Abstract
The present report describes an atypical Brucella ovis strain (Bo10) isolated from the epididymis and testis of an infected ram. Macroscopic and microscopic lesions characteristic for the infection, including positive Brucella immunostaining, were observed within lesions in the genital organs. Compared to other isolates, strain Bo10 required an additional day (a total of 96 hr) of incubation to form visible colonies, showed a distinct carbon source utilization profile, agglutinated only weakly with rough (R) serum, but showed a high capacity for autoagglutination. Isolate Bo10 failed to produce the 1,071-bp fragment in the outer membrane protein (omp) 31 gene–based part of the “Bruce-ladder” multiplex polymerase chain reaction system but did produce a 1,915-bp amplicon, thus presenting a profile similar to Brucella abortus. Sequence analysis of the 1,915-bp fragment revealed an 842-bp long insertion sequence (IS)711 transposon element inserted into the promoter region of the omp31 gene, immediately upstream from the ribosome binding site (-10 box/Pribnow box). Sodium dodecyl sulfate–polyacrylamide gel electrophoresis of a whole-cell lysate showed the absence in Bo10 of the approximately 31-kDa protein fragment associated with omp31. The results demonstrate a natural inactivation of omp31 and, consequently, the absence of the Omp31 protein in this B. ovis isolate. The novel location of IS711 within the genome of a naturally occurring B. ovis strain supports the hypothesis that IS711 could be an active transposon in this Brucella species.
Ovine epididymitis, caused by Brucella ovis, can lead to severe economic losses in sheep-producing countries.2,5,20 However, B. ovis can be successfully eradicated from flocks using the test and cull approach by focusing only on rams in the testing program.3,21 Brucella ovis is a small coccobacillus that lacks an O-polysaccharide chain, and thus expresses rough lipopolysaccharide as a surface antigen. Beside lipopolysaccharide, the outer membrane proteins (OMPs) are the other major components of the cell wall of B. ovis, which serve as protective antigens.4,7 A major OMP is Omp31, a target for B. ovis vaccine development.6,9
Brucella ovis has an approximately 3.27 mega base pair (bp) genome structured into 2 chromosomes (CP000708 and CP000709). Genomes of all Brucella species contain the 842-bp insertion sequence (IS)711 in varying copy numbers; IS711 also serves as a target to identify different Brucella species.13,16,19 The genome of B. ovis bears 38 copies of IS711. 22 Although IS711 is very stable, mobility of IS711 has been demonstrated in vitro in Brucella pinnipedialis and B. ovis by a transposon trap. 18 It has also been hypothesized that IS711 could be an active genome element. 23 The current study describes an inactivation of the omp31 gene caused by IS711 in a B. ovis field isolate obtained during the eradication of ovine epididymitis from a flock in Hungary, central Europe.
