Abstract
Members of the genus Francisella (viz., F. noatunensis subsp. orientalis [Fno] and F. noatunensis subsp. noatunensis) have been described as causative agents of chronic granulomatous and pyogranulomatous lesions in wild and cultured fish species. In the present study, 68 archived formalin-fixed, paraffin-embedded (FFPE) tissues from several fish species, collected at different geographical locations from 2000 to 2011, were analyzed using a real-time polymerase chain reaction assay for the detection of the Fno intracellular growth loci C (iglC) gene and by immunohistochemistry for the demonstration of Fno antigens. The results revealed a high correlation between these 2 diagnostic techniques validating their use for the diagnosis of Fno infection in archived FFPE tissues and confirming the presence of Fno in fish species from the Cari y years of the present century.
Introduction
During the late 1990s and early 2000s, fish disease diagnosticians reported the presence of small, Gram-negative, pleomorphic coccobacilli associated with granulomatous and pyogranulomatous lesions in cultured fish submitted during high mortality peaks. The organisms were acid-fast negative and resembled the salmonid pathogen Piscirickettsia salmonis; thus, they were referred to as Rickettsia-like organisms (RLOs) and/or Piscirickettsia-like organisms (PLOs) for several years.3,7,8 In Taiwan, reports of RLOs causing disease in fresh-, brackish-, and salt water pond–cultured tilapia (Oreochromis spp.) can be tracked to the early 1990s with farms reporting mortalities of up to 95% due to this pathogen.2,3,9 During the past decade, similar outbreaks suspected to be the result of infection with RLOs and PLOs were reported in cultured tilapia from Hawaii, Florida, Texas, and Georgia.7,8 Unfortunately, isolation and/or genetic characterization of the causative organism was not made at the time; thus, diagnostic confirmation of the cause of mortality was not achieved. With the recent description of Francisella noatunensis subsp. orientalis (Fno; synonym F. asiatica) as an etiologic agent of pyogranulomatous and granulomatous infections in wild and cultured fish, many of the previously diagnosed cases of RLOs and PLOs are thought to have been caused by Fno.2,4
In the past decade (2001–2011), Fno has caused substantial mortality in tilapia and in other important warm-water fish species cultured in the United States, Taiwan, the United Kingdom, Japan, Costa Rica, and other regions of Latin America.2,4,10,12 The bacterium not only has been isolated and found to cause disease in important worldwide culture species such as tilapia, three line grunt (or, chicken grunt; Parapristipoma trilineatum), and hybrid striped bass (Morone chrysops × M. saxatilis) but has been found in wild fish such as the guapote tigre (Parachromis managuensis) in Costa Rica and other parts of Latin America.2,4,10,12 The disease can present as an acute syndrome with few clinical signs and high mortality or as a subacute to chronic syndrome with nonspecific clinical signs, including anorexia, exophthalmia, and anemia. Upon macroscopic and microscopic examination, internal organs are enlarged and contain widespread, multifocal, slightly raised, white nodules. Histological examination of affected tissues reveals multifocal granulomatous inflammatory lesions containing numerous small, pleomorphic coccobacilli.2,4 The aim of the current study was to investigate the presence of Fno in archived formalin-fixed, paraffin-embedded (FFPE) tissues collected during the last 2 decades from different geographic locations and fish species utilizing immunohistochemistry (IHC) and a molecular diagnostic approach.
Materials and methods
Samples
Formalin-fixed, paraffin-embedded fish tissue samples from 68 diagnostic cases submitted between 2001 and 2011 were selected from the archives of University of Arkansas–Pine Bluff (California samples), Louisiana State University, School of Veterinary Medicine (Haiti, northwestern and southeastern U.S. samples), University of Florida, College of Veterinary Medicine (Jamaica samples), University of Florida Tropical Aquaculture Laboratory (Florida samples), Universidad Nacional de Costa Rica–Escuela de Medicina Veterinaria (Latin America samples), and Hawaii Department of Agriculture (Hawaiian samples). Selected cases had been previously diagnosed with chronic (often granulomatous) inflammatory lesions of either unknown etiology or likely the result of infection with RLOs or PLOs.
