Abstract
Toxoplasma gondii and Chlamydophila abortus are the 2 most common infectious causes of ovine abortion worldwide. These obligate intracellular pathogens are associated with severe placentitis leading to abortion or stillbirth in pregnant ewes, and resulting in significant economic losses. The objectives of the current study were the development, validation, and application of a duplex real-time polymerase chain reaction (PCR) assay capable of quantifying the burden of infection by T. gondii and C. abortus in material submitted for diagnostic purposes. The validation was carried out using samples from ewes experimentally infected with these organisms. Based on the numbers of genome copies detected, an arbitrary cutoff level was established to correlate with significant pathological changes sufficient to give rise to abortion. When the PCR assay was applied to samples from 66 Irish farms with naturally occurring outbreaks of ovine abortion, toxoplasmosis and enzootic abortion of ewes (EAE) accounted for 14% and 20% of the farms, respectively, while on 6% of the farms, there was evidence of dual infection. When standard diagnostic techniques including histopathological examination, serological analysis, chlamydial antigen detection, and bacteriological culture, were used on samples from the same farms, toxoplasmosis was diagnosed in 17% of farms, and EAE in 12%; dual infection was diagnosed on 3% of the farms. In general, good agreement was found between the PCR and the standard methods. The duplex real-time PCR assay developed in this study has proved to be a very sensitive and rapid tool that might provide a valuable addition to the methods currently available for routine diagnosis of ovine abortions.
Keywords
Introduction
Toxoplasma gondii and Chlamydophila abortus are the 2 most common causes of infectious abortion in sheep worldwide.4,5,28 Clinical ovine toxoplasmosis occurs following primary infection of pregnant sheep as a result of the ingestion of sporulated oocysts. 24 Infected animals may subsequently abort or produce stillborn and/or weak lambs, often along with a small, mummified fetus. 24 With C. abortus, the major sources of infection are vaginal discharges, placentas, fetuses, and coats of lambs. 15 Infected pregnant ewes usually abort but may on occasion give birth to live lambs, which are generally weak and do not survive. 15
The cost of toxoplasmosis to the sheep industry is difficult to determine because, in addition to causing abortion during the later stages of gestation, T. gondii can also cause fetal loss at earlier stages of pregnancy, which may go undetected. 3 The best estimate of the annual cost of toxoplasmosis to the U.K. sheep industry based on lost output (abortions/stillbirths), input costs, and the cost of control measures is £12.4 million while the corresponding figure for enzootic abortion of ewes (EAE) is £23.8 (Bennet R, Ijpelaar J: 2003, Economic assessment of livestock diseases in Britain. Final report to the Department for Environment, Food and Rural Affairs; http://randd.defra.gov.uk/Document.aspx?Document=ZZ0102_1215_FRP.doc). If the diagnosis is confirmed rapidly and reliably, then control measures can be implemented to reduce the impact of infection in current and future years, thus protecting the economic viability of the flocks.
The definitive method for diagnosis of T. gondii and chlamydial infection is isolation of the pathogen from infected tissues.10,32 However, this is labor-intensive, time-consuming, and expensive, relies on the submission of fresh material to the diagnostic laboratory, and is frequently impractical in diagnostic situations. 34 Histological examination of placenta and fetal brain, often combined with immunohistochemical examination of suspect lesions, is a recommended method for the diagnosis of toxoplasmosis. 36 In addition, serological testing can help confirm infection in ewes that have aborted due to toxoplasmosis because the majority of ewes remain seropositive for at least 6 months following infection. 34 Serum agglutination tests and commercial T. gondii antibody enzyme-linked immunosorbent assays (ELISAs) are currently being used in diagnostic laboratories. Serological diagnosis of EAE can be carried out using complement fixation and ELISA,12,22 but they may exhibit varying degrees of sensitivity and specificity leading to false-positive results in sheep previously vaccinated or infected with Chlamydophila pecorum.11,35
Inconclusive results or a failure to reach a diagnosis are frequently reported by diagnostic veterinary laboratories for ovine abortion cases despite detailed investigation and testing. 34 Although the current diagnostic techniques can provide an accurate diagnosis of ovine toxoplasmosis and enzootic abortion, results may be missed as they are dependent on factors such as the level of infection, the host immune responses, sample selection, and in the case of histopathology, the particular sections examined. Recently, molecular techniques such as the real-time polymerase chain reaction (PCR) have emerged as highly sensitive and reliable methods for T. gondii and C. abortus DNA detection.27,28,31 The real-time PCR method is a very sensitive and rapid technique that allows direct detection and quantification of the pathogen in biological samples. 31 A validated PCR test for C. abortus or T. gondii in ovine tissues might help identify infected fetuses that are “missed” due to the absence of typical histological lesions, or the absence of fetal membranes from the sample submission. 34 Ideally, placental material should be submitted for PCR-based diagnostics. However, these pathogens have also been identified in fetal hair coat swabs, maternal vaginal swabs, and fetal organs including liver, lung, spleen, and brain.14,15 Such specimens may be suitable for diagnosis using highly sensitive molecular assays.
The objective of the current study was to test the usefulness of a quantitative real-time PCR as a diagnostic method for the detection of C. abortus and T. gondii in different field material submitted from cases of ovine abortion. The results were compared to those obtained using standard methods available in the Irish Regional Veterinary Laboratories.
Materials and methods
DNA purification and standard curve preparation for the duplex real-time PCR
DNA from T. gondii and C. abortus was purified following manufacturer’s instructions.a,b Toxoplasma gondii DNA was obtained from a commercial vaccine, c while C. abortus DNA was obtained from chlamydial elementary bodies (EBs) purified from C. abortus–infected McCoy cells as previously described. 15 The concentration of DNA was determined by a spectrophotometer d and related to copy numbers based on the molecular weight of the complete genome sequence of C. abortus and T. gondii. Standard curves were then constructed using 10-fold serial dilutions of known copy numbers of either C. abortus or T. gondii DNA. The standard curves were used as templates in the real-time PCR assay to determine the limit of detection and efficiency.
