Abstract
Sustained transmission of canine
Keywords
Canine
Two studies were completed to assess the performance of various CIV diagnostic methods. For the first study, a total of 124 shelter and household dogs demonstrating symptoms of CIRDC were swabbed between March 2006 and April 2007 by clinicians at Colorado State University Veterinary Teaching Hospital (CSU-VTH; Fort Collins, CO) or on-site at shelters reporting CIRDC. Briefly, a sterile polyester-tipped swab was inserted into the nostril of each dog and, if possible, a second swab was collected from the pharynx of each animal. The swab samples were immediately placed in different vials of 1 ml of viral transport medium (phosphate buffered saline, 5 mg/ml of bovine serum albumin, 2,000 U/ml of potassium penicillin G, 4 mg/ml of streptomycin, 16 µg/ml of gentamicin, and 100 U/ml of nystatin) and stored at −80°C until VI, FluAB RIDT, and real-time RT-PCR assay could be performed. For the second study, 2 nasal swabs were simultaneously collected between December 2009 and November 2010 from 1,372 dogs housed in 1 of 6 different humane shelters participating in a large-scale U.S. CIV surveillance program. One swab was processed on-site according to the recommendation of the manufacturer of Flu Detect. The second nasal swab sample was stored at 4°C before being shipped to the laboratory overnight on ice for real-time RT-PCR assay. All studies were reviewed and approved for conduct by the CSU Institutional Animal Care and Use Committee prior to initiation.
