spiked with known quantities of BTV or EHDV. Of the blood extraction kits, the MagMAX Viral RNA Isolation kit c used with Applied Biosystem cycling parameters was more sensitive, with a LOD of 10 CCID50/ml for BTV and 1 CCID50/ml for EHDV in comparison with the original EHDV parameters. This extraction kit was also easy to use, because it did not require phenol extraction or individual tubes when processing high-volume samples. The AgPath-ID kit c provided greater sensitivity for real-time RT-PCR and uses fewer reagents, so it is a faster option, with less chance of error. A comparison of the original assay reaction, consisting of the TaqMan EZ e versus Ag-Path ID, c indicated that the Ag-Path ID reagents had a 1,000-fold increase in sensitivity for both BTV and EHDV RNA detection (Table 4).
Discussion
Exotic serotypes of BTV continue to be introduced into countries where there are naïve livestock. This has been a significant issue in Europe with the introduction of BTV into southern Europe,
17
and, more recently, the introduction of BTV-8 in northern Europe, which is causing significant disease in cattle.
17
The introduction of exotic BTV or EHDV serotypes, and the potential effect on livestock health and trade, is a concern to the U.S. livestock industry.
The limit of detection (LOD) was determined for signatures A7, A8, B1, and B3 by running each serotype in at least triplicates from 2 pg to 2 fg.*
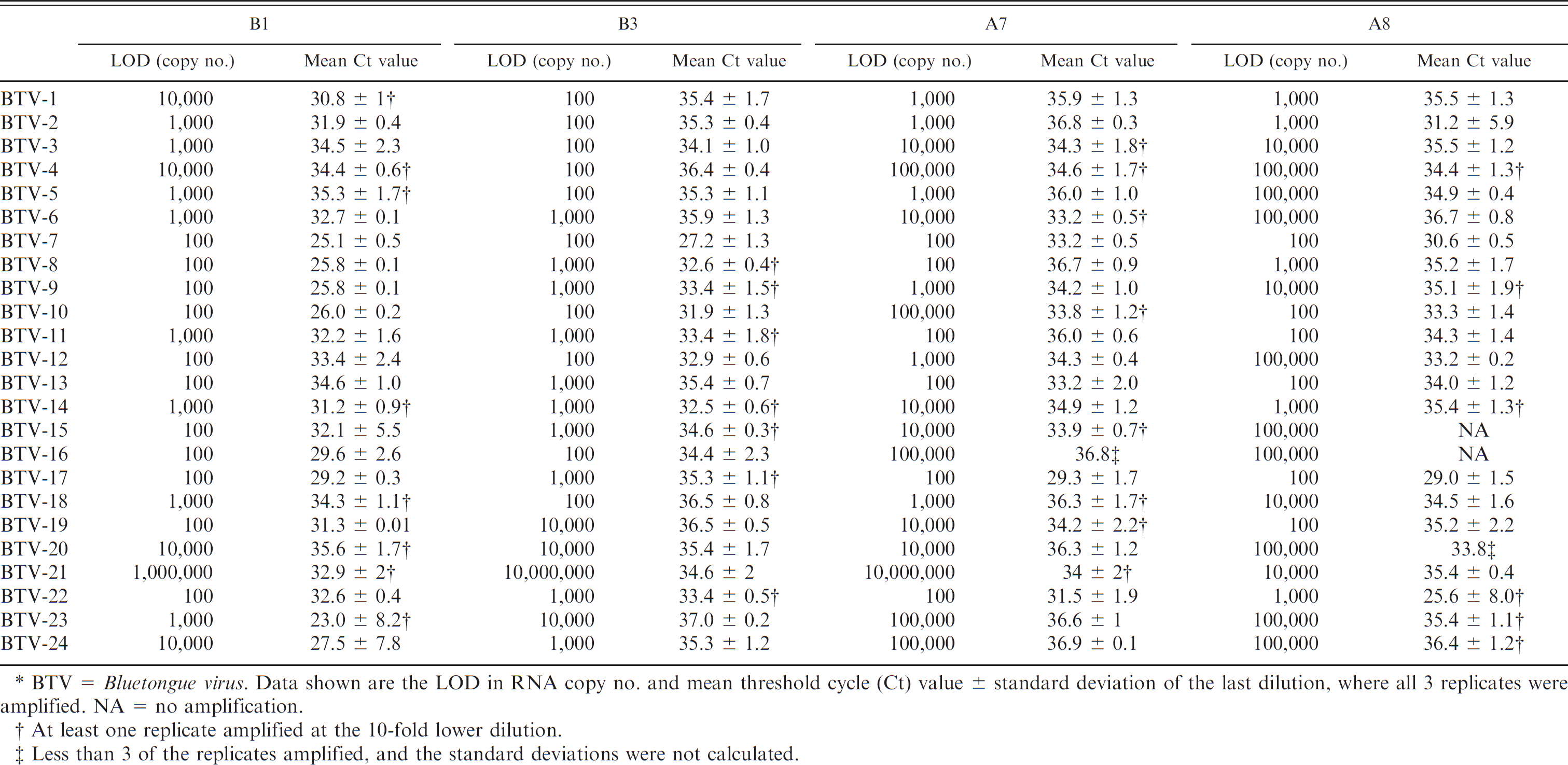
BTV =
At least one replicate amplified at the 10-fold lower dilution.