Positive and negative control tissue collection
Naïve healthy Nile tilapia (Oreochromis niloticus) fingerlings (mean weight: 20.5 g) were experimentally infected with Fno, Edwardsiella ictaluri, or Edwardsiella tarda following previously published protocols.12,13 The fish were obtained from a source with no history of Francisella infection and a subsample of the population was confirmed as negative for bacteria by culture on agar media a and real-time polymerase chain reaction (PCR) assay prior to use in the study.10,11 One hundred fish were maintained at 10 fish per tank in approximately 37-liter aquarium tanks with individual filters, air stones, and approximately 36 liters of unchlorinated fresh water at a temperature of 25°C. Three tanks were used per treatment, and 1 tank was used as a control. Fish in each tank were acclimatized for 1 week prior to challenge. The day of the challenge, approximately 108 CFU of Fno, E. ictaluri, or E. tarda were added to 3 tanks. Noninfected control fish were treated in a similar manner, but received sterile phosphate buffered saline (PBS). b Mortality was recorded every 12 hr for 8 days. Dead fish and survivors from each challenge were subjected to a complete clinical, bacteriological, molecular, and histopathological examination following previously published methods.10,11 For each treatment, spleen and gills samples were obtained from 6 different surviving fish at 8 days postchallenge. A small portion of each tissue (approximately 20 mg/tissue) served to extract DNA and quantify total genome equivalent (GE) loads of Fno per mg of tissue following previously published reports. 11 Another 2 portions of the tissues were fixed in formalin c for either 1 or 7 days. The fixed tissues were then processed with the rest of the unknown samples and served as either positive controls (Fno laboratory-infected tissues) or negative controls (noninfected naïve fish, E. ictaluri– or E. tarda–infected fish).
Data (both original and inverse sine transformed) obtained from experimental challenges were compared in an analysis of variance of a factorial arrangement of treatments using a commercial software package. d Where significance was found, post hoc pairwise comparisons were conducted with t-tests of least squares means. Differences were considered significant at P ≤ 0.05.
DNA isolation and Fno iglC gene real-time PCR
Ten serial sections, each 5 µm in thickness, were made from each paraffin block. Sections were cut with a microtome and collected separately into 1.5-ml tubes. To avoid DNA contamination during the collection, the microtome was cleaned with 70% ethanol between samples, and a new area of the blade was used for each sample. Sections were deparaffinized with xylene, and DNA was extracted with a commercial kit e according to manufacturer’s instructions for paraffin-embedded tissues with a slight modification: paraffin was initially separated from the tissue sections (held in a 1.5-ml tube) by heating at 65°C for 10 min. Three ethanol (100%) rinses were performed after the use of xylene for paraffin removal, followed by complete evaporation in a vacuum centrifuge for 15 min. Elution from spin filters was performed once to a total volume of 50 μl. Samples were screened for the presence of Fno using a real-time PCR assay targeting a species-specific region of the intracellular growth loci (iglC) gene.10,11
Generation of mouse anti-Fno polyclonal antibodies
An Fno-specific whole-cell antiserum (anti-Fno) was obtained by suspending formalin-killed Fno cells in an adjuvant f at a concentration of 108 bacteria/ml. Briefly, Fno LADL 07-285A 12 was cultivated on agar plates a at 27°C for 3 days, and bacteria were harvested by scraping, washing twice, and suspending in 1× PBS. b Formalin c was added to the bacterial suspension at a final concentration of 0.7% and, after storage overnight at 4°C, an aliquot was streaked onto the agar media to confirm inactivation. The bacterial cell suspension was washed twice with 1× PBS b and emulsified with an equal volume of adjuvant. f The suspension was used for subcutaneous immunization of 4 BALB/c mice according to a previously reported method. 14 Serum from mice was tested by enzyme-linked immunosorbent assay for the presence of antibodies to Fno sonicate as previously described. 13 Mice were boosted twice (weeks 2 and 4 post–initial immunization) and, at week 6 postinoculation of the first suspension, were humanely euthanized. Blood was collected by cardiocentesis immediately after mice were rendered unconscious by CO2. Death was confirmed by cervical dislocation. Serum from mice had titer ratios of >1:52,000 to Fno sonicate.