Influenza viruses from swabs were isolated by inoculating both Madin–Darby canine kidney (MDCK) cells and the allantoic cavity of 10-day-old embryonated chicken eggs with 200 µl of thawed viral transport medium and incubated for 48–72 hr at 37°C. Incubation for the inoculation of MDCK cells included 5% CO2. After cell culture inoculation, 2 ml of Eagle minimum essential medium c supplemented with 2.25 mg/ml of bovine serum albumin, 50 mg/ml penicillin–streptomycin, 250 μg/ml amphotericin B, and 1 µg/ml of tolylsulfonyl phenylalanyl chloromethyl ketone–treated trypsin d were added to each well. Cell and egg cultures were passaged up to 3 times. Influenza virus was confirmed by immunocytochemistry staining with an anti–influenza A nucleoprotein antibody e and RT-PCR amplification using SZAHA primers 20 for the hemagglutinin and neuraminidase genes. The PCR products were analyzed by direct cycle sequencing f at the CSU Proteomics and Metabolomics Facility (Fort Collins, CO). Nucleotide and amino acid sequences were assembled using commercial g software.
For the 1-step real-time RT-PCR assay, primers and probes were designed using commercial
h
software and based on 9 matrix (M) gene sequences from equine and canine
Sensitivity estimates for VI, FluAB, and real-time RT-PCR were determined by a stochastic latent class modeling approach described previously
2
using Gibbs sampling.
18
The CSU-VTH study data set was divided into shelter (
Comparison of canine
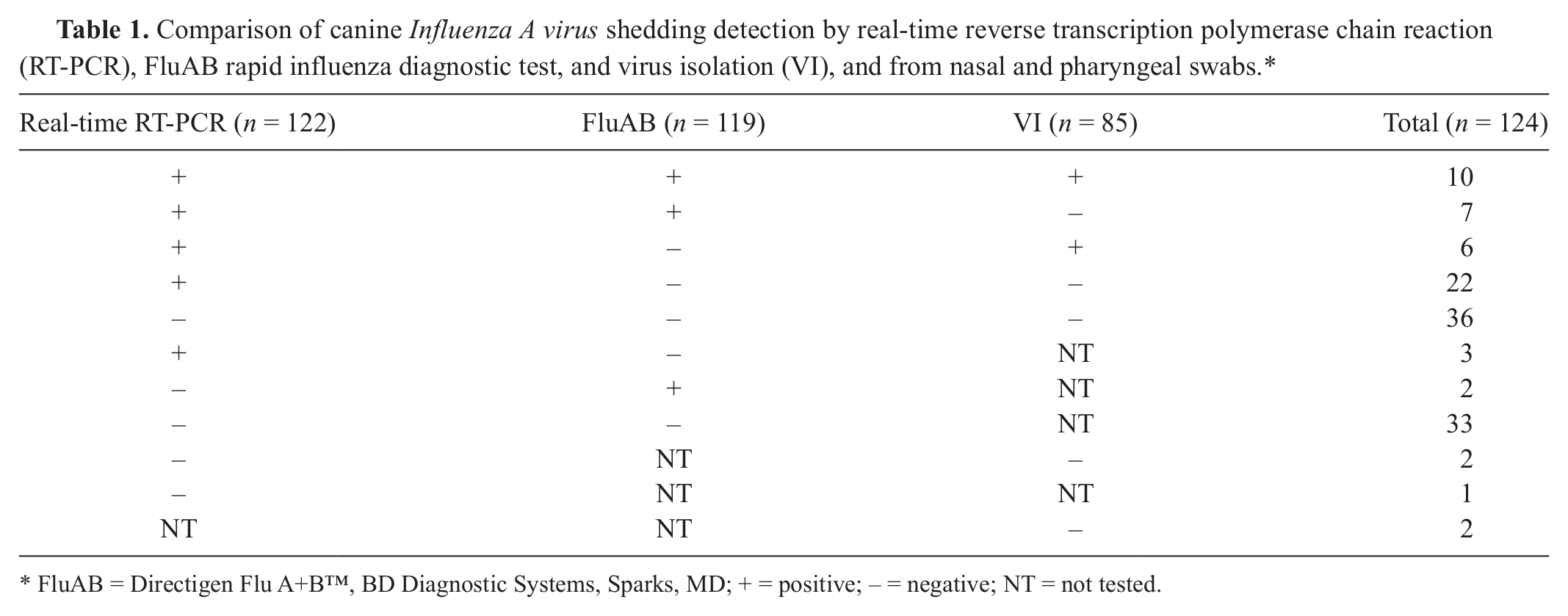

FluAB = Directigen Flu A+B™, BD Diagnostic Systems, Sparks, MD; + = positive; − = negative; NT = not tested.
Estimates, modes, 95% probability intervals (PI), and beta distributions used for prior distributions in the stochastic latent class models.
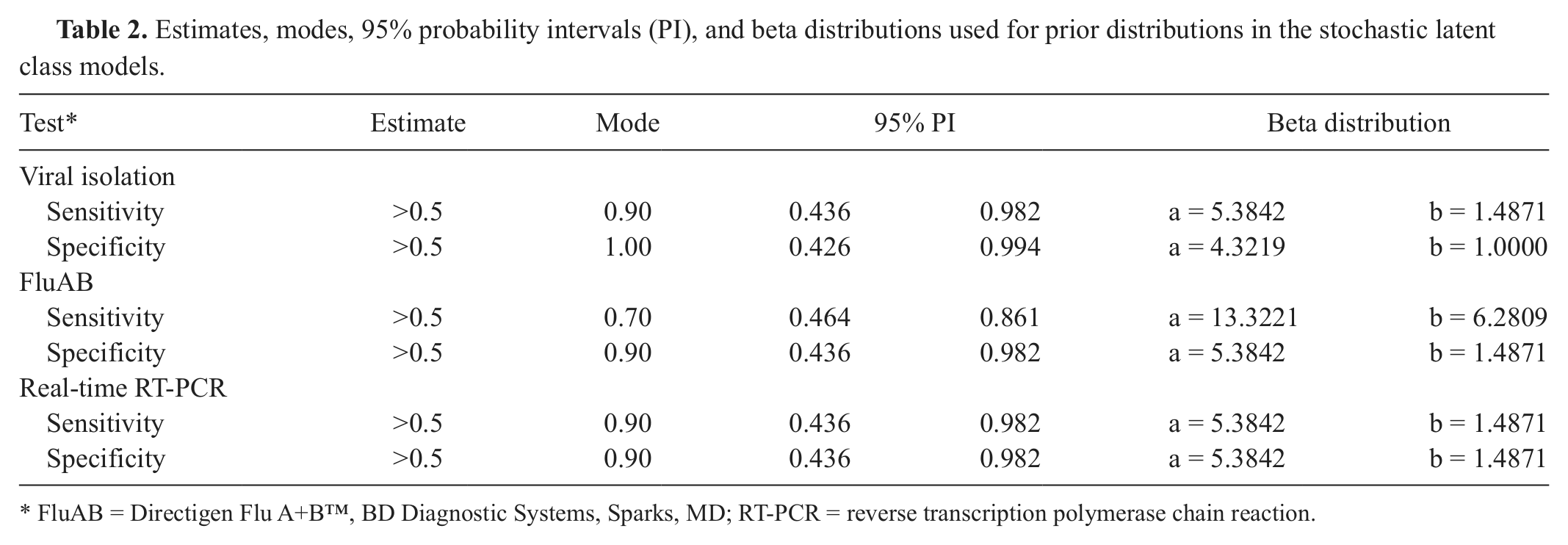
FluAB = Directigen Flu A+B™, BD Diagnostic Systems, Sparks, MD; RT-PCR = reverse transcription polymerase chain reaction.