Duplex real-time PCR including internal control
A duplex real-time PCR was developed to detect simultaneously T. gondii and C. abortus by using a specific commercial chemistry. e The selected target amplicons were the 529-bp repeat element RE and the ompA gene for T. gondii and C. abortus, respectively.8,27 An internal amplification control (IC1) 16 was included to increase confidence in the assay and to identify false negatives. All the primers and probes are shown in Table 1. Primers and probes were commercially provided, f and their specificity was tested using the ClustalW2 software (http://www.ebi.ac.uk/Tools/clustalw2/index.html) to avoid any cross-hybridization between amplicons. Amplifications were carried out on a thermocycler, g the threshold cycle (Ct) values were set up automatically, and data were analyzed using commercial software. h The slope (S) and the regression coefficient (R2) of the standard curves were calculated to estimate the efficiency of the assay using the following equation: [10(-1/slope) − 1] × 100.
Target genes, internal controls, and corresponding primers and probes selected for the development of the multiplex real-time polymerase chain reaction.*
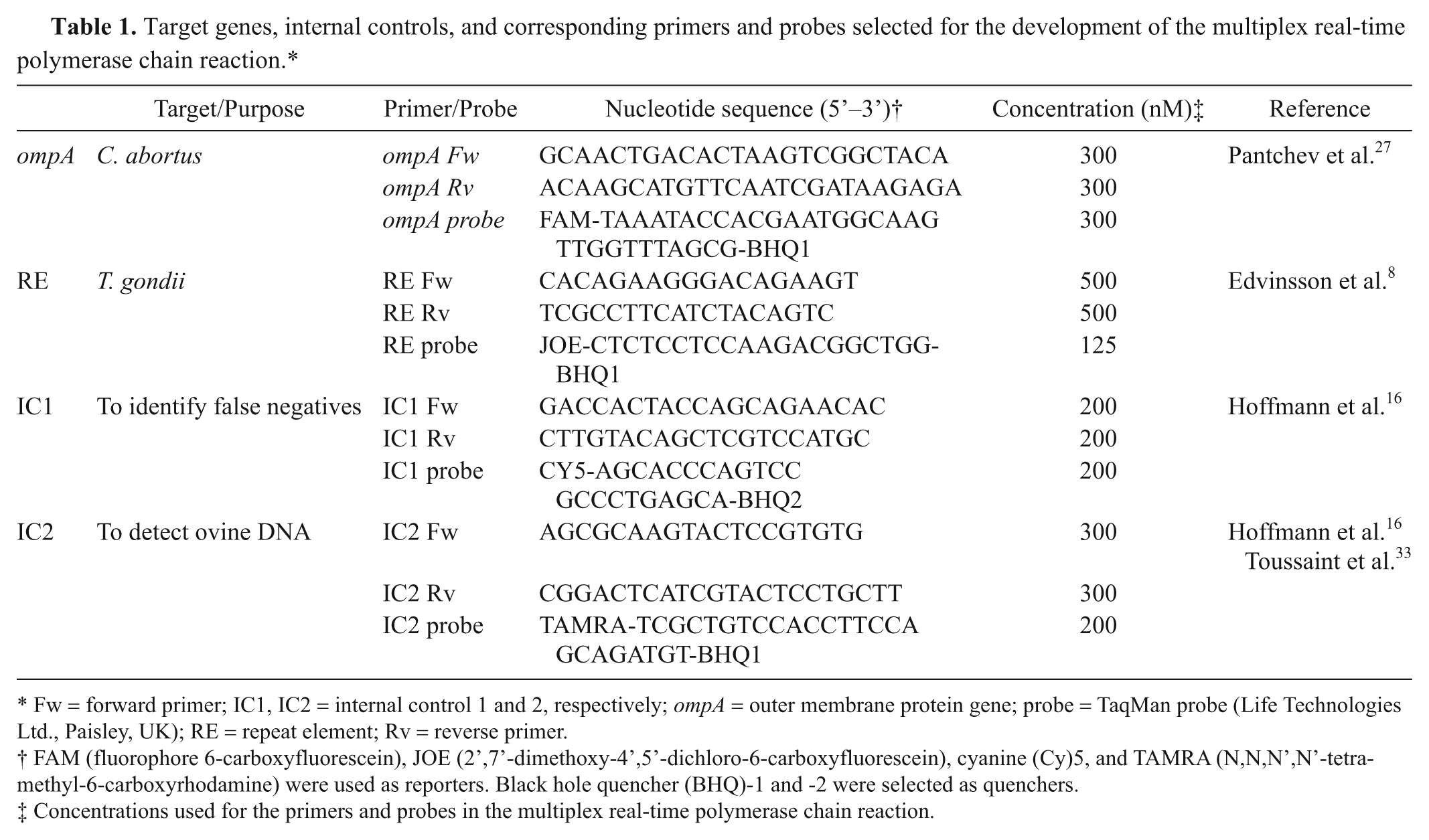

Fw = forward primer; IC1, IC2 = internal control 1 and 2, respectively; ompA = outer membrane protein gene; probe = TaqMan probe (Life Technologies Ltd., Paisley, UK); RE = repeat element; Rv = reverse primer.
FAM (fluorophore 6-carboxyfluorescein), JOE (2’,7’-dimethoxy-4’,5’-dichloro-6-carboxyfluorescein), cyanine (Cy)5, and TAMRA (N,N,N’,N’-tetramethyl-6-carboxyrhodamine) were used as reporters. Black hole quencher (BHQ)-1 and -2 were selected as quenchers.