Immunohistochemistry
Briefly, deparaffinized fish tissue sections were treated with 0.25% potassium permanganate g for 10 min to bleach melanin pigment present in melanomacrophage aggregates, followed by a rinse with 1% oxalic acid h to remove excess potassium permanganate. Tissues were washed with distilled water to clear potassium permanganate g from sections and incubated with peroxidase i for 5 min to block endogenous peroxidase activity. After the sections were rinsed in 0.05 M Tris buffered saline j (TBS), the sections were incubated in pronase k for 15 min at 37°C. Sections were rinsed 3 times with 1× TBS j and blocked utilizing a blocking agent l for 15 min, followed by 3 more TBS j washes. Tissue sections were suspended in TBS j containing the primary mouse anti-Fno diluted 1:10,000 overnight at 4°C. Dilutions of 1:1,000, 1:5,000, and 1:100,000 were also evaluated. The following morning, the samples were washed 3 times with TBS, j and a commercial kit m was utilized in accordance to manufacturer’s specifications for the detection and visualization of mouse antibodies. n The sections were then rinsed in running tap water, counterstained with hematoxylin, o coverslipped, and examined by light microscopy by 4 veterinary pathologists in a blinded study.
Results
Experimentally infected fish presented few mortality events by 8 days postchallenge. Fish challenged with Fno, E. ictaluri, and E. tarda experienced 13.3, 3.33, and 6.66 cumulative percentage mortality, respectively. No mortality events occurred in the PBS-treated fish. Surviving fish presented classical signs of francisellosis and edwardsiellosis in tilapia with the presence of splenomegaly and renomegaly with multifocal white nodulations.10,12 Fish challenged with Fno presented high Fno genome equivalents in the spleen (4.91 ± 0.77 log Fno GE/mg) and gills (2.48 ± 0.95 log Fno GE/mg). Fno was not detected in the fish challenged with either E. ictaluri or E. tarda. Upon real-time analysis of FFPE Fno-infected tissues, a significant difference was