As there was only 1 population (shelter dogs) sampled for the Flu Detect RIDT analysis, the sensitivity, specificity, positive predictive value, and negative predictive value were determined using traditional calculations from a 2 × 2 contingency table. Of the 1,372 shelter dog nasal swab samples used for the evaluation of the Flu Detect RIDT, only 3 of the 38 tests that were positive by real-time RT-PCR were also positive by Flu Detect, while Flu Detect was negative in 35 nasal swab samples that were positive by real-time RT-PCR. Conversely, Flu Detect was positive for 218 of the 1,334 samples that were negative by real-time RT-PCR. Using real-time RT-PCR as the reference, the Flu Detect RIDT had a sensitivity of 7.9% (3/38), specificity of 83.7% (1,116/1,334), positive predictive value of 1.4% (3/221), and negative predictive value of 97.0% (1,116/1,151). All 6 shelters in the study showed similar inconsistencies between real-time RT-PCR and Flu Detect results. Although FluAB was clearly better than the Flu Detect test at diagnosing canine influenza in dogs, the 95% PI for FluAB sensitivity ranged from 48% to 80%, suggesting that true-positive samples in the current study might have been misdiagnosed as negative. Similarly, the Flu Detect test is poorly suited for CIV diagnosis, and, in fact, was more likely to give a false-positive rather than a true-positive result. It should be noted, however, that PPV is generally low when there is low prevalence of disease and, indeed, the true prevalence of CIV is currently unknown in all dog populations.
In conclusion, the findings in the current study suggest that real-time RT-PCR has the highest sensitivity of the 3 methods evaluated for detecting CIV nasal shedding and can be used as a rapid diagnostic test for CIV. If real-time RT-PCR cannot be performed for a suspected case of CIV, isolation and quarantine remain the safest approaches. However, this strategy is impractical for many facilities, and the cost for real-time RT-PCR might be higher than RIDTs. Facilities with financial constraints could consider testing in series, starting with FluAB RIDT performed on-site, followed by real-time RT-PCR confirmation while the potentially infected dog is isolated to minimize CIV transmission to naïve dogs. As turnaround time for real-time RT-PCR is approximately 1 day, the quarantine period could range from 48 hr (for a negative real-time RT-PCR sample) to 7 days (for a positive real-time RT-PCR sample). Ultimately, the real-time assay has the highest sensitivity for detecting virus nasal shedding and is recommended as a rapid diagnostic test for CIV.
Footnotes
Acknowledgements
The authors thank the veterinarians and staff from CSU-VTH and from the participating humane shelters for sample collection and performing the on-site test strips, and Dr. Katharine Benedict for help with the stochastic latent class modeling.
a.
BD Diagnostic Systems, Sparks, MD.
b.
Synbiotics Corp., Kansas City, MO.
c.
Gibco, Invitrogen Corp., Carlsbad, CA.
d.
Worthington Biochemical Corp., Lakewood, NJ.
e.
Kindly provided by Drs. M. McGregor and Y. Kawaoka, University of Wisconsin–Madison, Madison, WI.
f.
PE, Applied Biosystems, Foster City, CA.
g.
Lasergene 7.0, DNASTAR Inc., Madison, WI.
h.
Beacon Designer, Premier Biosoft Inc., Palo Alto, CA.
i.
QIAamp Viral RNA Mini Kit, Qiagen GmBH, Hilden, Germany.
j.
iScript One-Step RT-PCR Kit, Bio-Rad Laboratories, Hercules, CA.
k.
Mastercycler Realplex, Eppendorf AG, Hamburg, Germany.
l.
pGEM-T Easy vector, Promega Corp., Madison, WI.
m.
Kindly provided by Dr. D. P. Lunn, North Carolina State University, Raleigh, NC.
n.
Kindly provided by Dr. C. W. Olsen, University of Wisconsin–Madison, Madison, WI.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: Funding was provided by a grant from the Morris Animal Foundation (D09CA-009).