Concentrations used for the primers and probes in the multiplex real-time polymerase chain reaction.
Duplex real-time PCR statistical analysis
The DNA concentrations used to establish the standard curves were tested in duplicate and replicated 3 times. Statistical analysis was performed using commercial software. i Threshold cycle values were compared using analysis of variance, followed by least significant difference testing at the P < 0.05 level.
Validation of the duplex real-time PCR
The validation of the duplex real-time PCR was carried out using samples obtained from pregnant sheep that were orally infected with C. abortus or T. gondii.14,15 Samples from noninfected pregnant sheep were also tested as control samples. The collected samples included placenta, fetal brain, fetal tissues (liver, lung, and spleen), fetal aqueous humor, amniotic and allantoic fluids, fetal hair coat swabs, and vaginal swabs. DNA was extracted following manufacturer’s instructions j and tested in duplicate using the duplex real-time PCR assay. A second internal control (IC2) was included when DNA from swabs was tested to confirm the presence of ovine DNA and verify the effectiveness of the nucleic acid extraction step (see primers and probes in Table 1). 33 Based on the range of genome copies detected for each type of sample and the presence or absence of significant lesions consistent with EAE or toxoplasmosis, a cutoff point was established for the PCR assay above which the pathogen burden was correlated with significant pathology.
Collection of field samples for the application of the duplex real-time PCR
Samples were collected from cases of ovine abortion that had occurred on 66 Irish farms and included placenta, fetal brain, fetal tissues, fetal stomach contents, fetal pleura fluid, fetal hair coat swabs, and vaginal swabs. Samples were obtained from at least 2 animals on each farm and from at least 2 different types of samples from the same animal. Collected samples were then tested using the duplex real-time PCR, and diagnosis was based on the estimated cutoff point described above. Samples were also tested by methods in routine use for diagnosis of ovine abortions, including histopathological examination, a commercial chlamydial lipopolysaccharide (LPS) detection test, k serological tests, and culture. Histopathological examination was carried out on placentomes and fetal brains for the presence of lesions consistent with EAE or toxoplasmosis. Vaginal swabs were tested using the commercial chlamydial LPS detection test. Sera were examined using an agglutination test for T. gondii antibodies l and an ELISA specific for antibodies to the polymorphic membrane proteins (PMP) of C. abortus. m Placenta, fetal liver, and fetal stomach contents were subjected to routine bacteriological culture methods.
Genotype analysis of T. gondii isolates by RFLP
Toxoplasma gondii strains detected in the field samples were subjected to genotyping as follows. Nested PCR was used to amplify the following loci of T. gondii: 5’-SAG2, 3’-SAG2, BTUB, GRA6, and SAG3.9,13,17,19 Restriction fragment length polymorphism (RFLP) analysis was then carried out by digestion of the amplification products with HhaI, Sau3AI, TaqI, MseI, and NciI, respectively. Restriction enzymes were commercially provided. n Restriction fragments were separated on 3% agarose gels and visualized under ultraviolet light. o The M4 strain of T. gondii was used as genotype II positive control. 14
Results
Development of the duplex real-time PCR
Threshold cycle values obtained for the standard curves were not significantly different when target genes were tested alone or in combination (data not shown). The limit of detection was established as 1 and 100 genome equivalents of T. gondii and C. abortus, respectively, and the efficiency of the assay was 91% for RE gene and 88% for the ompA.
Validation of the duplex real-time PCR
The consistency of the duplex real-time PCR was tested on samples obtained from 17 and 20 pregnant ewes orally infected with T. gondii and C. abortus, respectively.14,15 The vast majority of the collected samples were positive for T. gondii or C. abortus by the real-time PCR, but at different levels depending on the type of sample tested and the number and severity of the lesions present. As shown in Table 2, fewer than 103 genome copies were detected in placental samples from animals that did not display any significant histopathological lesion consistent with toxoplasmosis. A similar finding was observed when fetal brain and tissues were tested, but the threshold was found to be around 102 genome copies. Regarding C. abortus–infected ewes, fewer than 106 genome copies were detected in placental samples without lesions associated with EAE (Table 2). Results from fetal tissues were more variable but fetal hair coat and vaginal swabs from ewes that did not display lesions consistent with EAE contained equal or fewer than 103 genome copies. Based on these results, arbitrary cutoff points were established for the interpretation of PCR results (Table 3). All samples obtained from control animals were negative for T. gondii or C. abortus but positive for IC1 and IC2. The samples collected from animals displaying low or moderate levels of infection were also positive for the internal controls.
DNA copy numbers detected in ewes orally infected with Toxoplasma gondii or Chlamydophila abortus.