Less than 3 of the replicates amplified, and the standard deviations were not calculated.
Comparison of the TaqMan EZ RT-PCR core reagents e and the AgPath-ID One-Step RT-PCR reagents c paired with the MagMAX Viral RNA Isolation. c *
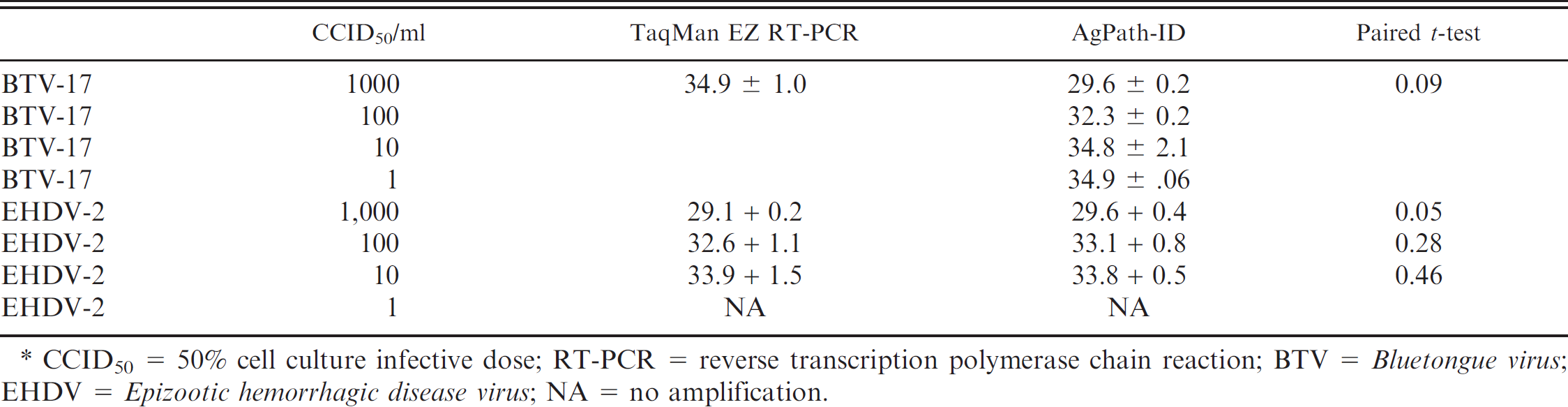
CCID50 = 50% cell culture infective dose; RT-PCR = reverse transcription polymerase chain reaction; BTV =
The sequence analysis performed in this study was done to assist in primer design and not for use in a comprehensive phylogenic analysis; however, sophisticated analysis was performed. The prototype viruses used in the current study were provided by or received by previous investigators, and their origins were unable to be confirmed. It was presumed that BTV prototype serotypes 1–9, 12, 14–16, 18, 19, 23, and 24 are of African, primarily southern African, origin. Serotypes 20 and 21 likely originated from Australia. These assumptions fit the phylogenetic analysis in that viruses tend to group according to geographic origin. This tendency has previously been shown for the S10 gene in a more comprehensive phylogenetic study. 3 A similar tendency for geographic topotypes was demonstrated for the S3 and S7 genes as well. 30,42 It is unfortunate that the origins of these viruses could not be confirmed, but the data suggest that the M5 gene may also display a tendency for geographical topotypes. The current approach for BTV and EHDV phylogenetic analysis is to use whole genome sequencing approaches. 18 This technology will rapidly advance our understanding of the molecular epidemiology of these viruses.