In 2011, a test and cull eradication program was conducted in a flock that consisted of 2,400 ewes and 93 rams in Hungary. In brief, rams were separated from the ewes, and blood samples were taken from each animal at monthly intervals and examined by B. ovis–specific indirect enzyme-linked immunosorbent assay (ELISA).a,21,25 Positive rams were removed from the flock after each test and stalled in a separate farm until their slaughter. 21 The epididymes and testes of seropositive rams were collected and submitted for laboratory examinations. Gross pathological, histopathological, immunohistochemical, bacteriological, and polymerase chain reaction (PCR) examinations were performed from samples of the epididymes and testes as previously described. 12
In brief, 4-μm thick sections of formalin-fixed and paraffin-embedded tissue samples were stained with hematoxylin and eosin, and examined by light microscopy. Hyperimmune rabbit serum was produced for immunohistochemistry by intravenous infection of an animal with 4 × 109 colony-forming units of live Brucella canis in accordance with all applicable institutional and national guidelines. Deparaffinized sections were incubated overnight at 4°C with the hyperimmune serum diluted to 1:20,000, and antibody binding was detected by a horseradish peroxidase–labeled polymer. b
Brucella-selective agar c containing 5% heat-inactivated horse serum, d trypticase soy agar, e and chocolate agar c containing 0.1% yeast extract c were inoculated with 100 µl of homogenized testes and epididymes. The plates were incubated at 37°C in 5% carbon dioxide for 7 days. In addition, 1-ml aliquots of homogenized organs were cultured at 37°C for 21 days in trypticase soy broth e before 100 µl of inoculum was transferred to Brucella agar plates on days 7, 14, and 21 and cultured as described above. Growth of B. ovis was checked daily. Routine biochemical and growth-based typing tests were performed on the isolated strains. 1 Carbon-source utilization of the strains was examined with a commercial system. f Slide agglutination tests of the isolated strains were carried out using A, M (monospecific sera against the A or M agglutinogen of Brucella spp. forming smooth colonies), and R (serum against Brucella spp. with rough colony form) sera. g The autoagglutination capacity of the isolated strains was tested as previously described, 4 with minor modifications. In brief, strains were grown on trypticase soy agar e for 48 hr, and a suspension with an optical density of 0.8 at 600 nm (OD600) was prepared in trypticase soy broth e containing 5% heat-inactivated horse serum d and Brucella-selective supplement. c A 2-ml aliquot of each suspension was left in a spectrophotometer microcuvette without agitation at room temperature, and the OD600 reading was scored for 48 hr at 6-hr intervals.