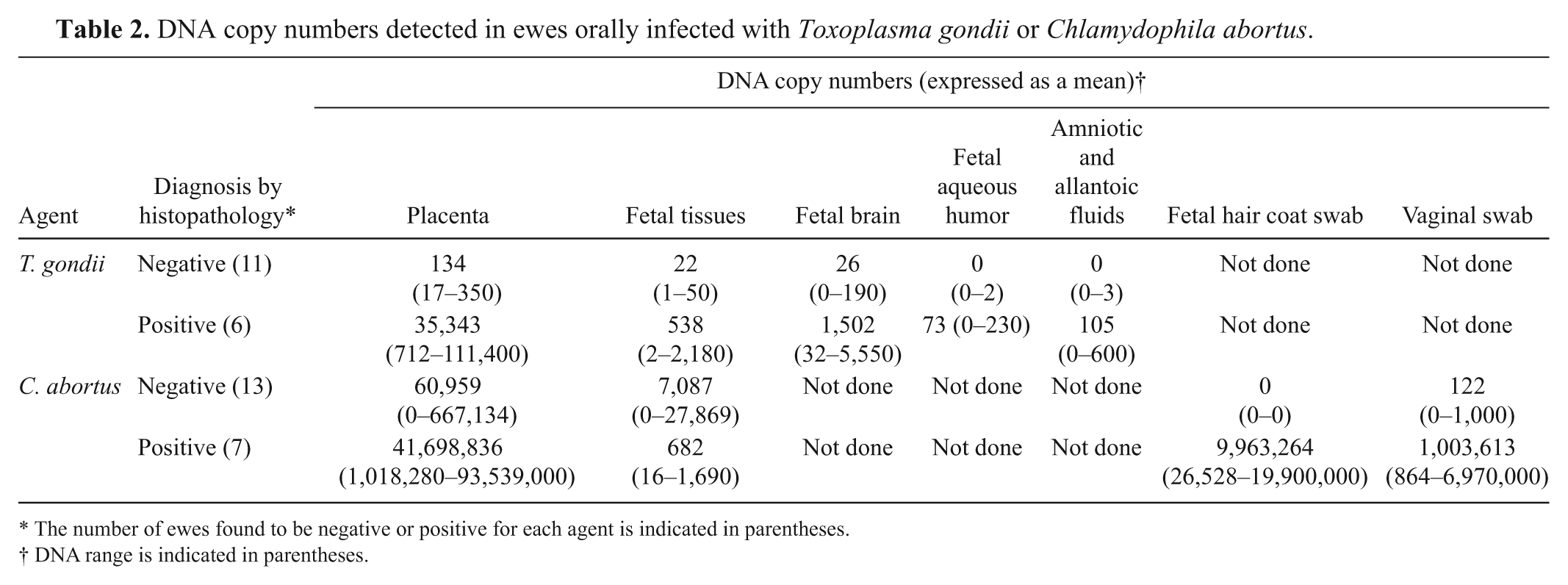
The number of ewes found to be negative or positive for each agent is indicated in parentheses.
DNA range is indicated in parentheses.
Criteria used for diagnosis of toxoplasmosis or enzootic abortion of ewes (EAE) based on detected DNA copy numbers and critical threshold cycle (Ct) values for each target gene.*
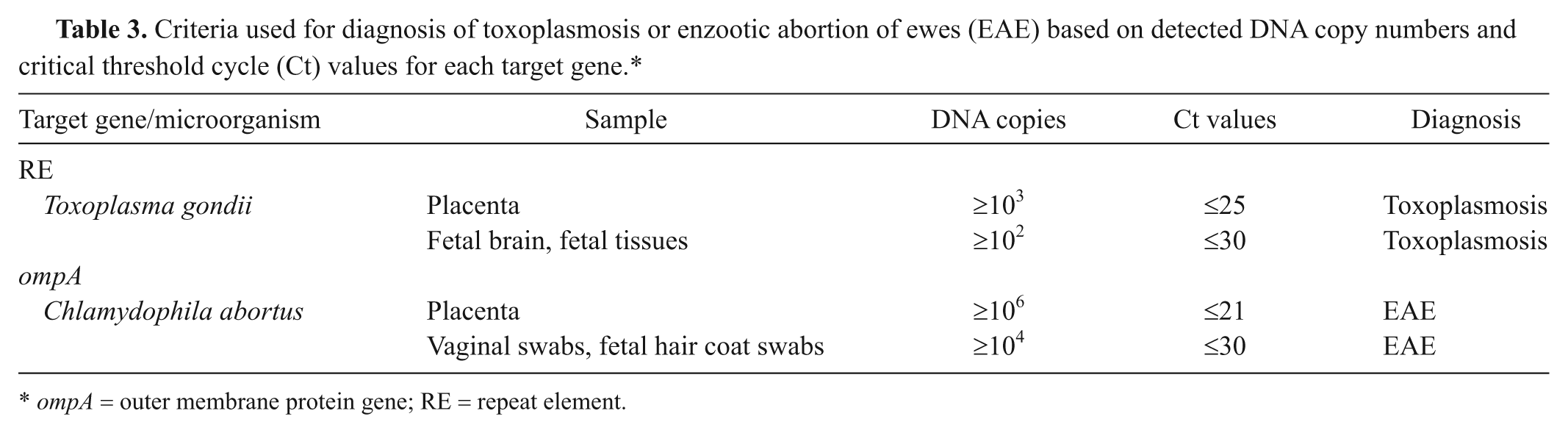

ompA = outer membrane protein gene; RE = repeat element.
Application of the duplex real-time PCR to field samples
A total of 489 samples were collected from animals associated with ovine abortion on 66 Irish farms. When DNA was extracted from the collected field samples and subjected to the duplex real-time PCR, it was found that 10% and 55% of the samples were positive for T. gondii and C. abortus, respectively (Table 4). Chlamydophila abortus was detected most frequently and in the greatest concentrations in placenta. For T. gondii, the most suitable samples were placenta, fetal hair coat swabs, fetal brain, and other fetal tissues such as lung, liver, and spleen.