observed in tissues fixed for 1 day or 7 days (P < 0.05). Detection was greater in tissues fixed for only 1 day in comparison to those fixed for 7 days (Table 1). However, positive detection was observed in all Fno-infected FFPE positive control tissues (Table 1). No detection of Fno was obtained in any of the negative control fixed tissues. Of the 68 archive samples analyzed by real-time PCR specific for the Fno iglC gene, 47 resulted in cycle threshold (Ct) values lower than 40 (range: 25–37 Ct), indicating detection of Fno DNA (Table 1).
Molecular (iglC real-time polymerase chain reaction [PCR]) and immunohistochemical identification (IHC) of Francisella noatunensis subsp. orientalis (Fno) from formalin-fixed, paraffin-embedded fish tissues.*
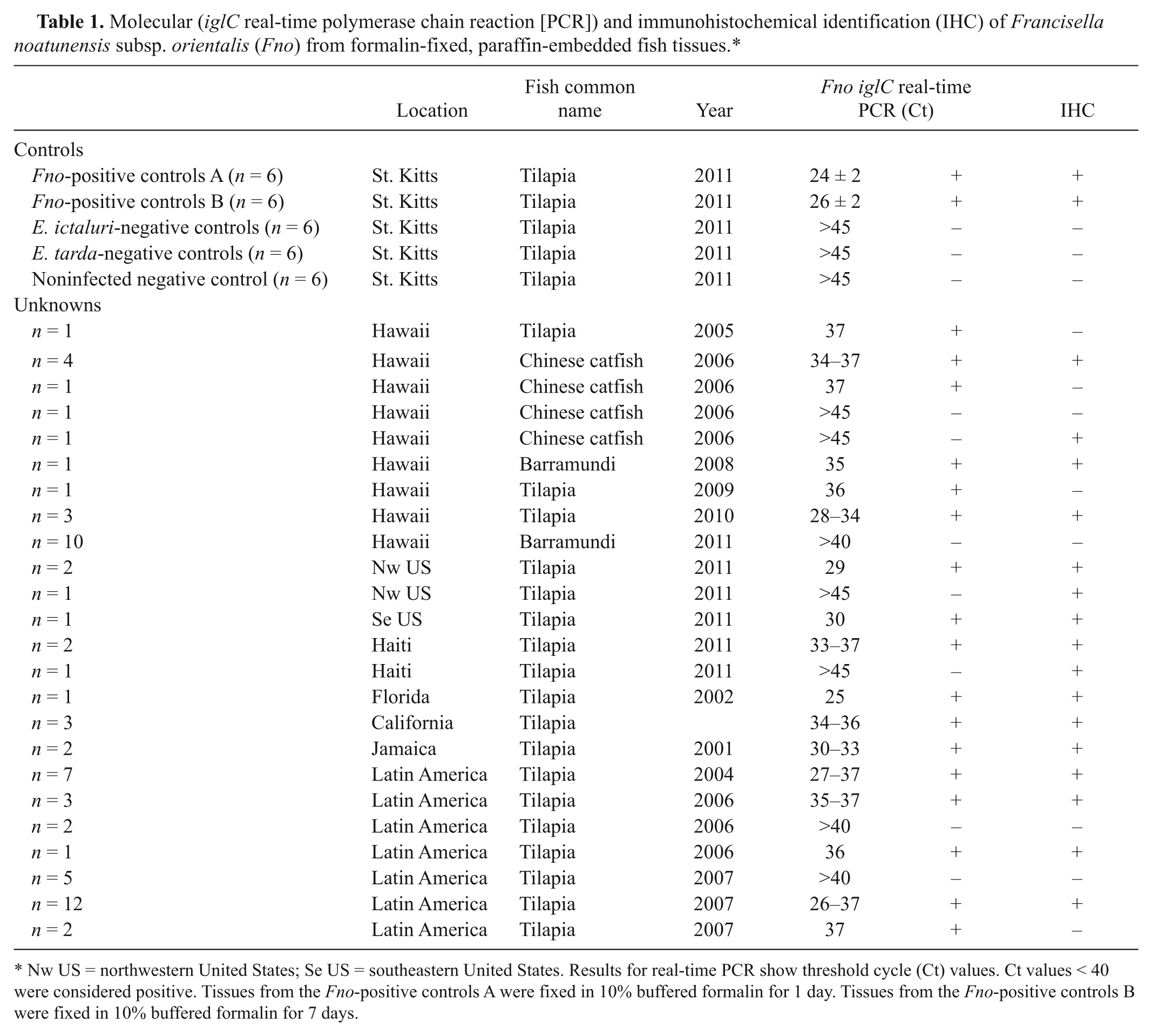
Nw US = northwestern United States; Se US = southeastern United States. Results for real-time PCR show threshold cycle (Ct) values. Ct values < 40 were considered positive. Tissues from the Fno-positive controls A were fixed in 10% buffered formalin for 1 day. Tissues from the Fno-positive controls B were fixed in 10% buffered formalin for 7 days.
Initially, different dilutions of the primary antibodies (mouse anti-Fno) were evaluated utilizing the experimentally infected positive and negative controls. The results indicated fewer background and good detection when using anti-Fno diluted 1:10,000. Histological and immunohistochemical analysis performed in the archive samples utilizing specific antibodies against Fno corroborated the results obtained from the real-time PCR assay. Of the 68 samples, 45 examined stained positively against Fno antigens by IHC (Table 1). Positive reaction was observed in different tissues including gills, liver, gastric epithelium, pericardium, endocardium, and meninges; however, the strongest and most consistent positive reaction was observed within and around splenic and renal granulomas (Fig. 1). Individual host cells contained numerous positive-stained organisms inside spacious intracytoplasmic vacuoles.
