Abstract
Introduction
Materials and methods
Identification of target sequences and generation of a control plasmid
Sequence analysis was performed from all BLV full-length
Primers and probes used in the current study.
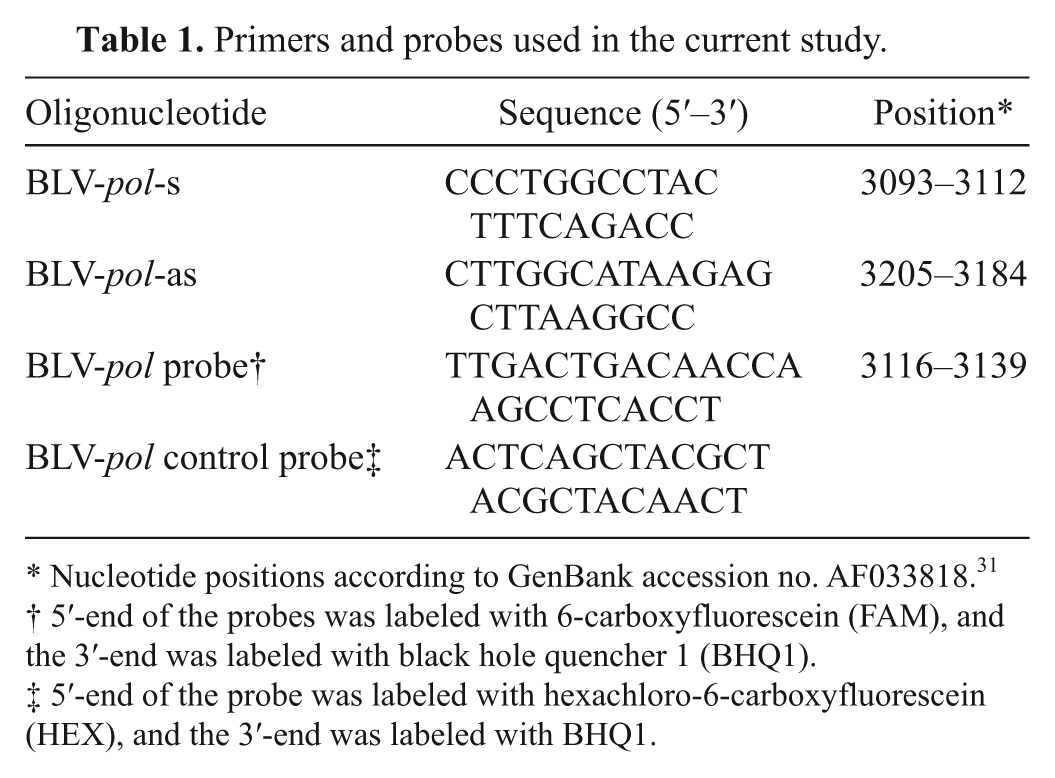

Nucleotide positions according to GenBank accession no. AF033818. 31
5′-end of the probes was labeled with 6-carboxyfluorescein (FAM), and the 3′-end was labeled with black hole quencher 1 (BHQ1).
5′-end of the probe was labeled with hexachloro-6-carboxyfluorescein (HEX), and the 3′-end was labeled with BHQ1.
A control plasmid was constructed for the BLV
The concentration of the plasmid was determined spectrophotometrically, and the exact number of plasmid molecules was calculated using the formula [X (g/µl DNA)/(length in nucleotides × 660)] × 6.022 × 1023 = number of molecules/µl. The lower detection limit of the real-time PCR was determined by amplifying serial 10-fold dilutions of the plasmid pCR2.1-BLV-
In addition to the control plasmid, DNA from persistently BLV-infected PO714 cells was used to investigate assay sensitivity by serial dilution of DNA equivalent to 1,000, 100, 10, 1, 0.75, 0.5, 0.25, and 0.1 cells in 100 ng/µl of DNA derived from the BLV-negative cell line PO. The dilutions performed in triplicate were subsequently investigated by real-time PCR.
Samples and DNA isolation
A total of 350 bovine PBMC samples were analyzed. Ethylenediamine tetra-acetic acid–blood samples were collected from 144 serologically positive animals (Table 2), which included 133 samples from naturally infected animals from Europe (Croatia, Germany, Poland, Russia, Slovenia, Sweden, Ukraine), Argentina, New Zealand, and Saudi Arabia and 11 samples from experimentally BLV-infected cattle in Germany. A second set of EDTA-blood samples were derived from 168 animals in Europe (Croatia, Germany, Poland, Slovenia, Sweden) and Argentina. No serological test could be performed from 38 samples from Brazil and Egypt as only PBMC-derived DNA had been obtained (Table 2).