DNA was extracted from homogenized testes and epididymes of seropositive rams with a commercial DNA isolation kit, h and the samples were screened by PCR using the primer pair BMEI0535f/BMEI0536r from the “Bruce-ladder” method.12,14 DNA was extracted from 1 distinct colony of each isolated strain with a commercial kit h according to the manufacturer’s recommendations. The molecular typing of the strains was based on the Bruce-ladder multiplex PCR system following the recommendations of the OIE Manual of Diagnostic Tests and Vaccines for Terrestrial Animals.10,14,24 The PCR reactions were performed as previously described. 11 In order to amplify a larger fragment of the omp31 gene, a PCR assay was designed utilizing the primer pair BtransF2 (5′-CGCCGAATGTGATTAGGT-3′) and BtransR2 (5′-GGACGCCAAAATTACGGACTTC AT-3′; Fig. 1). The PCR reaction was performed in a 25-µl volume, containing 1 µl of target DNA, 5 µl of buffer, i 2 µl of 25 mM magnesium chloride, 1 µl of 10 mM deoxyribonucleotide triphosphate, 1 µl of each 10 pmol/µl primer, and 0.2 µl of Taq polymerase (5 U/µl). i The PCR assay consisted of initial denaturation for 5 min at 95°C followed by 35 amplification cycles consisting of denaturation for 30 sec at 95°C, primer annealing at 55°C for 30 sec, and extension at 72°C for 60 sec. Following agarose gel electrophoresis, direct cycle sequencing was performed with both primer pairs used for amplification on an automated DNA sequencer. j The reading errors of the chromatograms were corrected, and an annotated PCR product was constructed. k
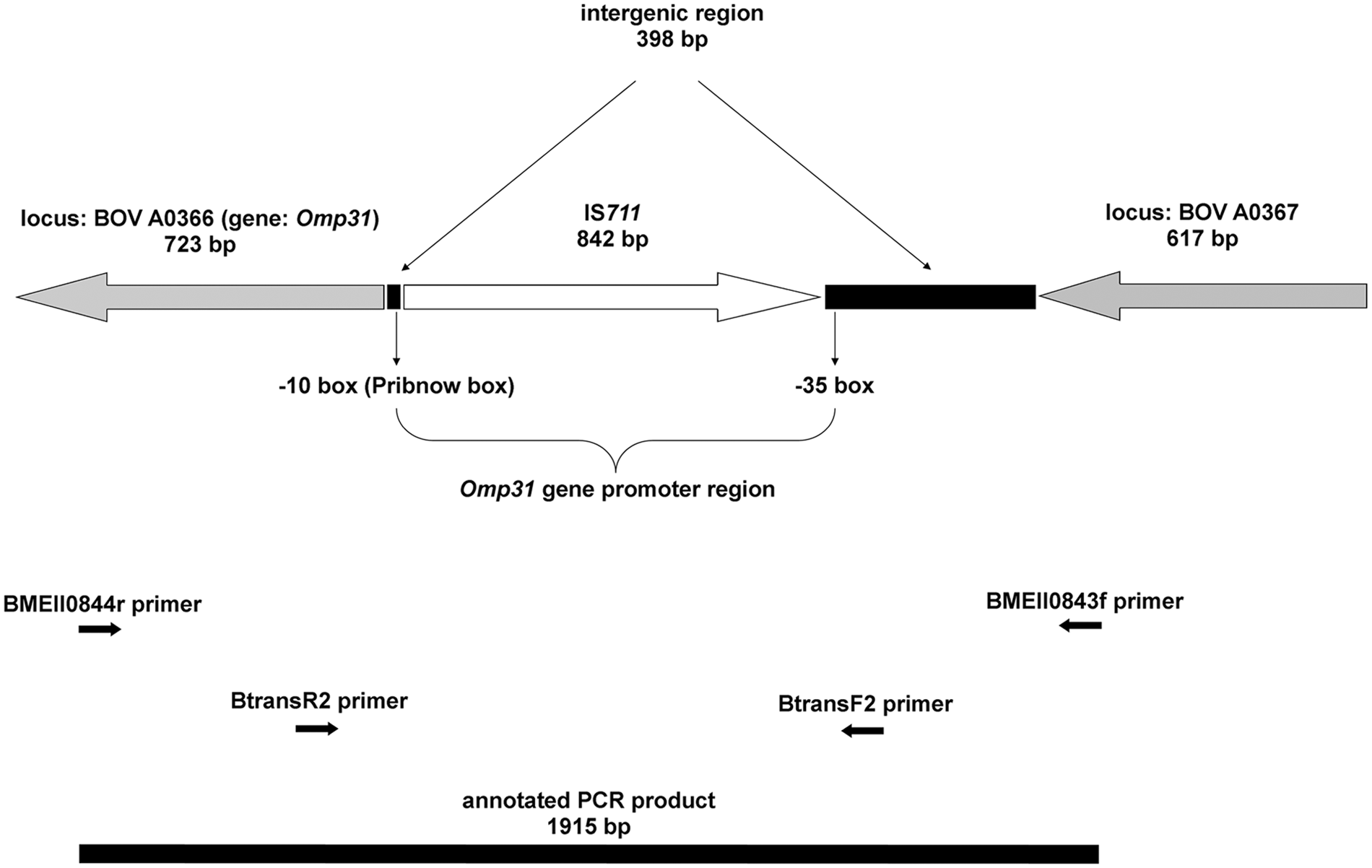
Genomic location of the newly identified copy of IS711 transposon, omp31 gene, and the promoter region; polymerase chain reaction (PCR) primers used; and the amplified PCR product on chromosome II of Brucella ovis strain Bo10.
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) was performed with whole-cell lysates as previously described. 17 Briefly, sonicated bacteria were treated with an equal volume of lysis buffer (50 mM Tris-HCl, 100 mM NaCl, 0.1% Triton X-100 m ) for 10 min at 25°C. After incubation, samples were diluted 1:2 in Laemmli buffer l and boiled for 10 min. The proteins were visualized by Coomassie brilliant blue staining. m A low-range SDS-PAGE standard d was used to determine the molecular mass of the proteins of the Omp25/Omp31 protein family.