Results of field samples tested by the duplex real-time polymerase chain reaction.*
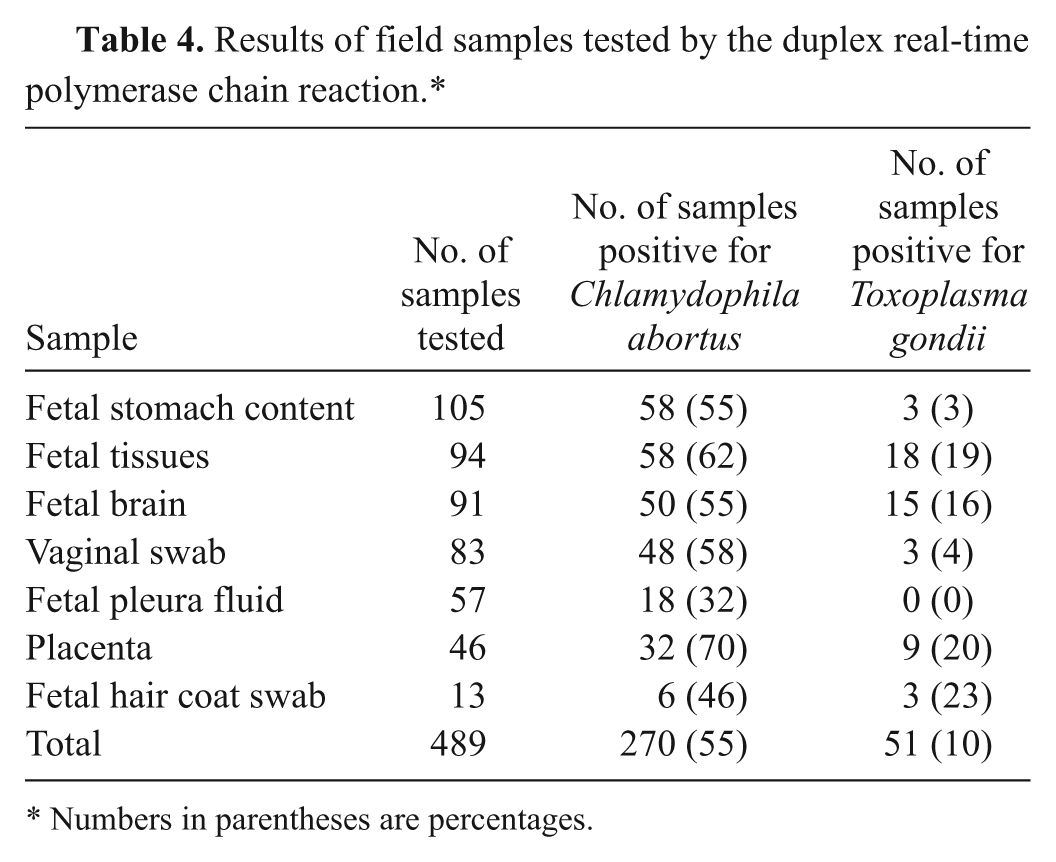
Numbers in parentheses are percentages.
Diagnosis of farm abortions: real-time PCR versus standard methods
Based on the real-time PCR results, EAE was diagnosed in 20% of the 66 farms, whereas 14% of the investigations indicated toxoplasmosis (Table 5). There was evidence of dual infection on 6% of the farms. Furthermore, C. abortus was detected in samples from another 23% of the farms, while 9% of the farms had evidence of the presence of both T. gondii and C. abortus. However, the PCR test did not detect any DNA copy number in 29% of the farms. When the diagnosis was reached by the standard methods, EAE was diagnosed in 12% of the Irish farms investigated, while 17% of the farms were positive for toxoplasmosis (Table 5). Dual infection was diagnosed in 3% of the farms. Campylobacter was associated with abortion on 6% of farms, while Escherichia coli and Salmonella accounted for 3% of abortions each, and 2% of abortions were associated with Listeria monocytogenes. A diagnosis was not reached in cases from 54% of the farms.
Causes of ovine abortion on 66 Irish farms using the standard methods and the duplex real-time polymerase chain reaction (PCR).*
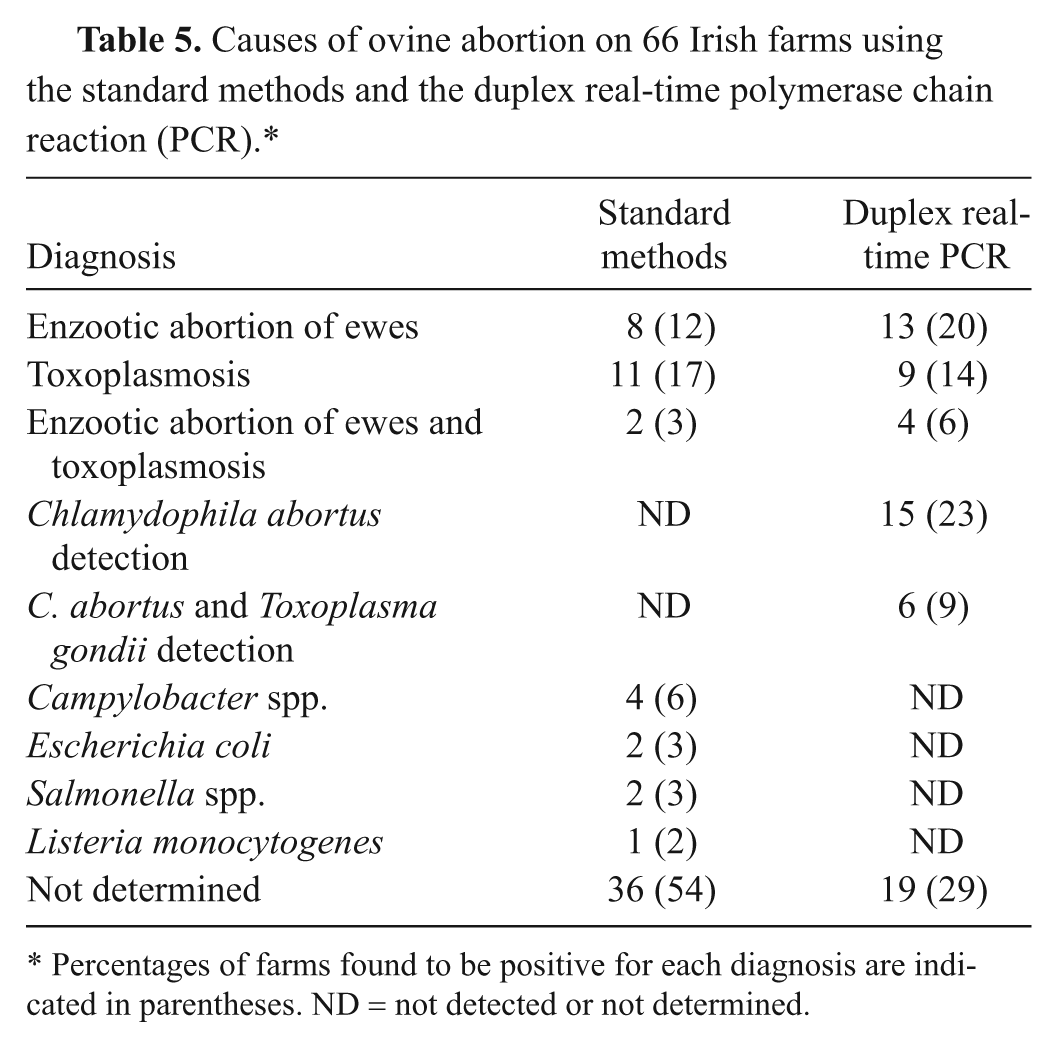
Percentages of farms found to be positive for each diagnosis are indicated in parentheses. ND = not detected or not determined.