Samples analyzed with the
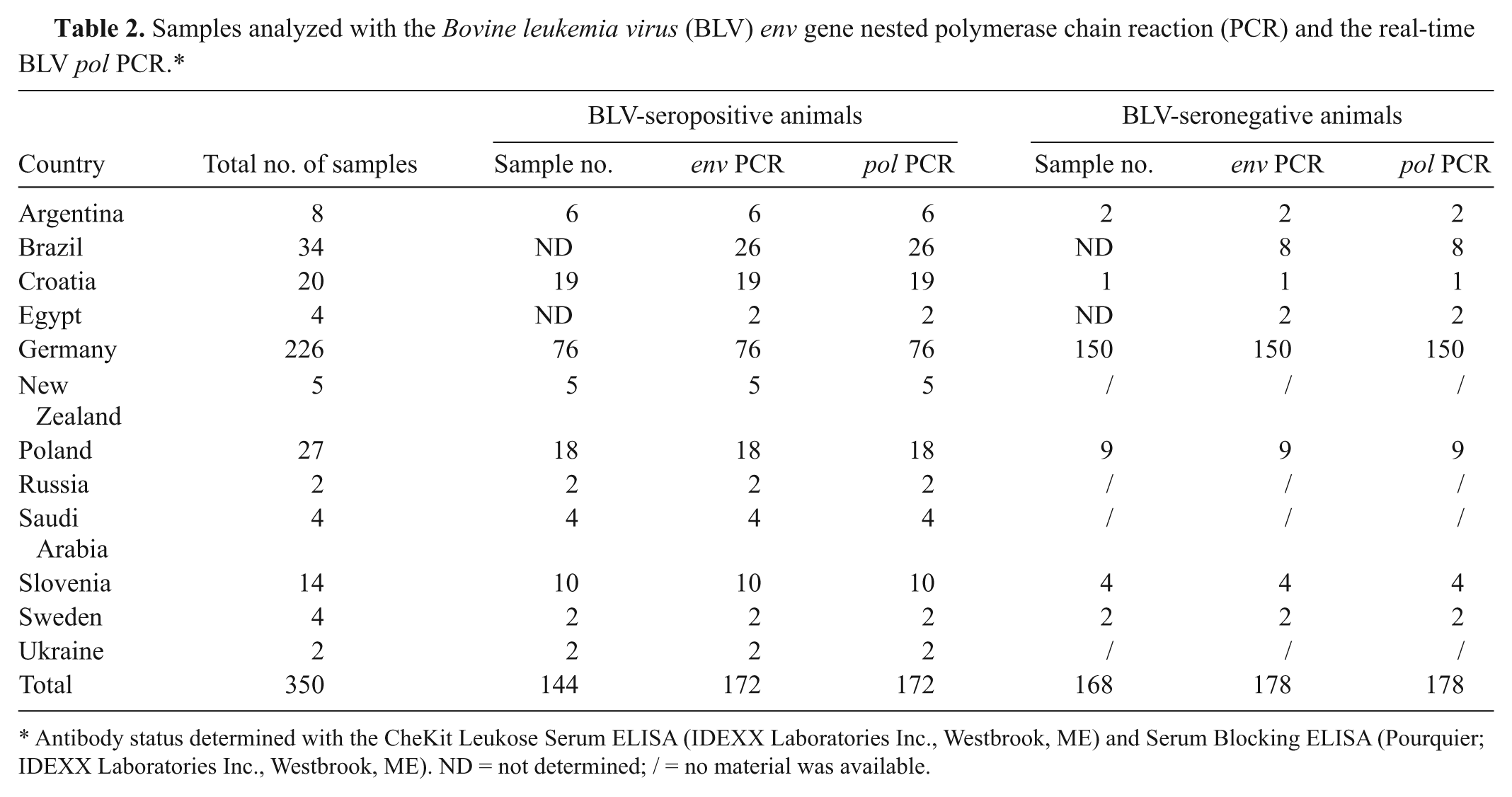
Antibody status determined with the CheKit Leukose Serum ELISA (IDEXX Laboratories Inc., Westbrook, ME) and Serum Blocking ELISA (Pourquier; IDEXX Laboratories Inc., Westbrook, ME). ND = not determined; / = no material was available.