Altogether, 42 seropositive rams were culled during monthly ELISA examinations until the flock reached seronegative (“disease-free”) status after 8 months. In total, B. ovis was cultured from 27 of the 42 seropositive rams (64%). These 27 and 1 additional (altogether 28) out of the 42 tested animals (67%) were PCR positive. The other 14 seropositive rams were negative by both culture and PCR. Except for 1 isolate, named Bo10, the strains were clearly identified as B. ovis both with classical and molecular methods (data not shown). Important agents causing ovine epididymitis other than Brucella species (Actinobacillus seminis, Histophilus somni, Corynebacterium pseudotuberculosis, and Trueperella pyogenes) were ruled out by bacteriological examination.
The ram, infected with strain Bo10, was also ELISA and PCR positive. During gross pathological examination of this animal, macroscopic lesions were identified as well-demarcated, spherical foci with a diameter of 1.5–2 cm in the epididymis, containing a yellowish-white, creamy material. Histopathology revealed severe, diffuse, chronic, purulent epididymitis and the presence of abscesses and chronic lymphohistiocytic inflammation in the testes. Brucella antigen was found in the cytoplasm of macrophages located in and around the ductus epididymidis and in abscesses, as well as extracellularly within the central necrotic core of abscesses (Fig. 2). Infrequent immunostaining was also observed in other cell types such as ductus epididymidis epithelial cells, lymphocytes, plasma cells, and intravascular monocytes.
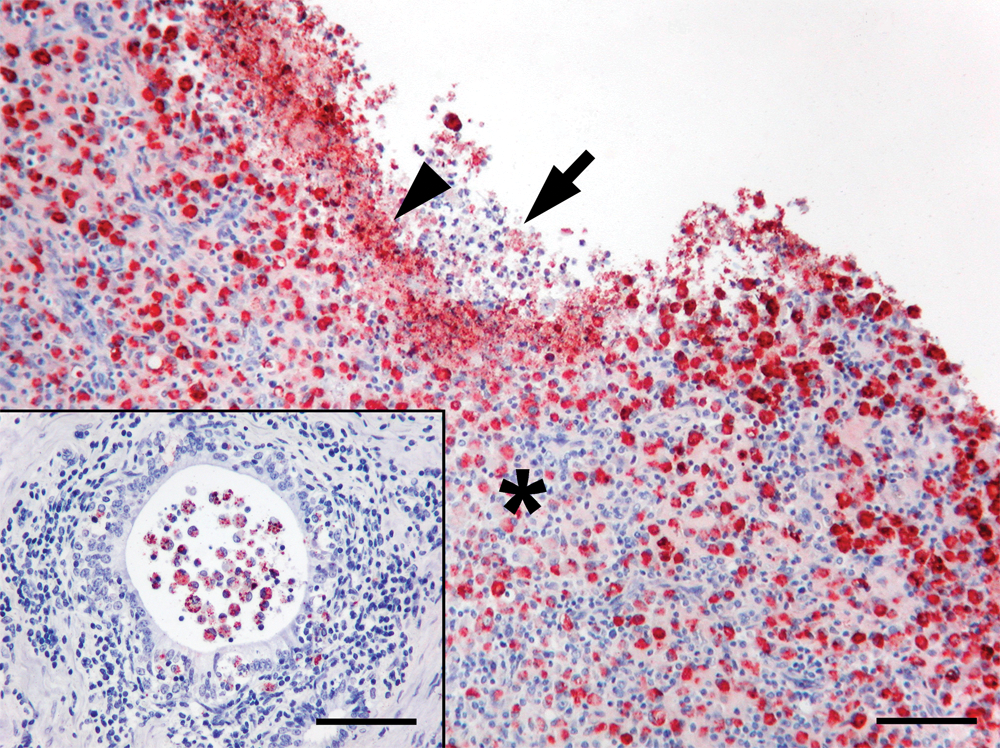
Ram; epididymis; case 10 (Brucella ovis strain Bo10). Remnants of necrotic inflammatory cells in the centre of abscess with moderate amounts of intra- and extracellular Brucella antigen (arrow), and a narrow layer of large amounts of extracellular Brucella antigen at the border of necrotic central area (arrowhead) are shown. The zone of lymphocytes, plasma cells, and large numbers of macrophages is also evident, containing large numbers of macrophages immunostained for Brucella (asterisk). Immunohistochemistry using hyperimmune rabbit anti–Brucella canis primary antibodies cross-reacting with B. ovis, hematoxylin counterstain. Bar = 60 µm. Inset: chronic inflammation with the dilatation of ductus epididymidis by inflammatory cells. Strong immunostaining is seen for Brucella in macrophages and in a few epithelial cells and lymphocytes. Immunohistochemistry using hyperimmune rabbit anti–B. canis primary antibodies cross-reacting with B. ovis, hematoxylin counterstain. Bar = 100 µm.