Correlation between the standard methods and the duplex real-time PCR
Nine out of the 10 (90%) cases diagnosed as EAE by the standard methods were positive by the duplex real-time PCR assay (Table 6). All the cases diagnosed as EAE following histopathological examination or ELISA testing (cases 1–6, 14, 15) were positive by the duplex real-time PCR. Seven out of the 8 cases found to be positive by the chlamydial LPS detection test (cases 2–8, 14) were also diagnosed as EAE by the duplex real-time PCR. However, only 10 of the 17 cases (59%) classed as EAE using duplex real-time PCR were diagnosed as EAE using standard methods. The duplex real-time PCR test detected 8 additional cases of EAE (cases 9–13, 24–26).
Details of diagnostic results for the 66 Irish farms based on standard methods and the duplex real-time polymerase chain reaction (PCR).*
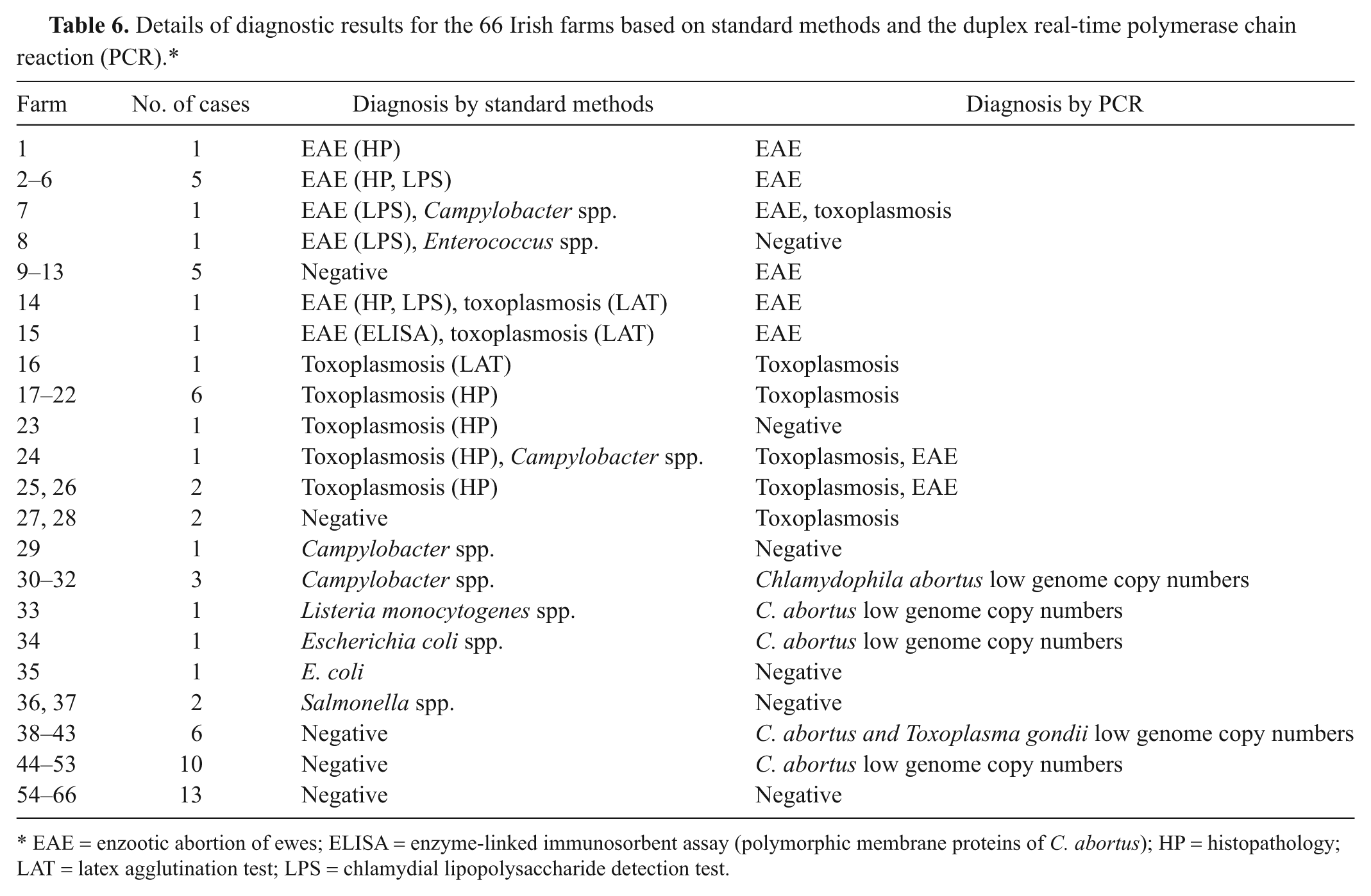
EAE = enzootic abortion of ewes; ELISA = enzyme-linked immunosorbent assay (polymorphic membrane proteins of C. abortus); HP = histopathology; LAT = latex agglutination test; LPS = chlamydial lipopolysaccharide detection test.
