Abstract
In September 2010, an outbreak of disease in 2 wild bird species (red-legged partridge,
Viruses belonging to the
In September 2010, BAGV was detected and identified in the southernmost province of Spain (Cádiz), as the etiological agent of an outbreak in wild birds.
1
The outbreak caused mortality in 2 bird species (red-legged partridges,
BAGV was identified as the causative agent of the outbreak during laboratory investigations using a pan-flaviviral hemi-nested reverse transcription polymerase chain reaction (RT-PCR) assay, amplifying a fragment of the nonstructural NS5 protein coding region of the viral RNA, 14 which was subsequently sequenced to confirm the identity of the viral genome detected. 1 However, this method is cumbersome, time consuming, and, as often happens with nested PCR methods, 9 prone to cross-contamination, and therefore is not suitable for efficient diagnosis and disease surveillance. With the aim to improve the efficacy and speed of the diagnosis of this disease, and to enable surveillance based on the detection of the BAGV genome, a real-time RT-PCR method specific for this virus was developed, and its diagnostic performance was evaluated using field samples and virus isolates from the first outbreak of this disease in Spain. The present article describes the development and evaluation of this new technique.
As a first step, nucleotide primers and a commercial probe aimed at the specific detection of BAGV were designed based on multiple alignments comprising a highly conserved sequence within the NS5 region of different flaviviruses. The alignment included sequences from 3 different BAGV strains (African strain Dak Ar B209, Indian strain 96363, and Spanish strain H/2010), 4 different WNV strains (Kunjin strain MRM61C, NY99, PT6.39, and Eg101), 1

Sequence alignment showing the position of the primers and probe designed in the current study with regard to the nucleotide sequence of
The probe and 1 of the selected primers contained, respectively, 1 and 3 degenerate positions to improve the specificity of the real-time RT-PCR for BAGV (Fig. 1). To achieve the best conditions for RT-PCR amplification, the protocol was optimized by testing different concentrations of primers and probe. The final protocol consisted of a 1-step RT-PCR with the following mixture: RT-PCR buffer, d RT-PCR enzyme mix, d forward primer (0.7 µM final concentration), reverse primer (0.7 µM final concentration), probe (0.15 µM final concentration), and 3 µl of template in a total of 20 µl of reaction volume. Amplification and fluorescence detection were conducted in real-time PCR equipment e using a program consisting of a reverse transcription step at 48ºC for 25 min followed by inactivation and denaturation at 95ºC for 10 min, and a PCR amplification cycle of 40 cycles of 95ºC for 15 sec, 52ºC for 30 sec, and 60ºC for 30 sec. Fluorescence data were acquired at the end of the 60ºC step. A hemi-nested RT-PCR specific for a broad range of flaviviruses 14 (the same used in the diagnosis of the outbreak) was performed in parallel in all the samples analyzed in this work, to allow a comparison.
The sensitivity of the new method was analyzed by testing either serial dilutions of viral RNA or viral suspensions (extracted using an automated procedure, f according to the manufacturer’s protocol) from 200-µl aliquots of cell-culture propagated virus seeds (titrated in median tissue culture infectious doses [TCID50]/ml by a standard limiting dilution titration assay 13 using BSR cells, virus titer: 104 TCID50/ml) of a cell culture-grown BAGV isolate Spain H/2010 from the heart of an affected partridge in the 2010 outbreak in Spain. 1 As a result, the real-time RT-PCR method for BAGV genome detection showed a detection limit of 10-1 TCID50/ml, which was comparable to the sensitivity achieved by the pan-flaviviral hemi-nested RT-PCR method performed in parallel (Fig. 2).