Large numbers of visible colonies of strain Bo10 appeared 1 day later (at 96 hr) than the other isolates. The colonies were small and yellow-colored, had a creamy consistency, and showed rough colony morphology with crystal violet staining. The Gram- and Köster-stain negative, small coccobacilli demonstrated catalase activity, but did not produce oxidase, nor urease or hydrogen sulfide. The Bo10 strain grew in the presence of 20 mg/ml of thionin but was inhibited by basic fuchsin. As the commercial identification system b used in the current study is unable to identify B. ovis, all isolates were identified or presumed to be Brucella melitensis (probability range: not measured to 100, similarity range: 0.404–0.557) except for Bo10, which was presumed to be Brucella suis (probability: not measured, similarity: 0.219). Strain Bo10 showed very weak agglutination with serum R, and agglutination was not observed with sera A and M. Strain Bo10 showed high capacity for autoagglutination; the OD600 decreased to 0.149, 0.040, and 0.000 after 6, 12, and 48 hr of incubation, respectively, while all other isolates remained in suspension (OD600 of approximately 0.8) throughout the experiment.
Isolate Bo10 failed to produce the 1,071-bp fragment in the omp31 gene–based part of the Bruce-ladder PCR system but it yielded an approximately 1,900-bp long amplicon, thus presenting a profile similar to Brucella abortus (Fig. 3). This fragment was successfully amplified and sequenced with the BMEII0843f/BMEII0844r primer pair of the Bruce-ladder method and the BtransF2/BtransR2 primer pair designed in the current study. The exact size of the fragment (JX081250) was 1,915 bp and contained the 842-bp insertion sequence, IS711 (Fig. 1). The sequence of the IS711 was identical to the IS711 sequence in GenBank (accession no. M94960) except the mismatch (C-T) of the 5th nucleotide at 5′-end. This transposon element was located in an intergenic region, exactly in the promoter region of omp31, immediately upstream from the ribosome binding site (-10 box/Pribnow box) and 388-bp downstream from the BOV A0367 locus. Thus, the presence of IS711 enlarged the original 1,071-bp PCR product to a 1,915-bp amplicon. Contrary to the other B. ovis strains, the approximately 31-kDa protein fragment typical of Omp31 was not detected in strain Bo10 by SDS-PAGE (Fig. 4).