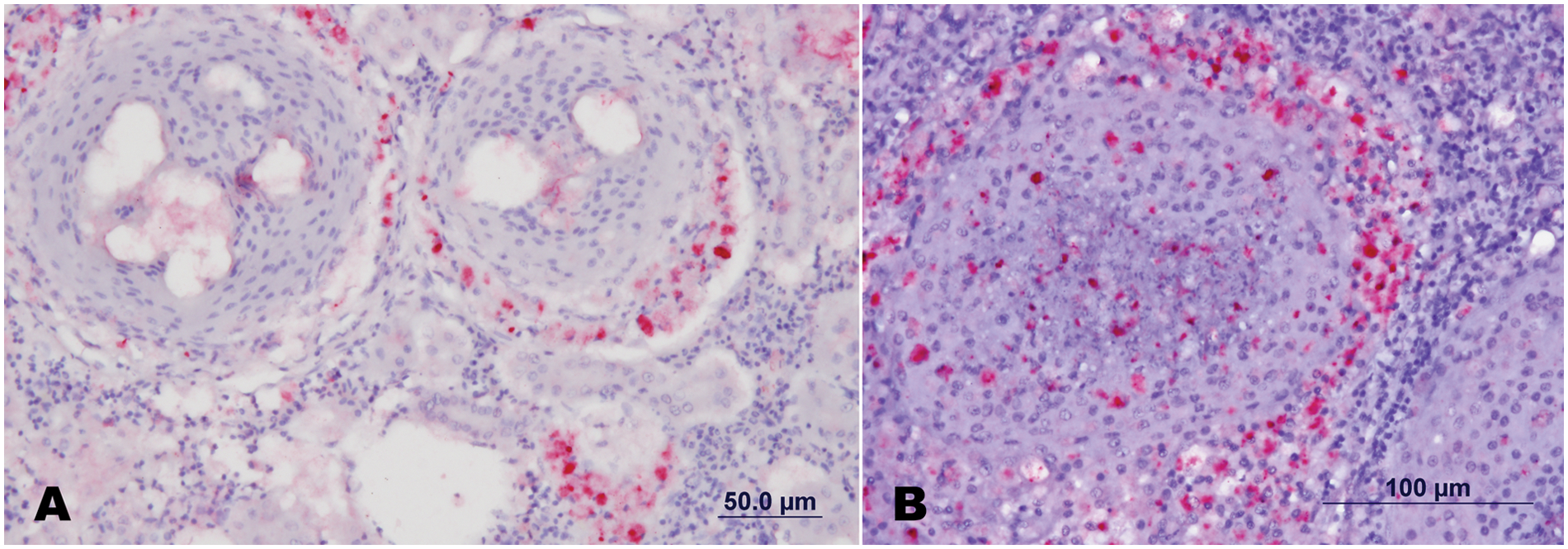
Francisella noatunensis subsp. orientalis (Fno)–infected fish tissue sections, showing positive reaction (red) for the polyvalent anti-Fno mouse serum. Spleen (
Discussion
Members of genera Mycobacteria, Nocardia, Francisella, Edwardsiella, Photobacterium, Piscirickettsia, Renibacterium, and Streptococcus have been described as etiological agents capable of inducing granulomatous disease in wild and cultured fish species worldwide.1,2,4,9 Although diagnostic methods have been described for a few of them, culture and identification of these and other fish pathogens is usually hampered by their fastidious nature, coinfection with other fast-growing bacteria, low levels of the organism present in the tissues, and/or the submission of inadequate samples to the diagnostic center.1,6,9,16 The current availability of highly specific and sensitive molecular and serological diagnostic techniques has facilitated the detection, description, and characterization of many previously unidentified or misdiagnosed pathogens from archived samples.5,6,16
In the current study, archived fish tissues (obtained from 8 different locations and more than 5 fish species and their hybrids) collected from the Caribbean, North and South America, and Hawaii during the last 2 decades and previously diagnosed with granulomatous inflammatory disease caused by “unknown,” “Rickettsia-like,” or “Piscirickettsia-like” organisms were subjected to Fno-specific IHC and real-time PCR analysis. In the majority of the cases, previous histological analysis revealed the presence of small pleomorphic bacteria within vacuolated macrophages located in areas of moderate to severe chronic inflammatory cell infiltration.7,8 Although conventional bacteriological approaches, like inoculation of agar media (including tryptic soy agar with 5% sheep blood, MacConkey agar, and brain heart infusion agar) resulted in no consistent bacterial isolation, the involvement of a bacterial pathogen was evident utilizing nonspecific histochemical (Gram and Giemsa) stains.
In the present study, real-time PCR analysis targeting the Fno iglC gene demonstrated the presence of high bacterial GE loads in 47 of the 68 analyzed samples. The specificity of this molecular assay for Fno was previously validated utilizing a wide range of warm-water fish pathogens (Mycobacterium marinum, Nocardia spp., Streptococcus agalactiae, Streptococcus iniae, E. ictaluri, E. tarda, Aeromonas spp., Vibrio spp., and Flavobacterium columnare). 11 As previously validated, when utilizing spiked tissues, the detection limit for the assay was determined to be approximately 1 organism, resulting in Ct of 38. 11 Although specific quantification of the bacterial load present in the archive samples utilized in the current study cannot be made, the data obtained not only confirms the causative agent of the previously described granulomatous inflammatory lesions as Fno, but also highlights the potential usefulness of this molecular technique for the retrospective diagnosis of fastidious bacteria in archived tissues.