Cell lines persistently infected with BLV (PO714),
Peripheral blood mononuclear cells were isolated from EDTA-blood samples using density gradient
d
centrifugation according to the manufacturer’s instruction. DNA isolation from the PBMC and cell lines was performed using a commercial kit
e
following the manufacturer’s instructions. The amount of DNA was measured spectrophotometrically
f
and by gel electrophoresis; 0.5 µg was used in the BLV
Real-time PCR based on the conserved BLV pol gene
The PCR was performed using a 20-µl reaction volume and the TaqMan universal PCR master mix
g
including AmpliTaq Gold DNA Polymerase, AMPErase UNG, deoxyribonucleotide triphosphate with 2′-deoxyuridine 5′-triphosphate, and passive reference dye (5-carboxy-X-rhodamine [ROX]). For one reaction, the assay was optimized to 10 µl of universal PCR master mix, 2 µl of DNA, 0.6 µl each of the
BLV env nested PCR, RFLP analysis, and ELISA
Using the primers BLV-
Results
Identification of target sequences
The comparative sequence analysis performed with the available full-length BLV
Sequence alignment of the
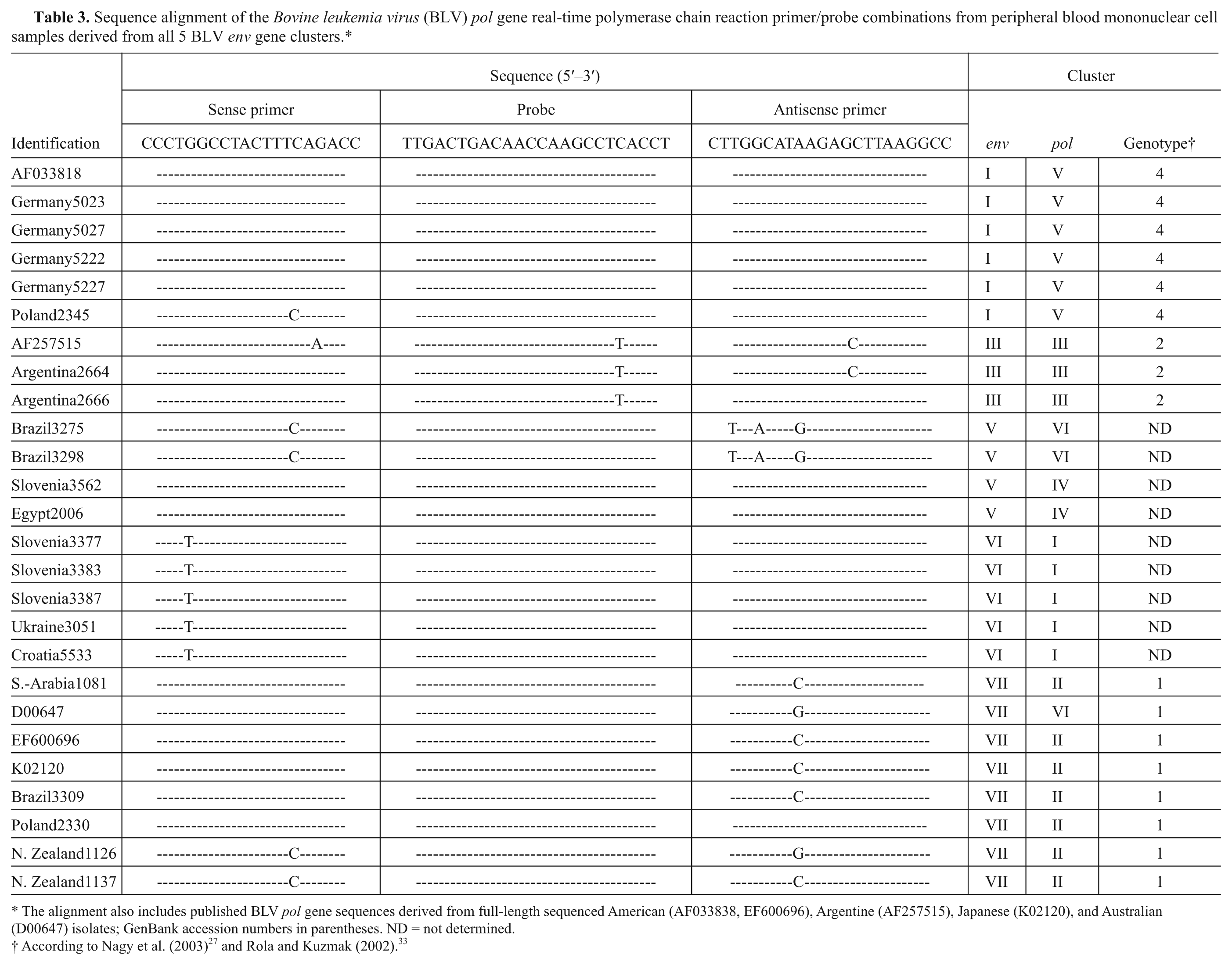
The alignment also includes published BLV
According to Nagy et al. (2003) 27 and Rola and Kuzmak (2002). 33
As control plasmid, a 112-bp fragment (nt 3088–3205, AF033818) corresponding to the BLV
Sensitivity and specificity of the real-time BLV pol PCR
The annealing temperature of the real-time BLV
For further assay analysis, serial dilutions of DNA derived from persistently infected PO714 cells in 100 ng/µl of BLV-negative DNA revealed that a DNA equivalent of one PO714 cell could still be detected. Fewer amounts of PO714 cell–derived DNA failed to reproducibly generate a positive reaction. Additional serial dilution experiments using DNA derived from PBMCs of a BLV-infected animal with PL showed that dilutions (in 100 ng/µl of BLV-negative DNA) containing 1 × 10−6 to 1 × 10−8 µg DNA were still positive.