Regarding the diagnosis of toxoplasmosis, 10 out of the 13 (77%) cases diagnosed as toxoplasmosis using standard methods were also found to be positive by the PCR (Table 6). Nine out of the 10 cases (17–26) found to be positive based on histopathological findings were also diagnosed as toxoplasmosis by the duplex real-time PCR. Only 1 out of the 3 cases (14–16) diagnosed as toxoplasmosis by the LAT method was positive by PCR. The duplex real-time PCR gave a positive result for 3 additional cases of toxoplasmosis.
Significantly, C. abortus was detected at low numbers in samples from 21 farms (cases 30–34, 38–53). Toxoplasma gondii DNA was also found at low levels in 6 of these 21 farms (cases 38–43).
Genotype analysis of T. gondii isolates by RFLP
Toxoplasma gondii DNA isolated from the field samples was subjected to genotype analysis by RFLP. Fifteen out of the 19 (79%) isolates were found to be genotype II. The remainder of the isolates (21%) was type III. The genotype III isolates were not associated with any histopathological lesions, but 60% of the genotype II isolates were related to cases with significant lesions consistent with toxoplasmosis.
Discussion
In the current study, a duplex real-time PCR test capable of detecting DNA from T. gondii and C. abortus was first validated using samples from experimentally infected sheep prior to its application to field material. All samples obtained from control animals were negative, while all the samples obtained from the experimentally infected animals were positive. In general, high DNA copy numbers were detected in samples from ewes displaying lesions consistent with the challenge pathogen.
When the duplex real-time PCR was applied to cases of ovine abortion, T. gondii and C. abortus DNA was detected in samples from 29% and 55% of the farms, respectively. Standard diagnostic methods available in the Irish Regional Veterinary Laboratories were able to diagnose infection related to toxoplasmosis and EAE in 17% and 12% of the farms, respectively. The higher detection of these pathogens using real-time PCR over standard diagnostic techniques may be due to the identification of an increased number of true positives identified by PCR and not detected by less sensitive methods, subclinical infection, contamination during sampling, or samples from vaccinated sheep as recently described. 21
Asymptomatic shedding may occur as a result of subclinical infection or following clinical recovery, so results of molecular methods with high sensitivity must be assessed with caution regarding the clinical relevance of the finding. 2 In this respect, real-time PCR has been proposed to play a role in distinguishing between colonization and clinical disease. 20 Indeed, a significant advantage of real-time PCR technology is the ability to quantify genome copy number. In the present study, a cutoff point was established for the PCR assay to differentiate detection of the pathogen from a pathogen burden associated with significant pathological lesions. The cutoff values were derived from a series of experimental infections where pregnant ewes were challenged with either T. gondii or C. abortus, and genome copy numbers were correlated with the presence of characteristic lesions. When these cutoff values were applied to field cases, the number of abortion outbreaks diagnosed as toxoplasmosis or EAE fell from 29% to 14% and from 55% to 20% of the farms, respectively. On 6% of these farms, there was evidence of dual infection. Clearly, these figures are much closer to those obtained using standard diagnostic methods (17% for toxoplasmosis, 12% for EAE, and 3% for dual infection), with a high correlation between the PCR assay results and those of diagnostic methods currently in use.
Although there was a very good correlation between high PCR copy numbers and the presence of characteristic histopathological lesions in the placenta, there was 1 case in which lesions associated with toxoplasmosis were observed but no diagnosis was reached based on the PCR results. In this respect, there are some issues to be considered as sources of possible errors in connection with preanalytical procedures for PCR assays, such as sampling, aliquoting, and splitting. 1 It has been reported previously by the authors that the pooling of placental samples does not improve the detection rate of T. gondii by real-time PCR. 14 In addition, if samples are not taken from the most infected locations, then the yield of pathogen DNA may be insufficient for a diagnosis.
All the samples that displayed significant lesions consistent with EAE as well as those found to be positive by the ELISA were confirmed as EAE by the PCR. Similarly, most of the cases (7/8) diagnosed as EAE by the commercial LPS detection test were also positive by the PCR. The only case found to be negative did not display the typical histopathological lesions either, indicating that the LPS detection test may provide false-positive results, as antibodies used in this method are not 100% specific for C. abortus. The commercial LPS detection test is an antigen detection test used to diagnose human urogenital Chlamydia trachomatis infections and is not optimized for ovine samples. False-positive results are usually associated with cross-reactivity with other bacteria present in the reproductive tract. In the context of ovine abortions, false-positive reactions can arise where swabs are taken from fetal membranes, vaginal discharge, or wool coat that are contaminated with a range of the bacteria known to cause problems such as C. pecorum and Chlamydophila psittaci or other bacteria including Staphylococcus aureus, Haemophilus aegyptius, Klebsiella pneumoniae, Gardnerella vaginalis, Neisseria gonorrhoeae, E. coli, Streptococcus agalactiae, Moraxella lacunata, Salmonella enterica, Acinetobacter lwoffii, and Acinetobacter calcoaceticus. 30
Regarding diagnosis of toxoplasmosis, there was a good correlation between the results of histopathological examination and PCR, but only 1 of the 3 farms found to be positive by the LAT were diagnosed as toxoplasmosis using PCR. No histopathological lesions consistent with a diagnosis of toxoplasmosis were reported in samples from these farms, indicating that the presence of antibodies is not always indicative of active infection. Some studies have reported that LAT is less sensitive than other serological assays.23,29
As previously reported, the placenta is the sample of choice for the diagnosis of EAE. 15 However, other samples such as fetal stomach contents, fetal organs including brain, as well as fetal hair coat and vaginal swabs could be considered as alternatives when placenta is not submitted to the laboratory. For T. gondii, the most reliable samples were not only placenta and fetal hair coat swabs but also fetal tissues. As a result of using a wider range of samples and a highly sensitive PCR assay, 3 and 8 more cases of toxoplasmosis and EAE, respectively, were identified when compared with standard methods.