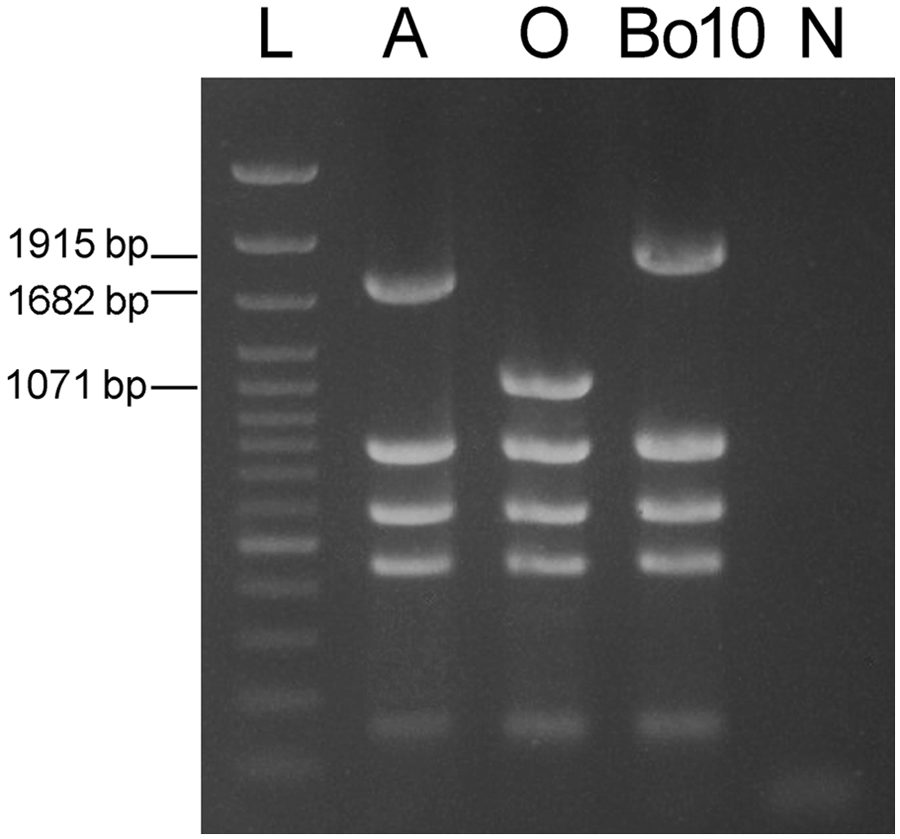
Differentiation of Brucella abortus (A), Brucella ovis (O), and B. ovis strain Bo10 by “Bruce-ladder” multiplex polymerase chain reaction. L = DNA ladder n ; N = negative control.
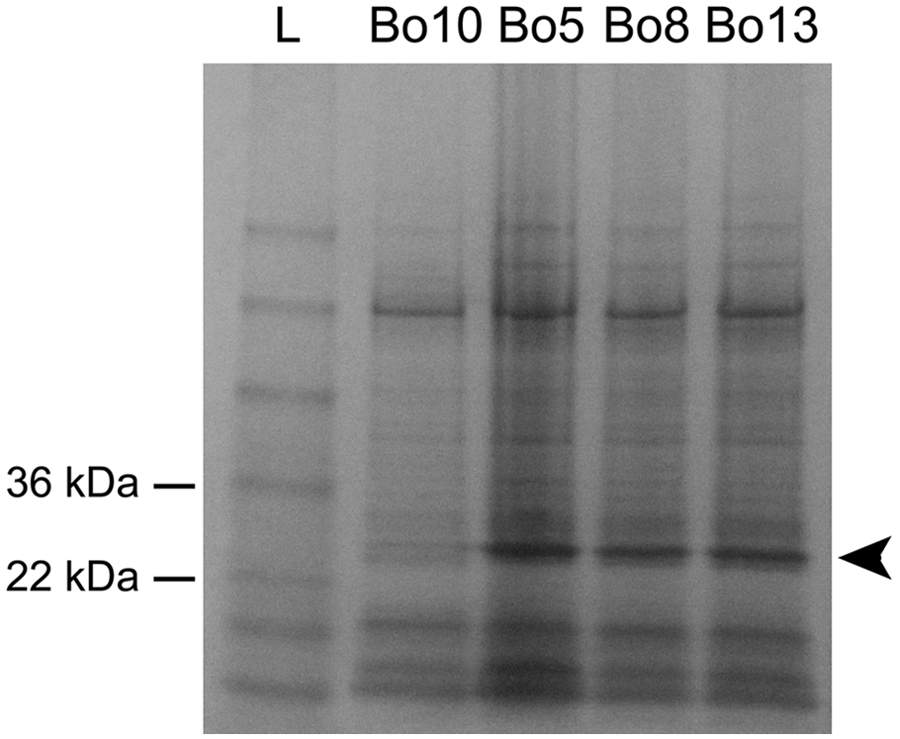
Sodium dodecyl sulfate–polyacrylamide gel electrophoresis analysis of a whole-cell lysate of Brucella ovis strain Bo10 and 3 control B. ovis strains (Bo 5, Bo8, and Bo13) isolated during the eradication program. Note the lack of the strong protein band at approximately 31 kDa in Bo10. The 31-kDa large protein fragment typical of Omp31 is marked by an arrowhead in the 3 control B. ovis strains. L = protein ladder. e
Brucella ovis was successfully eradicated from the sheep flock following the principles of the test and cull approach. Gross pathological, histopathological, and immunohistochemical lesions that presented in both the host of the Bo10 isolate and in the other rams were typical for B. ovis infection. The lack of Omp31 protein in isolate Bo10 did not appear to influence the use of immunohistochemistry, as this method uses a polyclonal rabbit serum. The colony and microscopic morphology, staining, biochemical profile, and lack of agglutination with A and M sera of both strain Bo10 and all other isolates were typical for B. ovis. 1 With the exception of the Bo10 strain, the strong agglutination of the strains with R sera were also typical for B. ovis. The Bruce-ladder method was effective for the identification of the other isolates associated with this outbreak. However, in the case of the atypical Bo10 strain, the irregular PCR product could have led to the misidentification of the isolate as B. abortus, which is a rare cause of reproductive disorders in sheep.8,11,15