To exclude the amplification of other bovine retroviruses, BIV- (FBS-BIV 198) and BFV- (BHK21-C13) infected cell cultures were investigated. The results proved that BIV and BFV proviral DNAs are not detected by the assay. Comparison of the primers and probes to the 2 published JDV sequences Pul01 and Kal07 revealed that no more than 5 bases are complementary, which makes it highly unlikely that the real-time PCR will amplify JDV proviral DNA.
Real-time BLV pol PCR
From 312 samples with known serological BLV status as determined by commercial ELISA,
l
all 144 samples derived from BLV-seropositive animals were detected by the real-time BLV
BLV env nested PCR and RFLP analysis
All DNA samples were also investigated with the BLV
From the 144 BLV-seropositive samples, 29 were selected for RFLP analysis. The results revealed that 5 of the 7 described RFLP-based BLV genotypes
13
were present in the sample collection. Ten samples belonged to variant A (Belgian subtype) as the 444-bp BLV
Discussion
The current study introduces a fast, sensitive, and specific method for the detection of BLV provirus sequences in PBMC of BLV-infected cattle. In contrast to most of the previous PCR assays that targeted the BLV
The lack of amplification using DNA from persistently BIV- and BFV-infected cells verified the specificity of the assay with regard to other bovine retroviruses. Low sequence homologies between the primer and probe sequences to JDV also seem to exclude the artificial detection of JDV with the BLV
Real-time PCR has many advantages over nested PCR. Real-time PCR is a shorter and less cumbersome procedure and is more amenable to high throughput testing. Furthermore, real-time PCR is less prone to cross-contamination. In summary, the described real-time method is suitable for detecting BLV provirus in infected animals.
Footnotes
Acknowledgements
The authors thank Kerstin Wink-Kruschke and Gabriele Czerwinski for technical support; Asia Gorska for the primer design; Martin Beer, Martin Groschup, and Anne Balkema-Buschmann for providing PBMC samples from BLV-negative animals; Roland Riebe for providing the cell lines PO714, FBS-BIV 198, and BHK21-C13; Mario Ziller for statistical analysis; and Shane Peterson, Murdoch University in Washington, for performing the sequence comparison analysis with regard to JDV.
a.
GCG Wisconsin software package, Accelrys Inc., San Diego, CA.
b.
Beacon Designer 2.06, PRIEMER Biosoft International, Palo Alto, CA.
c.
Eurofins MWG Operon Ebersberg, Bavaria, Germany.
d.
Histopaque-1077, Sigma-Aldrich, St. Louis, MO.
e.
QIAamp® DNA Blood Mini Kit, Qiagen, Hilden, North Rhine-Westphalia, Germany.
f.
Spectrophotometer DU 640 Beckman Coulter Inc., Fullerton, CA.
g.
TaqMan® universal PCR master mix, Applied Biosystems, Foster City, CA.
h.
Mx3000P real-time PCR system, Agilent Technologies Inc., Santa Clara, CA.
i.
MXPro4 software, Agilent Technologies Inc., Santa Clara, CA.
j.
Thermocycler Primus 25/96, MWG Biotech AG, Ebersberg, Bavaria, Germany.
k.
Restriction endonucleases, New England Biolabs, Ipswich, Suffolk, UK.
l.
CheKit Leukose Serum ELISA and Serum Blocking ELISA [Pourquier] by IDEXX Laboratories Inc., Westbrook, ME.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: The work was supported by the Federal Government of Germany and by the EU Network of Excellence, EPIZONE (contract no. FOOD-CT-2006-016236).