Identification of the genetic group of T. gondii involved in clinical cases is a useful adjunct to diagnosis. 31 Little genetic typing has been performed on T. gondii isolates from sheep to date. 5 In the current study, genotyping of the toxoplasmosis cases has shown that the majority of infections (79%) were caused by genotype II of T. gondii. The rest of the isolates (21%) were identified as genotype III. Similarly, the clonal type II lineage and its closely related genotypes have accounted for 68% of T. gondii isolates in the United States. 6 The type III lineage accounted for 14% of the strains and was the second most prevalent genotype. Furthermore, it was observed in the current study that the genotype II isolates tended to be associated with significant lesions consistent with toxoplasmosis. In contrast, typical histopathological lesions were not observed in the current study in the cases where genotype III isolates were identified. Two studies carried out in the United Kingdom and France indicated that all the isolates from adult sheep that aborted or were persistently infected with T. gondii were clonal type II.7,26 In Denmark, 54% of the type II isolates of T. gondii were reported to be from aborted lambs. 18 The remaining isolates were obtained from healthy sheep. These results suggest that type II is the predominant lineage in sheep, and is capable of causing abortion.
There are few detailed epidemiological studies documenting the frequency of ovine chlamydial infections. Most studies to date have been based on materials submitted for routine laboratory examination by standard methods. 21 However, it is difficult to make accurate comparisons between such studies due to differences in study design and criteria, flock size and management, prevalence of other infectious agents, and the detection methods applied. In 2011, the seroprevalence of Chlamydophila was reported in 32 randomly selected German sheep flocks with an average abortion rate lower than 1%. 21 It was estimated in the 2011 study that 94% of the tested flocks were serologically positive with ongoing infection in nearly half (47%) of the flocks, and that PCR and DNA microarray testing, together with sequencing, revealed the presence of phylum Chlamydiae in 78% of the flocks. The species most frequently detected was C. abortus (50%), followed by C. pecorum (47%) and C. psittaci genotype A (25%), representing 39%, 37%, and 19% of the flocks, respectively. 21 Previously, a PCR assay was used for the detection of C. abortus in samples of uterine tissue collected from pregnant sheep in an abattoir and resulted in a prevalence of 31%, with a significantly higher prevalence in pregnant animals (47%). 25 In the current study, the duplex real-time PCR assay detected C. abortus in 55% of the studied farms, suggesting that the bacterium may be more widely disseminated in sheep flocks than previously documented.
The future for improvements in ovine abortion diagnosis lies in the development of tests based on real-time PCR, which provide a quantitative result and require few handling steps. 31 While further field studies are required, the duplex real-time PCR described in the current study has proved to be a useful tool, not only for detection of both C. abortus and T. gondii, but also for determining pathogen burden levels. Quantification of the pathogen load provided a significant advantage in terms of assigning significance to results compared to detection alone.
Footnotes
Acknowledgements
The authors wish to express their gratitude to Dr. Elaine McCarthy for assistance with the T. gondii genotyping. The authors would also like to acknowledge Dr. Erin Williams and Dr. Mary Sekiya for their helpful discussions.
a.
DNeasy Blood and Tissue Kit, Ireland Qiagen Ltd., West Sussex, UK.
b.
High Pure PCR Template Preparation Kit, Roche Diagnostics, Mannheim, Germany.
c.
Toxovax, Intervet Ireland Ltd., Dublin, Ireland.
d.
NanoDrop 1000 spectrophotometer, Fisher Scientific Ireland, Dublin, Ireland.
e.
TaqMan Probe, Life Technologies Ltd., Paisley, UK.
f.
Eurofins MWG Operon, Ebersberg, Germany.
g.
ABI PRISM 7500 thermocycler, Life Technologies Ltd., Paisley, UK.
h.
7500 System software, Life Technologies Ltd., Paisley, UK.
i.
SPSS 15.0, SPSS Inc., Chicago, IL.
j.
High Pure PCR Template Preparation Kit, Roche Diagnostics Ltd., Burgess Hill, UK.
k.
Clearview Chlamydia Test, Unipath, Bedford, UK.
l.
Latex Agglutination Test (LAT) Toxoreagent, Mast Group Ltd., Bootle, Merseyside, UK.
m.
Pourquier ELISA, Pourquier Institute, Montpellier, France.
n.
New England BioLabs, Beverly, MA.
o.
SYBR Safe, Life Technologies Ltd., Paisley, UK.
Declaration of conflicting interests
The author(s) declared the following potential conflicts of interest with respect to the research, authorship, and/or publication of this article: An invention disclosure based on this work, titled “Multiplex Diagnostic Real Time PCR Test for Ovine Abortions,” was registered by the UCD Innovation and Technology Transfer Centre (NovaUCD) on March 10, 2010 (reference no. 2005402).
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This work was supported by the Research Stimulus Fund of the Irish Department of Agriculture, Fisheries and Food.