The current study describes a new location for IS711 in the genome of B. ovis. Previous studies suggested that IS711 could be an active element, but mobility of IS711 in B. ovis has only been proven in vitro. Under field conditions, transposition was shown solely in B. abortus.16,18,23 The findings support the previous hypothesis that IS711 could be an active transposon in B. ovis.
The slower growth ability, autoagglutination capacity, genomic localization of IS711, and absence of the typical 31-kDa protein fragment of Omp31 in the Bo10 strain confirm that this is indeed a case of natural inactivation of the omp31 gene in B. ovis. This finding is consistent with the results of an earlier study, in which identical properties were described in an artificially created, omp31 gene–deleted B. ovis mutant strain. 4 Similar to the artificial omp31 mutant B. ovis strain, the Hungarian Bo10 strain is a virulent isolate as shown by the significant pathological and histopathological lesions observed in the genital organs of the affected ram. In conclusion, a natural IS711 insertion-caused inactivation of the omp31 gene was detected in a virulent B. ovis strain.
Footnotes
a.
Diavet Ltd., Budapest, Hungary.
b.
EnVision+ Kit, Dako Denmark A/S, Glostrup, Denmark.
c.
Oxoid Ltd., Cambridge, United Kingdom.
d.
Invitrogen Corp., Carlsbad, CA.
e.
Difco, BD Diagnostic Systems, Sparks, MD.
f.
MicroLog MicroStation System, GN2 Microplates; Biolog Inc., Hayward, CA.
g.
French Agency for Food, Environmental & Occupational Health Safety/ANSES/, Maisons-Alfort Cedex, France.
h.
QIAamp DNA Mini Kit, Qiagen Inc., Valencia, CA.
i.
Promega Inc., Madison, WI.
j.
ABI PRISM 3100-Avant Genetic Analyzer, Applied Biosystems Inc., Foster City, CA.
k.
BioEdit v. 7.1.3.0, Sequence Alignment Editor; Ibis Biosciences, Carlsbad, CA.
l.
Bio-Rad Laboratories, Hercules, CA.
m.
Sigma-Aldrich Inc., St. Louis, MO.
n.
GeneRuler 100 bp Plus DNA Ladder, Fermentas Inc., Glen Burnie, MD.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Lendület program (grant no. LP2012-22) of the Hungarian Academy of Sciences.