Similarly, immunohistochemical analysis indicated positively stained tissues with Fno antigens in 45 of the 68 samples examined. The few samples (n = 3) that were positive by IHC but negative using real-time PCR were likely a reflection of the well-known limitations in the quality of DNA that can be extracted from archived paraffin-embedded tissue samples. The duration of fixation, age of paraffin blocks, sample size, storage environmental conditions, bacterial tissue load, and distribution are known to affect DNA extraction procedures, length of PCR targets, and the concentration of target DNA amplified.5,6,16 The effect of duration of fixation on DNA integrity was also demonstrated in the present study, when significantly higher Ct values were detected in tissues fixed in formalin for 7 days when compared to those same tissues fixed for only 1 day (Table 1).
Molecular tools for assessing the quantity and quality of nuclear DNA in a given forensic sample are currently available. 15 Similarly, future development of molecular diagnostic methods for the detection of fish “housekeeping genes” could potentially aid in the assessment of DNA degradation in samples and could facilitate the identification of false negatives.
The most likely explanation for the lack of correlation observed in the 5 IHC-negative and real-time PCR–positive samples may be the masking and/or destruction of Fno antigens during tissue preparation for IHC; this artifact has been reported in similar retrospective studies.5,6,16
In summary, the current study demonstrates the presence of Fno in FFPE fish tissues dating back to the early years of the current century utilizing a molecular and histological approach that can be successfully applied to archive tissues. These diagnostic techniques appear to have enormous potential in retrospective epidemiological investigations. In the present study, by successfully using these diagnostic methodologies, it was possible to confirm the presence of Fno-induced granulomatous disease in fish from 3 new geographical locations (Jamaica, Haiti, and Florida) as well as in a previously unreported fish species (Chinese catfish). Further research is necessary to elucidate the pathogenesis of Fno-induced granulomatous disease in fish and to understand the parameters that determine the virulence, clinical presentation, and the geographical and host species range of this bacterium.
Footnotes
Acknowledgements
The authors thank Dr. Kevin R. Macaluso at the Department of Pathobiological Science of Louisiana State University, School of Veterinary Medicine for the assistance in the creation of the antibodies utilized in this study, Dr. Matt Griffin at the Thad Cochran Warmwater Aquaculture Center and Mississippi State University for the critical review of this article, and Drs. Nigel Elliott and Gavin Bellamy from Jamaica for their assistance collecting and submitting samples.
a.
BBL Modified Thayer-Martin (MTM II) Agar, BD Diagnostic Systems, Sparks, MD.
b.
Phosphate Buffer Saline, Fisher Chemical, Fairlawn, NJ.
c.
10% buffered formalin, Fisher Chemical, Fairlawn, NJ.
d.
SAS statistical program, version 9.1.3, SAS Institute Inc., Cary, NC.
e.
DNeasy kit, Qiagen Inc., Valencia, CA.
f.
Titer-Max Gold adjuvant, TiterMax USA Inc., Norcross, GA.
g.
Potassium Permanganate, Fisher Chemical, Fairlawn, NJ.
h.
Oxalic Acid, Fisher Chemical, Fairlawn, NJ.
i.
Peroxidazed 1, Biocare Medical LLC, Concord, CA.
j.
Tris Buffered Saline, Fisher Chemical, Fairlawn, NJ.
k.
Carezyme III-Pronase, Biocare Medical LLC, Concord, CA.
l.
Biocare Background Sniper, Biocare Medical LLC, Concord, CA.
m.
MACH 4 Universal AP Polymer Kit, Biocare Medical LLC, Concord, CA.
n.
Vulcan Fast Red Chromogen, Biocare Medical LLC, Concord, CA.
o.
Harris Hematoxylin, Anatech Ltd., Battle Creek, MI.
Declaration of conflicting interests
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Funding
The author(s) disclosed receipt of the following financial support for the research, authorship, and/or publication of this article: This project was supported by Ross University, School of Veterinary Medicine (RUSVM) Research Program.