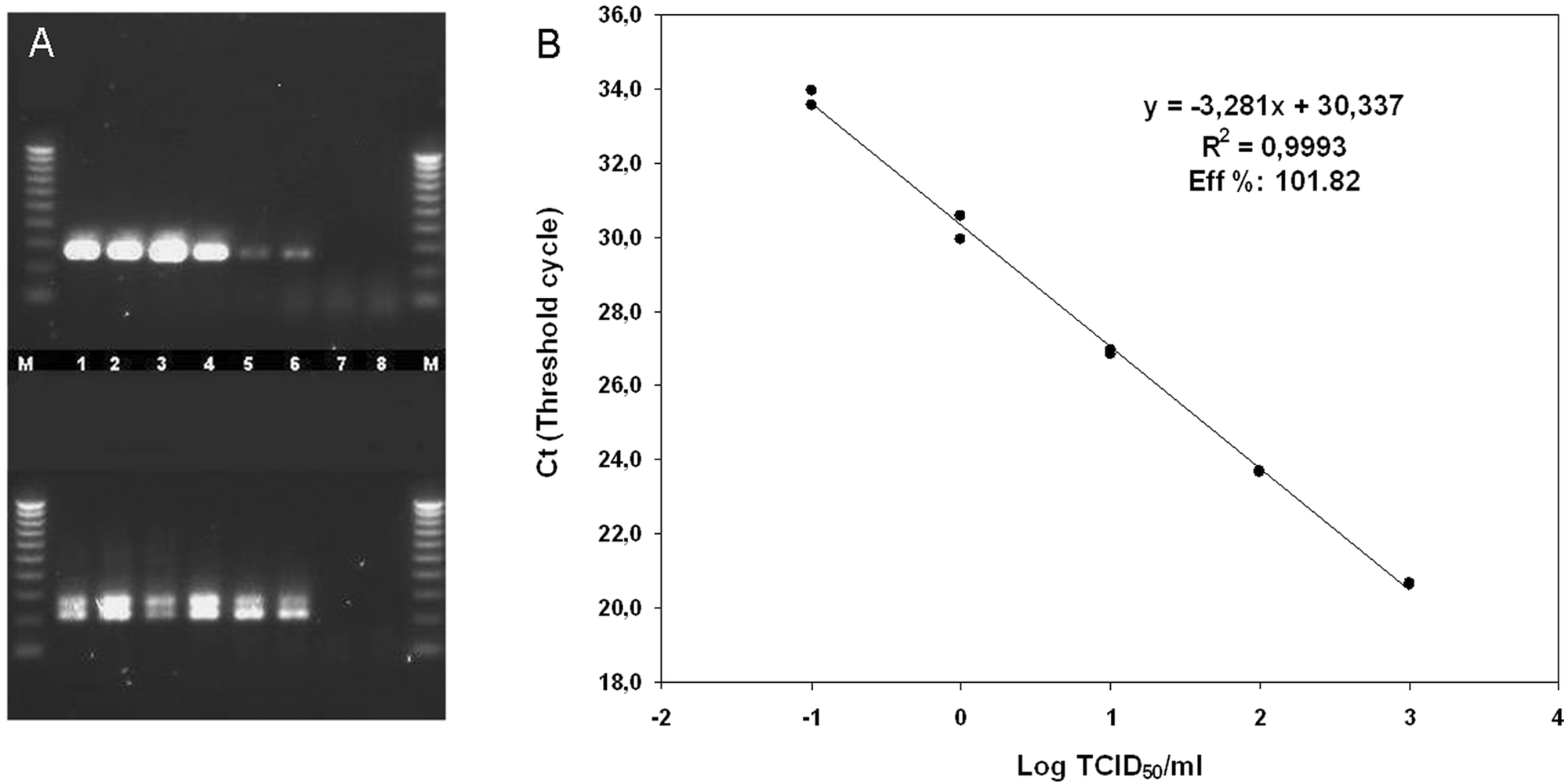
Comparison of sensitivity for
To study the specificity of the BAGV real-time RT-PCR, purified RNA samples from different flaviviruses, including BAGV isolate Spain H/2010, as well as non-BAGV flaviviruses (
To evaluate the performance of the newly developed real-time RT-PCR method in clinical samples, specimens were obtained from red-legged partridges (
Comparative analysis of clinical samples from affected red-legged partridges (
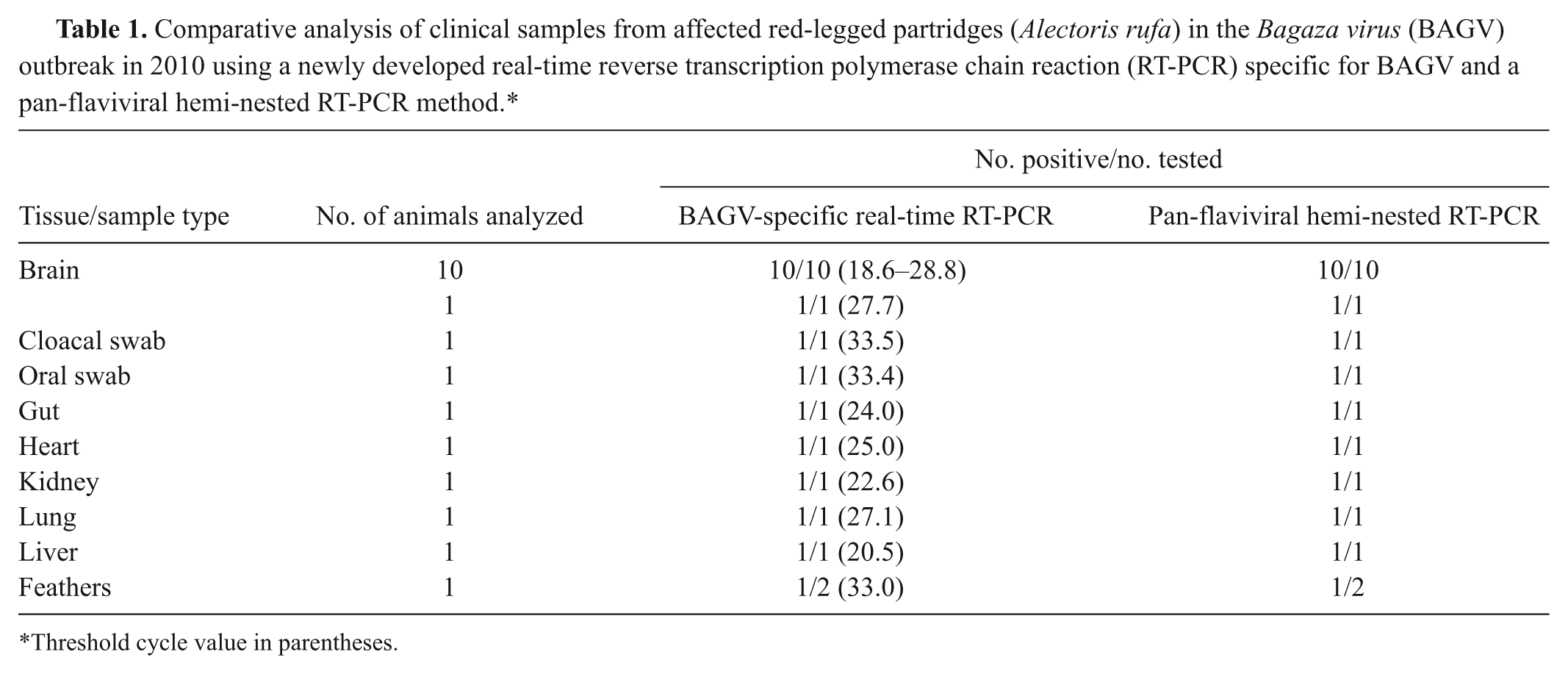
Threshold cycle value in parentheses.
In summary, a real-time RT-PCR method was developed that was specifically designed to detect the presence of BAGV genome in clinical samples with a high sensitivity. This new method proved to be valid for the detection of BAGV genome either in virus isolates or in clinical samples from affected animals, providing a sensitivity equivalent to the gel-based pan-flaviviral RT-PCR method previously employed. As compared to the gel-based RT-PCR, the new method offers several advantages, besides its specificity to BAGV. The new method reduces the time required for analysis (less than 4 hr following receipt of the samples) without any loss of sensitivity or specificity. The assay also significantly reduces the risk of false-positive results due to cross-contamination, which is a well-known drawback of nested PCR protocols. 9 In addition, in combination with an automated extraction system, this new method would enable large-scale analysis of the virus in animal populations potentially affected with the disease, a feature that may be useful in case of recurrence in the area and/or spread to other areas. This method provides the diagnostic laboratory with an effective and rapid analytical tool, useful in differential diagnosis and surveillance of this flaviviral disease of birds, which could represent an emerging animal health risk in Europe.
Footnotes
b.
Primer Express (version 2.0.0) software, Applied Biosystems, Branchburg, NJ.
c.
TaqMan–MGB Probe, Applied Biosystems, Branchburg, NJ.
d.
AGPATH-ID ONE RT-PCR kit, Applied Biosystems, Branchburg, NJ.
e.
7500 Fast Real Time PCR System, Applied Biosystems, Branchburg, NJ.
f.
BioSprint DNA Blood Kit, BioSprint 96 Workstation; Qiagen Inc., Valencia, CA.
g.
MagNA Lyser, Roche Diagnostics, Indianapolis, IN.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This study has been funded by the Ministries of Agriculture, Food and Environment, and Economy and Competitiveness of Spain (grant AGL2011-13634-E).