The sequence and phylogenetic analyses reported here demonstrated greater variability of the 2 target nonstructural genes than was shown in previous studies. 39 Understanding genetic diversity of potential viral genome targets is essential in designing more robust real-time RT-PCR assays that rapidly detect both domestic and exotic BTV and EHDV strains. The multiplex BTV and/or EHDV real-time RT-PCR assay reported in this study incorporates multiple RNA targets, thus it is less likely to be adversely affected by genetic variation than previous real-time RT-PCR assays for BTV or EHDV. Use of a sophisticated bioinformatic design approach has allowed the development of an assay that detects an estimated 10 CCID50/ml for BTV and 1 CCID50/ml for EHDV in spiked blood. A future improvement of this assay would be to include an internal or positive RNA control as in the multiplex vesicular disease panel. 16 However, to do so at this time would significantly delay release of the assay. The current assay is advantageous over previously reported assays, because it is the first reported real-time RT-PCR assay to detect and distinguish both BTV and EHDV in a single-tube format. In fact, the presence of both BTV and EHDV RNA in a clinical sample was detected. The assay was proven to be adaptable to genetic variation discovered in field isolates; thus, it provides a convenient and robust tool to orbivirus researches as well as veterinary diagnostic laboratories.
Acknowledgements
The authors thank Donna Johnson and Dr. Eileen Ostlund from APHIS-NVSL (Ames, IA) for providing RNA from exotic strains of BTV and EHDV necessary to complete this study and Ambion Inc. for technical advice. The authors thank Dr. Barbara Drolet, USDA, ARS, for reviewing an earlier version of this manuscript. This project was funded by USDA-ARS projects 5410-32000-015-00D and 5410-32000-018-00D. Bioinformatic analysis was supported by NIH Grant P20 RR016474 from the INBRE Program of the National Center for Research Resources. The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views of NIH or the USDA.
Footnotes
a.
Purescript®, Gentra Systems Inc., Minneapolis, MN.
b.
Implen Inc., Westlake Village, CA
c.
Ambion® MagMAX™-96 for Microarrays Total RNA Isolation, Ambion® MagMAX™ Viral RNA Isolation, Ambion® AgPath-ID™ One Step RT-PCR Reagents; Applied Biosystems, Foster City, CA.
d.
Invitrogen Inc., Carlsbad, CA.
e.
TaqMan EZ RT-PCR core reagents, Applied Biosystems, Foster City, CA.
f.
DNA Star Inc., Madison, WI.
g.
MacVector Inc., Cary, NC.
h.
MyiQ Single-Color Real-Time PCR Detection System, Bio-Rad Laboratories, Hercules, CA.
i.
Cepheid, Sunnyvale, CA.
j.
Biosearch Technologies Inc., Novato, CA.
